Abstract
Brain metastasis (BM) is one of the main complications of many cancers, and the most frequent malignancy of the central nervous system. Imaging studies of BMs are routinely used for diagnosis of disease, treatment planning and follow-up. Artificial Intelligence (AI) has great potential to provide automated tools to assist in the management of disease. However, AI methods require large datasets for training and validation, and to date there have been just one publicly available imaging dataset of 156 BMs. This paper publishes 637 high-resolution imaging studies of 75 patients harboring 260 BM lesions, and their respective clinical data. It also includes semi-automatic segmentations of 593 BMs, including pre- and post-treatment T1-weighted cases, and a set of morphological and radiomic features for the cases segmented. This data-sharing initiative is expected to enable research into and performance evaluation of automatic BM detection, lesion segmentation, disease status evaluation and treatment planning methods for BMs, as well as the development and validation of predictive and prognostic tools with clinical applicability.
Similar content being viewed by others
Background & Summary
Brain metastases (BMs) represent the most common intracranial neoplasm in adults. They affect around 20% of all cancer patients1,2,3,4,5,6, and are among the main complications of lung, breast and colorectal cancers, melanoma or renal cell carcinomas1,2,3,4. The increasing availability of systemic treatments has improved the prognosis of patients with primary tumors, leading to an increase in the probability of developing BMs2,3,6,7.
BMs often appear as multiple lesions, with only around 25% of patients harboring a single BM2,8. On magnetic resonance imaging (MRI) studies, they are found to present contrast-enhancing features. Contrast-enhanced T1-weighted (CE-T1-W) MRI is the gold standard imaging sequence for BMs, providing information about lesion size, morphology and surrounding healthy structures7,9. T2-weighted imaging and fluid attenuation inversion recovery (FLAIR) MRI sequences are also used to help in identifying BMs, due to the surrounding edema found in many BM lesions1,5,7.
Treatment of BMs typically includes a combination of radiotherapy, chemotherapy, immunotherapy, targeted therapies, and/or surgery1,2,3. Radiotherapy schemes include whole brain radiation therapy and stereotactic radiosurgery (SRS). SRS is considered the standard of care in patients with limited metastatic burden6,7,9,10,11.
The clinical management of BMs undergoing radiotherapy requires time-consuming processes such as lesion identification and segmentation2,3,12. Time spent on those tasks could be reduced with the aid of semi-automatic or automatic computer-guided algorithms. Machine learning (ML) and deep learning (DL) techniques are being developed for different problems related to BMs, such as: automatic BM detection5,6,7,12,13,14, segmentation11,13,14,15 and differential diagnosis of BMs from other brain tumors7,12,16. AI algorithms may also reduce human errors in all of those jobs that result from heavy workloads, allowing for increased reproducibility6,12.
Another problem in which AI can be helpful is the differentiation between post-treatment BM progression and radiation necrosis, a transient inflammatory effect after SRS. These two situations have overlapping features on MRI sequences, which makes it challenging to distinguish them visually7,9,10. Incorrect classification leads to unnecessary treatments and substantial patient harm. For this reason, AI methods have have been developed to automatically distinguish them7,9.
Finally, the development of prognostic and predictive metrics using the information contained in medical images is of the utmost importance because of the clinical implications. For BMs, the Graded Prognostic Assessment (GPA) index is the most popular clinically-validated prognostic scale1,3. However, it does not use any imaging information, but only clinical variables. In this sense, the field of Radiomics has the potential to improve the prognostic and predictive value of GPA and set the ground for novel indexes17,18. Radiomic-based research in brain tumors has been huge, and a variety of parameters have been studied4,7,16,19,20,21,22. Additionally, while morphological features obtained from MRI have proven effective in the setting of other brain tumors, little research has been done on their utility for BMs.23,24,25,26,27,28,29. The calculation of those biomarkers relies on brain tumor segmentations. Several approaches constructed using ML and DL algorithms have been proposed in the literature to automate this procedure11,12,30,31,32,33,34. However, due to the lack of large BM public datasets, there is no common ground on which they can be properly compared.
Publicly available datasets of BMs are limited. The most popular repository of images for cancer research is The Cancer Imaging Archive (TCIA)35, including more than 140 imaging repositories of different human cancers. However, in the case of BMs, only one database including 156 whole brain MRI studies have been found available14. This leads to the fact that while there is a good amount of public data for the much less frequent primary brain tumors such as glioblastoma, available datasets for BMs are scarce.
This study tries to solve that problem by contributing longitudinal magnetic resonance imaging studies of 75 BM patients, harboring 260 BM lesions, for a total of 637 imaging studies. Imaging studies include pretreatment post-contrast T1-w sequences, and most of them include other sequences such as T1, T2, FLAIR, DWI, etc. Semi-automatic segmentations of 154 different BMs for a total of 593 post-contrast T1-W segmentations are also provided with the dataset. These data are accompanied by an extensive database including clinical data and a set of morphological and radiomic-based features obtained from the segmentations.
MRI studies in our dataset have four times the number of segmentations than those currently publicly available14. Additionally, we make public three excel files, one of which contains clinical data, including patient information, details about the primary tumor, details about treatments, and the date of the patient’s death, as opposed to the already published one, which only contains information about the histology of the primary tumor.
Methods
Subject characteristics
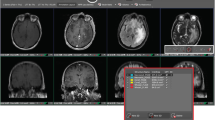
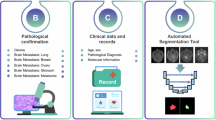
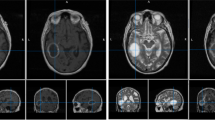
Data collected include the follow-up imaging studies and clinical data of 75 BM patients from 5 different medical institutions. Inclusion criteria was defined as: deceased adult patients with pathologically confirmed diagnosis of BM between January 1, 2005 and December 31, 2021, availability of imaging studies with at least the post-contrast T1-w high-resolution sequence (pixel spacing ≤2 mm., slice thickness ≤2 mm., no gap between slices), no noise or artifacts in the images, and availability of basic clinical data (age at diagnosis, sex, treatment schemes followed, survival, etc.). Primary tumors were: Non-small cell lung cancer (NSCLC) (n = 38), small cell lung cancer (SCLC) (n = 5), breast cancer (n = 22), melanoma (n = 6), ovarian cancer (n = 2), kidney cancer (n = 1) and uterine cancer (n = 1).
The 75 patients included had a total of 260 BMs with a total of 637 imaging studies. Of those, 593 studies were semi-automatically segmented as described below.
Image acquisition
All post-contrast T1-W sequences were obtained after intravenous administration of a single dose of contrast. The 593 imaging sequences segmented were acquired with a 1-T (n = 8), 1.5-T (n = 550) or 3.0-T (n = 35) MR imaging scanners. Regarding the MR imaging vendors, General Electric (n = 225), Philips (n = 197), and Siemens (n = 171) medical systems were used. Other image parameters are described in Table 1.
Segmentation procedure
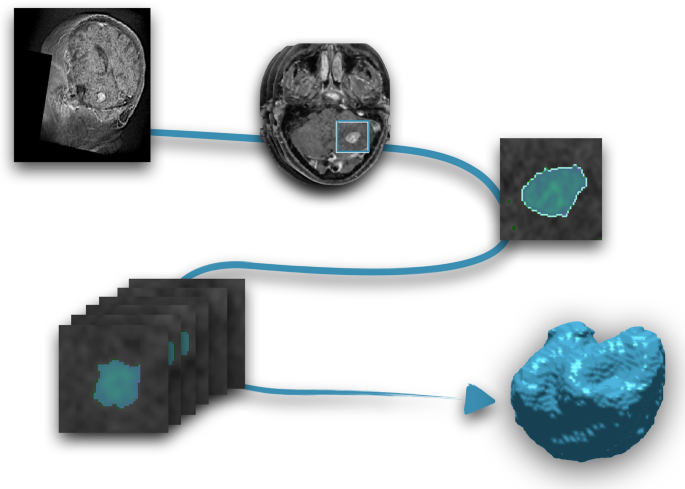
Segmentation was performed using an in-house semi-automatic segmentation procedure26,28. Tumors were automatically delineated by using a gray-level threshold chosen to identify the largest contrast-enhancing tumoral volume. Then, a biomedical engineer/applied mathematician (B. O.-T.) carefully corrected each segmentation, slice by slice, using a brushing/pixel-removing tool. The segmentation process is summarized in Fig. 1. The outcome was cross-checked by three researchers with more than seven years of expertise on MRI (D. M.-G., J. P.-B., V. M. P.-G.) and then corrected by one of the radiologists participating in the study (B.A, A.O.M, D.A, L.A.P.-R., E.A.). The raw medical images in DICOM format were used in this procedure, so they were not modified to perform the tumor segmentations.
Clinical data and anonymization
Clinical data were collected for the 75 patients. For each patient, age at diagnosis and sex, primary tumor type and subtype, molecular markers (e.g. EGFR, ALK and ROS1 for lung cancer) and tumor stage were taken. Also, the GPA index1,3, was included for a subset of institutions. Regarding each BM, the ID (a number to differentiate it from other BMs in the same patient), location in the brain (frontal, temporal, parietal and occipital, right and left side), date of appearance on MRI, and treatments received were recorded. For each treatment, the type of treatment, doses, fractions, date of start and date of end were recorded. The dates of follow-up MRI studies available were also included. Radionecrosis was confirmed for 39 lesions.
The first step of the data anonymization was performed at the institutions of origin of the data. Such a step included patient and center data anonymization. An additional more profound anonymization was performed using the clinical trials processor from the medical imaging resource center36. Within that step, all private DICOM tags and all tags containing sensitive or identifying information as well as all dates were modified such that for every subject, the imaging study where the first BM was initially identified corresponds to January 1st, 1900. The anonymized times were computed taking as reference that time point, in days, which means that negative numbers identified treatments prior to the diagnosis of the BM. The relative differences in times for the different events for each patient were preserved. The last anonymization step was a defacing process that made impossible the facial reconstruction. After this whole process, patient records were finally reviewed independently by three of the authors (B. O.-T., J. P.-B., and J. A. R.-R.).
Morphological parameters
Different morphological parameters were computed from the segmentations and gathered in the database, including the following:
Volumes
For each focus, three different types of volumes were computed: the contrast-enhancing (VCE), necrotic (or non-enhancing) (VN) and total volume (V = VCE + VN).
Contrast-enhancing spherical rim width (CE rim width)
Obtained for each focus from the CE and necrotic volumes as
By assuming that the areas of necrotic tissue and the entire tumor are spherical, this feature calculates the average width of the CE areas. Additional information and illustrations of tumors with high and low CE rim widths, can be found in29.
Surface
Obtained by reconstructing the tumor surface using the Matlab “isosurface” command from the discrete sets of voxels characterizing the tumor.
Surface regularity
It is a dimensionless ratio between the volume of the segmented tumor divided by the volume of a spherical tumor with the same surface. For each focus, it was calculated as
The range for this parameter is 0 (for tumors with highly uneven surfaces) and 1 (for spherical tumors). Additional information and illustrations of tumors with high and low CE rim widths, can be found in17.
Maximum diameter
It provides the largest longitudinal measure of the tumor and is computed for each focus as the maximum distance between two points located on the surface of the CE tumor.
Radiomic-based features
A total of 110 different features were extracted with the open-source Python package PyRadiomics version 2.2.037. This feature dataset includes 16 shape descriptors and different measures of the intensity distribution and texture within the segmentation labels. The intensity features include simple first-order statistics (19 features), those derived from the gray-level co-occurrence matrix (GLCM, 24 features), gray-level run-length matrix (GLRLM, 16 features), gray-level size-zone matrix (GLSZM, 16 features), neighboring gray-tone difference matrix (NGTDM, 5 features), and gray-level dependence matrix (14 features). The features were extracted from the original image sequence after z-score normalization, intensity scaling by a factor of 100 and subsequently shifting by 300 (i. e. three standard deviations) to ensure most intensity values are positive for the first-order features and geometry tolerance 0.04. Other specific tasks may require different feature extraction procedures18.
No voxel resampling prior to feature extraction was used to maintain the information as unaltered as possible. Since the algorithm to extract image features is shared, any user can redo the extraction by applying any resampling.
Atlas location features
Affine registration was used to align all subjects to MNI atlas space38 using the mri_robust_register39. The centroid of each separate metastasis lesion was listed and may be used to efficiently identify the location and affected brain region.
Ethical approval
We have complied with all relevant ethical regulation and all subjects included in the study are deceased. Human data were obtained in the framework of the study OpenBTAI (Open database of Brain Tumors for studies in Artificial Intelligence), a retrospective, multicenter, nonrandomized study approved by the corresponding institutional review boards: Fundación Instituto Valenciano de Oncología (2021-05), Hospital Universitario HM Sanchinarro (21.06.1858-GHM), Hospital Universitario 12 de Octubre (21/711), Hospital General Universitario de Ciudad Real (12/2021), Hospital Regional Universitario de Málaga (24/06/2021), Hospital Universitario y Politécnico La Fe (2021-504-1), MD Anderson Cancer Center (01/06/2021), Hospital Universitario de Salamanca (2021 10 879), Complejo Hospitalario Universitario de Toledo (29/9/2021-770) and Hospital Universitario Marqués de Valdecilla (14/2021 – 10/09/2021).
Data Records
All data records collected for this manuscript are available at the Figshare Repository40 and on the webpage https://molab.es where the number of cases will be expanded.
Raw medical images for each follow-up study have been stored using the Digital Imaging and Communications in Medicine image file format (DICOM, ISO 12052). Tumor segmentations and the corresponding images have been stored in The Neuroimaging Informatics Technology Initiative (NIfTI) format, maintaining raw medical image coordinates, since no preprocessing was used to perform the manual segmentations. We have uploaded six zip files with the DICOMS images, one containing all the segmentations (files ended _msk.nii) and one containing the corresponding images (files ended _img.nii) to each of the segmentations available. Also, three excel files containing: (1) all the clinical data, (2) morphological parameters measured directly from the segmentations, and (3) radiomic-based features computed for each follow-up study segmented are included together with the imaging data.
Technical Validation
Data collection
The collaborating expert board-certified neuroradiologists identified and collected the 637 follow-up studies of the 75 BM patients included in the study. Only confirmed BM patients were included in the study, and primary tumors for each patient were pathologically confirmed and verified prior to inclusion in the study.
Data curation and testing of the inclusion criteria was performed by a biomedical engineer/applied mathematician with more than seven years experience in management of medical images (B. O.-T., D. M.-G., J. P.-B. and V. M. P.-G.) and then cross-checked by a different expert.
Segmentation method
All semi-automatic segmentations performed in this study were carefully validated by an expert radiologist after have been performed by experienced experts in the management of medical images and cross-checked by a different expert. A reproducibility study for the methodology was performed in26, showing its reliability.
Each segmentation mask contains two labels for each BM: labels ending in 1 correspond to contrast-enhancing (CE) parts of the tumor; labels ending in 2 represent the non-enhancing or necrotic area of the tumor. Features were extracted for CE and necrotic zones and also were computed for the combination of both.
Comparison between measurements obtained and radiomic features
Two excel files are provided with features from the segmented images. One of them contains some morphological variables computed directly from the manual segmentation while the other is a radiomic-based set of features.
Usage Notes
The whole dataset can be downloaded from the figshare repository40. To process the provided images and segmentations, it is highly recommended that medical imaging tools be used, which handle consistently the physical space and orientation of the images. We verified that all the Nifti files (segmentations and images) can be loaded correctly with FSLeyes v1.3.0 (https://www.fsl.fmrib.ox.ac.uk) (FMRIB Centre, Oxford, UK) and DICOM files could be easily loaded using Horos v3.3.6 (https://www.horosproject.org).
Code availability
We provide the code used to extract the features with PyRadiomics at https://github.com/ysuter/OpenBTAI-radiomics. For reproducibility and convenience in case any user wants to customize the extraction, all the.py files needed and a “readme” file are available.
References
Achrol, A. S. et al. Brain metastases. Nature Rev Dis Primers. 5(5), 1–26 (2019).
Nayak, L., Lee, E. Q. & Wen, P. Y. Epidemiology of brain metastases. Curr Oncol Rep. 14, 48–54 (2012).
Lignelli, A. & Khandji, A. G. Review of imaging techniques in the diagnosis and management of brain metastases. Neurosurg Clin N Am. 22, 15–25 (2011).
Kniep, H. C. et al. Radiomics of brain MRI: utility in prediction of metastatic tumor type. Radiology. 290, 479–487 (2019).
Dikici, E. et al. Automated brain metastases detection framework for T1-weighted contrast-enhanced 3D MRI. IEEE J Biomed Health Inform. 24(10), 2883–2893 (2020).
Cho, S. J. et al. Brain metastasis detection using machine learning: a systematic review and meta-analysis. Neuro Oncol. 23(2), 214–225 (2021).
Tong, E., McCullagh, K. L. & lv, M. Advanced imaging of brain metastases: from augmenting visualization and improving diagnosis to evaluating treatment response. Front Neurol. 11, 1–14 (2020).
Wolpert, F. et al. Risk factors for the development of epilepsy in patients with brain metastases. Neuro Oncol. 22(55), 718–728 (2020).
Kim, H. Y. et al. Classification of true progression after radiotherapy of brain metastasis on MRI using artificial intelligence: a systematic review and meta-analysis. Neuro Oncol Adv. 3(1), 1–12 (2021).
Gagliardi, F. et al. Role of stereotactic radiosurgery for the treatment of brain metastasis in the era of immunotherapy: A systematic review on current evidences and predicting factors. Critical Reviews in Oncology Hematology. 165, 103431 (2021).
Bousabarah, K. et al. Deep convolutional neural networks for automated segmentation of brain metastases trained on clinical data. Radiat Oncol. 15, 87 (2020).
Xue, J. et al. Deep learning-based detection and segmentation-assisted management of brain metastases. Neuro Oncol. 22(4), 505–514 (2020).
Charron, O. et al. Automatic detection and segmentation of brain metastases on multimodal MR images with a deep convolutional neural network. Comput Biol Med. 95, 43–54 (2018).
Grøvik, E. et al. Deep learning enables automatic detection and segmentation of brain metastases on multi-sequence MRI. J Magnet Reson Imag. 51(1), 175–182 (2019).
Liu, Y. et al. A deep convolutional neural network-based automatic delineation strategy for multiple brain metastases stereotactic radiosurgery. Plos One. 12(10), e0185844 (2017).
Bae, S. et al. Robust performance of deep learning for distinguishing glioblastoma from single brain metastasis using radiomic features: model development and validation. Sci Rep. 10(1), 12110 (2020).
Gillies, R. J., Kinahan, P. E. & Hricak, H. Radiomics: Images Are More than Pictures, They Are Data. Radiology. 278(2), 563–577 (2016).
Mouraviev, A. et al. Use of radiomics for the prediction of local control of brain metastases after stereotactic radiosurgery. Neuro-Oncology. 22(6), 797–805 (2020).
Molina, D. et al. Tumour heterogeneity in glioblastoma assessed by MRI texture analysis: a potential marker of survival. Br J Radiol. 89(1064), 20160242 (2016).
Suter, Y. et al. Radiomics for glioblastoma survival analysis in pre-operative MRI: exploring feature robustness, class boundaries, and machine learning techniques. Cancer Imaging 20, 55 (2020).
Baid, U. et al. Overall survival prediction in glioblastoma with radiomic features using machine learning. Front Comput Neurosci. 14, 61 (2020).
Narang, S., Lehrer, M., Yang, D., Lee, J. & Rao, A. Radiomics in glioblastoma: current status, challenges and opportunities. Trasl Cancer Res. 5(4), 383–397 (2016).
Pérez-Beteta, J. et al. Glioblastoma: does the pre-treatment geometry matter? A postcontrast T1 MRI-based study. Eur Radiol. 27(3), 1096–1104 (2017).
Wangaryattawanich, P. et al. Multicenter imaging outcomes study of The Cancer Genome Atlas glioblastoma patient cohort: imaging predictors of overall and progression-free survival. Neuro Oncol. 17(11), 1525–1537 (2015).
Grabowski, M. M. et al. Residual tumor volume versus extent of resection: predictors of survival after surgery for glioblastoma. J Neurosurg. 121(5), 1115–1123 (2014).
Pérez-Beteta, J. et al. Tumor surface regularity at MR imaging predicts survival and response to surgery in patients with glioblastoma. Radiology. 288(1), 218–225 (2018).
Ellingson, B. M., Bendszus, M., Sorensen, A. G. & Pope, W. B. Emerging techniques and technologies in brain tumor imaging. Neuro Oncol. 16(7), 12–23 (2014).
Pérez-Beteta, J. et al. Morphological MRI-based features provide pretreatment survival prediction in glioblastoma. Eur Radiol. 29(4), 1968–1977 (2019).
Cui, Y. et al. Prognostic imaging biomarkers in glioblastoma: Development and independent validation on the basis of multiregion and quantitative analysis of MR images. Radiology. 278(2), 546–553 (2016).
Menze, B. H. et al. The multimodal brain tumor image segmentation benchmark (BRATS). IEEE T Med Imaging. 34(10), 1993–2024 (2015).
Ermiş, E. et al. Fully automated brain resection cavity delineation for radiation target volume definition in glioblastoma patients using deep learning. Radiat Oncol. 15, 100 (2020).
Porz, N. et al. Multi-modal glioblastoma segmentation: man versus machine. Plos one. 9(5), e96873 (2014).
Kamnitsas, K. et al. Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Med Image Anal. 36, 61–78 (2017).
Meier, R. et al. Clinical evaluation of a fully-automatic segmentation method for longitudinal brain tumor volumetry. Sci Rep. 6, 23376 (2016).
Clark, K. et al. The Cancer Imaging Archive (TCIA): Maintaining and operating a public information repository. Journal of Digital Imaging. 26 (6), 1045–1057 (2013).
Aryanto, K. Y. E., Oudkerk, M. & van Ooijen, P. M. A. Free DICOM de-identification tools in clinical research: functioning and safety of patient privacy. European radiology 25(12), 3685–3695 (2015).
van Griethuysen, J. J. et al. Computational radiomics system to decode the radiographic phenotype. Cancer Res. 77(21), e104–e107 (2017).
Mazziotta, J. et al. A probabilistic atlas and reference system for the human brain: international consortium for brain mapping (ICBM). Philos. Trans. Roy. Soc. Lond. Ser. B: Biol. Sci. 356(1412), 1293–1322 (2001).
Reuter, M., Rosas, H. D. & Fischl, B. Highly accurate inverse consistent registration: a robust approach. NeuroImage 53(4), 1181–1196 (2010).
Ocaña-Tienda, B. et al. Brain Metastasis MR images with segmentations, clinical data, morphological measurements and radiomic features, Figshare, https://doi.org/10.6084/m9.figshare.c.6194104.v1 (2023).
Acknowledgements
This research has been supported by grants awarded to V.M. P.-G. by the James S. Mc. Donnell Foundation, United States of America, 21st Century Science Initiative in Mathematical and Complex Systems Approaches for Brain Cancer (collaborative award 220020560, https://doi.org/10.37717/220020560), Ministerio de Ciencia e Innovación, Spain (grant numbers PID2019-110895RB-I00 and PDC2022-133520-I00) and Junta de Comunidades de Castilla-La Mancha (SBPLY/21/180501/000145). BOT is supported by the Spanish Ministerio de Ciencia e Innovación (grant PRE2020-092178).
Author information
Authors and Affiliations
Contributions
B.O.-T., J.P.-B., D.M.-G., M. R., E.A. and V. M.P.-G. designed research; B.O.-T. performed the segmentations; Y.S. performed full data anonymization; B.A., D.A., A.O.M., L.A.P.-R., E.G.P., M.L., N.C., F.N.-R., M.V.-D., B.L. and E.A. collected data; B.O.-T., D.M.-G., J.P.-B., J.A.R.-R. and V.M.P.-G. analyzed data; D.M.-G. and V.M.P.-G. wrote the paper; All authors revised and corrected the manuscript. B.O.-T. and J.P.-B. contributed equally to the paper and V.M.P.-G and E. A. are both joint senior authors of this manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ocaña-Tienda, B., Pérez-Beteta, J., Villanueva-García, J.D. et al. A comprehensive dataset of annotated brain metastasis MR images with clinical and radiomic data. Sci Data 10, 208 (2023). https://doi.org/10.1038/s41597-023-02123-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02123-0
This article is cited by
-
Morphological MRI features as prognostic indicators in brain metastases
Cancer Imaging (2024)
-
A large open access dataset of brain metastasis 3D segmentations on MRI with clinical and imaging information
Scientific Data (2024)
-
A Multi-Center, Multi-Parametric MRI Dataset of Primary and Secondary Brain Tumors
Scientific Data (2024)
-
Leveraging radiomics and machine learning to differentiate radiation necrosis from recurrence in patients with brain metastases
Journal of Neuro-Oncology (2024)
-
Investigation of distributed learning for automated lesion detection in head MR images
Radiological Physics and Technology (2024)