Abstract
The biogeography of bacterial communities is a key topic in Microbial Ecology. Regarding continental water, most studies are carried out in the northern hemisphere, leaving a gap on microorganism’s diversity patterns on a global scale. South America harbours approximately one third of the world’s total freshwater resources, and is one of these understudied regions. To fill this gap, we compiled 16S rRNA amplicon sequencing data of microbial communities across South America continental water ecosystems, presenting the first database µSudAqua[db]. The database contains over 866 georeferenced samples from 9 different ecoregions with contextual environmental information. For its integration and validation we constructed a curated database (µSudAqua[db.sp]) using samples sequenced by Illumina MiSeq platform with commonly used prokaryote universal primers. This comprised ~60% of the total georeferenced samples of the µSudAqua[db]. This compilation was carried out in the scope of the µSudAqua collaborative network and represents one of the most complete databases of continental water microbial communities from South America.
Measurement(s) | taxonomic diversity assessment by targeted gene survey |
Technology Type(s) | next generation DNA sequencing |
Sample Characteristic - Organism | Bacteria • Archaea |
Sample Characteristic - Environment | aquatic environment |
Sample Characteristic - Location | South America |
Similar content being viewed by others
Background & Summary
Microorganisms are the main drivers of biogeochemical cycles in freshwater ecosystems1,2,3,4. Due to their high abundances and activities and to their collective metabolic and phylogenetic diversity, prokaryotes support aquatic food webs and regulate the magnitude and recycling rates of major elements5. Thus, understanding the microbial diversity patterns is a fundamental topic in modern Microbial Ecology and a key step for advancing our knowledge on bacterial-mediated processes across continental water ecosystems.
Despite the extensive application of amplicon sequencing by high-throughput technologies (HTS), there are still important gaps in the study of aquatic microbial diversity6,7,8. For example, a rough mapping of the worldwide distribution of amplicon sequencing studies (Fig. 1), clearly shows that most of them are from the northern hemisphere, particularly from Europe and the United States – Canada, while the Southern Hemisphere has a contrasting underrepresentation9,10. This is especially true in South America and Africa, where sequencing studies are still scarce and generated from isolated efforts.
Global distribution of amplicon sequencing samples from continental water systems using HTS. The information was acquired from MGnify (https://www.ebi.ac.uk/metagenomics/) resource by searching for non-marine aquatic samples, obtained by amplicon or metabarcoding experimental types. The geographical coordinates were retrieved for 4,691 samples from a total of 7,832 using the metadata available from each sample.
The Southern Hemisphere covers a comparatively high share of the surface and volume of the continental and marine ecosystems in the world. In particular, South America is considered the “continent of water”, harboring 6 out of the 10 largest rivers in the world in terms of water discharge, draining about 30% of the continental freshwater that reaches the ocean11. This water flows through five huge hydrological river basins: the Amazonas (6,000,000 km2), Del Plata-Paraná/Paraguay (2,600,000 km2), Orinoco (990,000 km2), Araguaia-Tocantins (757,000 km2), and São Francisco (634,000 km2)12,13. In addition, a great number and diversity of lentic water bodies are also prominent features that tend to occur in lake districts and wetlands as a result of the main climatic and geomorphological processes acting on regional scales14.
Furthermore, the South American continent comprises a large ecological heterogeneity15,16. South America covers about 15% of the global land area (17,870,218 km2) and spans a broad latitudinal range, extending from 12° 28′N (Punta Gallinas, Colombia) to 55° 59′S (Cabo de Hornos, Chile). According to the biogeographic regionalization by Cabrera & Willink15, South America belongs to the Neotropical region, except the southernmost area, which is assigned to the Antarctic region. Owing to the wide latitudinal coverage, a large variety of climates occur, with much of the continental mass located within the intertropical belt, large regions of Chile, Argentina, and Uruguay laying in the Southern Temperate Zone, and the southern tip of the continent extending into sub-Antarctic latitudes. Due to the high habitat heterogeneity and a complex geological history, South America is considered a hotspot of biodiversity, being the most species-rich region on Earth17,18. This fact, added to its location in a predominantly maritime hemisphere, offers a unique opportunity for comparing empirical patterns found originally for Northern Hemisphere aquatic ecosystems. Pioneering studies have shown a geographically biased picture of the aquatic microbiome, leading to overlook differences in structure and functioning of microbial communities19. That limits, for instance, our understanding of the expected scenarios resulting from the current pressures experienced by the aquatic ecosystems20.
In order to reduce this gap, we performed an exhaustive bibliographic search, collected and annotated data of bacterial communities from South American continental waters. We constructed the first 16S rRNA amplicon sequencing database of South America, the µSudAqua[db] containing 866 georeferenced samples. For its integration in further works, preventing biases by the sets of primers and sequencing methodology used, we constructed a curated database (µSudAqua[db.sp]) which contains over 509 samples of the V3-V4 region of the 16S rRNA gene sequenced by the Illumina MiSeq technology with the commonly used primers proposed by Herlemann & collaborators21.
This work is a result of the µSudAqua collaborative network, a Latin American network in Aquatic Microbial Ecology. This network emerged as an initiative to nucleate researchers of the field to join efforts in consolidating a regional critical mass. Its main objectives are to strengthen and expand the interactions between aquatic microbial ecologists, to contribute to the development of a community feeling at the regional level, and to provide a fruitful space for long-term collaboration in research and training of human resources. More information can be found at the µSudAqua website (https://microsudaqua.netlify.app/).
Methods
Data compilation
The µSudAqua[db] database was constructed with samples from published papers and new data generated in the scope of this work (Table 1). We only considered those studies fitting with the following criteria: 1) samples were obtained from continental water systems of South America; 2) the whole bacterial community was studied using high-throughput amplicon sequencing of the 16S rRNA gene; 3) the 16S rRNA gene was subject to amplification using universal primers (i.e. studies using group-specific primers or functional genes were not included); 4) sequencing data were publicly available or provided by the authors of the study upon request; 5) the samples could be georeferenced.
Sample metadata were collected from the published papers or provided by the authors of the current work. The altitude was automatically extracted based on the sampling location, using the QGIS geographic information system software (https://qgis.org/). Each sample was assigned to an environmental type (e.g. shallow and deep lakes, rivers, streams, reservoir, swamps) and an ecoregion (section Ecoregions description). Besides, the georeferenced location and procedures adopted for the sampling and sequencing were fully recovered. The complete list of metadata recovered and its description is presented in Table 2. The samples information used to build the database is available as an accessed as plaintext (TSV format) at Zenodo repository22.
The µSudAqua[db] was used as a seed to construct the curated database, µSudAqua[db.sp], which contains a subset those samples sequenced with 1) Illumina MiSeq technology and; 2) the commonly used set of primers proposed by Herlemann & collaborators21.
The microbial communities were obtained with different filtration strategies. In some environments, water samples were pre-filtered to exclude larger particles, or to split the microbial community in free-living and particle attached fractions (Table 3). Even though different DNA-extraction methods were used (Table 3), the V3-V4 regions of the 16S rRNA gene were amplified using the same set of bacterial universal primers 341 F (5′-CCTACGGGNGGCWGCAG-3′) and 805 R (5′-GACTACHVGGGTATCTAATCC-3′)21. Samples of each project were indexed with Nextera XT v2 kit, and sequenced using the Illumina MiSeq technology in different sequencing facilities. The samples were obtained mostly from surface waters (0–50 cm) of continental systems with different limnological characteristics and different spatial and temporal coverage through six ecoregions (Table 3).
Amplicon sequences from the µSudAqua[db.sp] were processed using DADA2 v1.10.023, after primers trimming by Cutadapt v1.1824. Each sequencing project was analyzed separately with the same filtering parameters as recommended by Callahan & collaborators23. The quality of the samples was explored using the functions fastx_eestat and fastx_info from USEARCH v10.0.24025 to define the filtering parameters. This was then performed using the filterAndTrim function from DADA2 with the following quality values: maxEE = c(2,2) and truncLen = c(250,220). Only samples with more than 10,000 reads were analyzed.
To increase sensitivity to rare variants and avoid chimeras and sequencing errors, we used the “pool” option from dada function. The chimera sequences were excluded after merging the different projects using the functions removeBimeraDenovo and mergeSequenceTables, respectively. The taxonomic classification was performed using BLAST v2.526 with the blastn algorithm (e-value = 0.0001) and the SILVA database (SSU Ref 132 NR 9927) as a reference. The Amplicon sequence variants (ASVs) were classified into 7 different taxonomic groups. The contribution of each group was calculated as their relative abundance to the total number of reads, and the richness was defined as the total number of ASVs. The scripts used for DADA2 and sample description are available in GitHub (https://github.com/microsudaqua/usudaquadb).
Ecoregions description
To define the ecoregions, we adopted the level II classification proposed by Griffith & collaborators28 for Central, South America and the Caribbean. The characteristics of each ecoregion and subregion are briefly described below.
18. Central Andes
18.1 Central High Andes, Chile
The Central High Andes ecoregion extends from southern Peru, through Chile and Bolivia, to northern Argentina (5.18°–38.44° S, 78.17°–70.24° W). The landscape is typically mountainous, with snow-capped peaks, plateaus and valleys29. The ecoregion occupies an area of 140,960 km2 and lies within the altitudinal range between 3,200 and 6,600 m15. Its climate varies from temperate to cold, with an annual average temperature between below zero and 15 °C. This region is dry, with precipitation between 250 and 500 mm per year29,30. It is considered as a transitional zone between the wet puna to the north and west, and the dry puna to the south. This ecoregion has several high-elevation wetlands comprising both fresh and saline lakes, salt flats, temporary endorheic basins, as well as permanent rivers and streams fed by snowmelt. They regulate water flow by retaining water during the wet season and releasing it during the dry season. The salt flats, or salares, represent remnants of extensive paleolakes29.
19. Southern Andes
19.2 Valdivian Forest Hill and Mountains
The Valdivian Temperate Forests ecoregion is in the southern cone of South America (33.02°–46.91° S, 70.55°–74.51° W). It covers a narrow continental strip between the western slope of the Andes and the Pacific Ocean (area: 248,100 km2). The climate is temperate cool (mean annual temperature is 8.7 °C) with predominance of westerly winds, and annual precipitation of 1,500 mm31. The ecoregion is characterized by a profuse hydrographic system including large and deep lakes (mainly glacial origin)32,33 and small and shallow lakes34,35. The main rivers fed from these Andean waters, run across the plateau steppe and outflow to the Atlantic Ocean, but there are also other rivers that cross the Andes flowing towards the Pacific Ocean. Deep lakes (Zmax > 100 m) have a warm monomictic thermal behavior36. Nevertheless, small and shallow lakes (Zmax ~12 m) are dimictic or polymictic34. These lakes have very low nutrient (ranging from ultra-oligotrophic to oligotrophic status) and dissolved organic carbon concentrations, and high transparency to different wavelengths, which would imply high exposure to ultraviolet radiation37,38,39,40.
20. Amazonian-Orinocan Lowland
20.4 Amazon and Coastal Lowlands
The Amazon river basin is the largest in the world, comprising an area over 6 million km2, extending from 5°N to 17°S, and 79°W to 46°W. Basin sources are mostly located in the northern region of Brazil, starting in the Andes mountains of Peru and end in the Atlantic Ocean in the Brazilian coast41. The climate in the basin is in general hot and humid with mean annual temperature between 24 to 28 °C42. The average annual precipitation is ~2,200 mm, ranging from ~3,000 mm in the west to ~1,700 mm over the southeast of the basin43. The Amazon basin comprise numerous large rivers, tributaries, and large extensions of floodplains with thousands of lakes and associated wetlands linked to each other44. These systems vary from permanent to periodically flooded depending on the hydrological cycle, namely the flood pulse38. This flood pulse has a profound effect on the productivity, transport of elements and biotic interactions within these ecosystems41,45.
21. Eastern Highlands
21.2 Cerrado
The Cerrado is the second largest ecoregion in South America. It comprises the Brazilian central region (2.05°–23.77° S, 45.29°–54.37° W), and covers an area over 2 million km2, 46. It is a savannah domain, characterized by a tropical climate (mean annual temperature average: 22–27 °C), with dry winters and rainy summers46. Annual precipitation typically ranges from 1,200 to 1,800 mm and soil is usually acid and nutrient-poor47. The Cerrado altitude has little variation, being maximum only in the central highlands, from where important springs come out and end up contributing to form the three largest water basins in South America (Amazon, São Francisco and Del Plata-Paraná/Paraguay)48. There are very few natural lakes in this region, and most water bodies are either dammed shallow lakes or large hydroelectric reservoirs. As reservoirs are mainly found near cities, the nutrient inputs, pH and trophic state can vary49,50.
21.4 Atlantic Forests
The Atlantic Forests region is mainly located in Brazil, spanning along the Atlantic coast, and extending inland to Argentina and Paraguay (distributed from 5.00° S to 28.00° S and 35.14° to 53.56° W51). This ecoregion is a wide tropical (mean annual temperature ~23 °C), humid biome known mainly by its long line of coastal rainforest51. The coast is humid all over the year, with an annual precipitation typically ranging from 1,800 to 3,600 mm. This ecoregion is characterized by different formations like deciduous and semi-deciduous continental forests, bogs and mangroves, and grasslands52. Landscape can be flat and lentic environments in the countryside are either human made dammed creeks used for cattle ranching and crop irrigation or large hydroelectric reservoirs. Along the Brazilian Atlantic coast, lentic ecosystems are shallow lakes dug into the mountainside, or squeezed into the narrow strip between the mountain chain and the ocean53. There are also some herbaceous/shrubby sand-dune ecosystems, called Restinga, that form perennial or temporary coastal shallow lagoons54, which encompass wide environmental gradients (e.g.: trophic state, humic substances, salinity) that greatly influence aquatic biodiversity55.
22. Gran Chaco
22.2 Humid Chaco
Lakes and rivers from the Paraná floodplain system
The Paraná River is the second largest river of South America with a mean annual discharge of ~17,000 m3 s−1 and a drainage area of 2.6 106 km2. The headwaters are fully developed in Brazil and it travels 3,800 km along a main north to south direction through tropical to temperate latitudes up to its mouth in the Río de la Plata Estuary with mean annual temperatures of ~12.5 °C56. The middle stretch of the river begins downstream from the confluence with the Paraguay River (Argentina). Climate is humid subtropical, with annual precipitation between 900 to 1,000 mm. At this stretch, the river is characterized by a well-defined main channel and a large floodplain about 20 to 40 km wide, located by its right margin. Thousands of permanent shallow lakes and temporary environments occupy the floodplain which is flooded and drained by a well-developed and relatively stable fluvial network57. The system dynamic is subject to hydro-sedimentological pulses that occur with different magnitudes and constitute the main driving factor of the limnological features and the biota38,58, particularly, the microbial communities59,60,61.
23. Pampas
23.1 Uruguayan savanna, Uruguay
The ecoregion Uruguayan savanna comprises an area of 355,605 km2 which includes the whole country of Uruguay (30°–34° S, 53°–58° W) and extends mostly towards the southern part of Brazil to a small section of the Argentina62. The climate of this region is temperate, without dry season, and with hot summer63. The mean annual temperature ranges between 16 and 20 °C. The mean annual rainfall lies between 1,100 and 1,400 mm and is highly variable between years. This ecoregion encompasses the outlet of the Río de la Plata basin where a dense fluvial network, along with a series of coastal lagoons and numerous artificial lakes can be found. Rivers and streams are characterized by small slopes and rapid filling and draining64. Coastal lagoons, formed due to marine regressions and transgressions in the Holocene, are located at the Atlantic coast65 and their size and age increase towards the East. They are characterized by large gradients in salinity, light penetration and nutrient concentrations, and their hydrological cycle strongly determines the composition and activity of the bacterial communities66,67.
23.2 Southern Flat Pampas
The Pampa ecoregion extends westward across central Argentina (30.37°–38.98° S, 57.60°–62.31° W), from the Atlantic coast to the Andean foothills32. It is an extensive plain area (398,966 km2), except for the two, almost parallel, hill systems that cross the area in a NW–SE orientation (Sierras de Tandilia and Sierras de Ventania). The climate of this region is temperate and humid, with mean annual temperatures varying from 14 to 20 °C. The precipitation is concentrated during spring and summer months, and decreases from NE to SW (from 1,000 to 400 mm)38. The ecoregion is dominated by a large number of fluvial-aeolic shallow lakes and low order rivers and streams that mostly belong to the Salado-Vallimanca basins32. Particularly, lakes are characterized by rounded contours and pan-shaped profiles. They are typically shallow, polymictic, eutrophic to hypertrophic, with highly variable water renewal time and salinity. Most of the surrounding land is devoted to agricultural practices36. This economic development directly affected shallow lakes, promoting shifts in many of them from clear regimes, characterized by the presence of submerged vegetation, to algal-dominated turbid states68.
24. Monte-Patagonian
24.2 Patagonian Tablelands
The Patagonian tablelands ecoregion (defined as “Patagonian plateau” by Quirós & Drago32, is a complex landscape of about 600,000 km2, located in Argentina (33.68°–54.52° S, 68.75°–66.35° W. It is delimited by the Colorado River to the North, the Atlantic Ocean to the East, the Andes to the West and parallel 54° to the South69. It is characterized by extreme conditions of cold and dry climate, with average maximum temperatures of 2.9 and 14.0 °C in winter and summer, respectively, and minimum temperatures can be below −19.0 °C in winter. The mean annual precipitation is ~300 mm. This ecoregion encompasses different types of water bodies, including reservoirs, permanent natural lakes and temporary ponds. Most water bodies are shallow lakes, typically ranging from mesotrophic to eutrophic. Climate conditions determine that small shallow lakes (i.e. less than 30 km2) usually remain frozen from early autumn throughout late spring, however during the ice-free period due to frequent strong winds, the water columns are continuously mixed, thus preventing the formation of stable thermoclines70,71,72.
Data Records
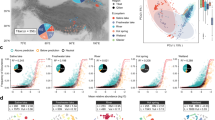
The µSudAqua[db] covers 866 individual samples of continental waters from South America (Table 1, Fig. 2). It contains samples sequenced using 454, Ion Torrent and Illumina technologies, and targeting different hypervariable regions of the 16S rRNA gene. The raw samples files are freely available in the European Nucleotide Archive (ENA) database73. They can be downloaded using the Run Accession Number from the metadata file provided in Zenodo repository22.
Sampling sites included in the µSudAqua[db] database by ecoregion. In color are highlighted the different ecoregions. The point size indicates the number of samples in the same sample site (e.g. time series). Triangles stand for samples sequenced with different primers from Herlemann & collaborators21. Those samples that constitute the µSudAqua[db.sp] database are indicated by circles.
The µSudAqua[db.sp] database is composed of 509 samples from 14 sequencing projects, representing ~60% of the data of the µSudAqua[db]. The ASVs’ information after sequencing processing using DADA2 pipeline (number of reads, nucleotide sequences and taxonomic classification) are also provided in different machine-reliable files at Zenodo22.
Technical Validation
The technical validation was performed using the µSudAqua[db.sp], that comprises the samples that were sequenced with the Illumina MiSeq technology, and targeted the V3-V4 regions of the 16S rRNA gene.
µSudAqua[db.sp]: Bacterial distribution among the ecoregions of South America
In total, 509 samples and 116,687,584 reads were processed with DADA2. In order to exclude possible remaining sequencing errors or chimeras, we filtered ASVs with less than 50 reads in less than 3 samples. Thus, the final ASV table consisted of 502 samples, 25,334 ASVs and 42,188,085 reads, from: Amazon and Coastal Lowlands (96 samples), Atlantic Forests (67), Cerrados (86), Humid Chaco (59), from Southern Flat Pampas (127), Valdivian Forested Hills and Mountains (7) and, Patagonian Tablelands (67). The information of sequence processing and quality check of samples is summarized in Table 3.
The µSudAqua[db.sp] database was mainly represented by Bacteria (24,279 ASVs, 97.7% reads) followed by chloroplasts (1,001 ASVs, 2.2% reads) and Archaea (54 ASVs, 0.03% reads). Within Bacteria, the reads were distributed in 6 principal taxonomic groups: Actinobacteria (33.4%), Proteobacteria (12.2% Betaproteobacteria, 7.6% Alphaproteobacteria and, 2.5% Gammaproteobacteria), Cyanobacteria (10.7%), Planctomycetes (9.5%), Bacteroidetes (8.6%) and Verrucomicrobia (6.7%). Bacteroidetes was the richest group (Bacteroidia, 3,359 ASVs), followed by Proteobacteria (Betaproteobacteria, 3,129 ASVs, Alphaproteobacteria, 2,133 ASVs, and Gammaproteobacteria, 1,648 ASVs) and Cyanobacteria (776 ASVs). The relative abundance and richness of each principal taxonomic group were notably different among the studied ecoregions (Fig. 3).
Usage Notes
The links to download the raw fastq data from µSudAqua[db] and µSudAqua[db.sp] are in the metadata file accessible in Zenodo22. In addition, other files associated with the µSudAqua[db.sp] are available in the same repository: ASVs table (number of reads in each sample), taxonomy, nucleotide sequences in fasta format and ASVs table filtered with only Bacteria. Importantly, the database will grow as new samples and sequencing projects from the µSudAqua network appear. This information will be uploaded in the repository and the tables will be updated in future versions of the database. A bibliography revision and open call for new data submission will be performed once a year, and the database will be updated after data quality check, processing and integration.
The µSudAqua[db] and µSudAqua[db.sp] databases are the first to integrate information of microbial diversity from continental systems of South America, an important region that has been overlooked comparing to other regions and environments worldwide. These databases will open new avenues for studies on the temporal patterns and spatial distributions of microbial communities among the different ecoregions of South America. Besides, the integration of the curated data to meta-analysis of microbial communities from different ecosystems (comparison between South America and well-studied regions of the world), will be particularly important for exploring the novel microbial diversity, allowing to reveal regions with unknown organisms and functions, as well as hotspots of microbial biodiversity.
Code availability
The workflow included several custom-made R and python scripts, which are accessible GitHub (https://github.com/microsudaqua/usudaquadb).
References
Cole, J., Findlay, S. & Pace, M. Bacterial production in fresh and saltwater ecosystems: a cross-system overview. Mar. Ecol. Prog. Ser. 43, 1–10 (1988).
Azam, F. et al. The Ecological Role of Water-Column Microbes in the Sea. Mar. Ecol. Prog. Ser. 10, 257–263 (1983).
Cotner, J. B. & Biddanda, B. A. Small players, large role: Microbial influence on biogeochemical processes in pelagic aquatic ecosystems. Ecosystems. 5, 105–121 (2002).
Falkowski, P. G., Fenchel, T. & Delong, E. F. The microbial engines that drive earth’s biogeochemical cycles. Science. 320, 1034–1039 (2008).
Newton, R. J., Jones, S. E., Eiler, A., McMahon, K. D. & Bertilsson, S. A Guide to the Natural History of Freshwater Lake Bacteria. Microbiol. Mol. Biol. Rev. 75, 14–49 (2011).
Coleman, M. L. et al. Genomic islands and the ecology and evolution of Prochlorococcus. Science. 311, 1768–1770 (2006).
Franzosa, E. A. et al. Sequencing and beyond: Integrating molecular ‘omics’ for microbial community profiling. Nat. Rev. Microbiol. 13, 360–372 (2015).
Hanson, C., Fuhrman, J., Horner-Devine, M. & Martiny, J. Beyond biogeographic patterns: processes shaping the microbial landscape. Nat. Rev. Microbiol. 10, 497–506 (2012).
Dai, A. & Trenberth, K. E. Estimates of freshwater discharge from continents: Latitudinal and seasonal variations. J. Hydrometeorol. 3, 660–687 (2002).
White, W. R. World water: resources, usage and the role of man-made reservoirs. Report No. FR/R0012. Fundation for Water Research, (2010).
Clark, E. A., Sheffield, J., van Vliet, M. T. H., Nijssen, B. & Lettenmaier, D. P. Continental runoff into the oceans (1950–2008). J. Hydrometeorol. 16, 1502–1520 (2015).
Stevaux, J. C., Paes, R. J., Franco, A. A., Mário, M. L. & Fujita, R. H. Morphodynamics in the confluence of large regulated rivers: The case of Paraná and Paranapanema Rivers. Lat. Am. J. Sedimentol. Basin Anal. 16, 101–109 (2009).
Brêda, J. P. L. F. et al. Climate change impacts on South American water balance from a continental-scale hydrological model driven by CMIP5 projections. Clim. Change 159, 503–522 (2020).
Llames, M. E. & Zagarese, H. E. Lakes and Reservoirs of South America. In Encyclopedia of Inland Waters vol.2 (ed. Linkens, G. E.). (Oxford: Elsevier, 2009).
Cabrera, A. L. & Willink, A. Biogeografia De America Latina 2da edn (Organización de los Estados Americanos, 1980).
Morrone, J. J. Biogeografía de América Latina y el Caribe 1st edn. (Nature, 2001).
Morrone, J. J. Biogeographical regionalisation of the neotropical region. Zootaxa 3782, 1–110 (2014).
Antonelli, A. et al. Amazonia is the primary source of Neotropical biodiversity. Proc. Natl. Acad. Sci. USA 115, 6034–6039 (2018).
Sarmento, H. New paradigms in tropical limnology: The importance of the microbial food web. Hydrobiologia 686, 1–14 (2012).
Meerhoff, M. et al. Environmental Warming in Shallow Lakes. A Review of Potential Changes in Community Structure as Evidenced from Space-for-Time Substitution Approaches. Adv. Ecol. Res. 46, 259–349 (2012).
Herlemann, D. P. R. et al. Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J. 5, 1571–1579 (2011).
Metz, S. & Huber, P. et al. A georeferenced rRNA amplicon database of aquatic microbiomes from South America (Dataset), Zenodo, https://doi.org/10.5281/zenodo.6802178 (2022).
Callahan, B. J., Sankaran, K., Fukuyama, J. A., McMurdie, P. J. & Holmes, S. P. Bioconductor Workflow for Microbiome Data Analysis: from raw reads to community analyses. F1000 Research 5, 1492 (2016).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10 (2011).
Edgar, R. C. UNOISE2: improved error-correction for Illumina 16S and ITS amplicon sequencing. Preprint at https://www.biorxiv.org/content/10.1101/081257v1 (2016).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Quast, C. et al. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Griffith, G. E., Omernik, J. M. & Azevedo, S. H. Ecological classification of the Western Hemisphere http://ecologicalregions.info/htm/ecoregions.htm (1998).
Salcedo, J. C. R. South America: Argentina, Bolivia, and Peru https://www.worldwildlife.org/ecoregions/nt1002 Accessed (2018).
Vidal, J. Geografía del Perú: las ocho regiones naturales, la regionalización transversal, la microregionalización 9th edn (PEISA, 1987).
Paruelo, J. M., Beltran, A., Jobbagy, E., Sala, O. E. & Golluscio, R. A. The climate of Patagonia: General patterns and controls on biotic processes. Ecol. Austral 8, 85–101 (1998).
Iriondo, M. Quaternary lakes of Argentina. Palaeogeogr. Palaeoclimatol. Palaeoecol. 70, 81–88 (1989).
Soto, D. & Campos, H. in Ecología de los bosques templados de Chile vol. 1 (eds. Khalin, J. M. & Villagrán, C.) (Editorial Universitaria, 1995).
Modenutti, B. et al. Structure and dynamic of food webs in Andean North Patagonian freshwater systems: Organic matter, light and nutrient relationships. Ecol. Austral 20, 95–114 (2010).
Modenutti, B. E. et al. Structure and dynamics of food webs in Andean lakes. Lakes Reserv. Res. Manag. 3, 179–186 (1998).
Quirós, R. & Drago, E. The environmental state of Argentinean lakes: An overview. Lakes Reserv. Res. Manag. 4, 55–64 (1999).
Morris, D. P. et al. The attenuation of solar UV radiation in lakes and the role of dissolved organic carbon. Limnol. Oceanogr. 40, 1381–1391 (1995).
Bastidas Navarro, M., Balseiro, E. & Modenutti, B. Bacterial Community Structure in Patagonian Andean Lakes Above and Below Timberline: From Community Composition to Community Function. Microb. Ecol. 68, 528–541 (2014).
Modenutti, B. et al. Environmental changes affecting light climate in oligotrophic mountain lakes: The deep chlorophyll maxima as a sensitive variable. Aquat. Sci. 75, 361–371 (2013).
Bastidas Navarro, M., Martyniuk, N., Balseiro, E. & Modenutti, B. Effect of glacial lake outburst floods on the light climate in an Andean Patagonian lake: implications for planktonic phototrophs. Hydrobiologia 816, 39–48 (2018).
Sioli, H. Hydrochemistry and Geology in the Brazilian Amazon Region. Amazoniana 1, 267–277 (1968).
Fick, S. E. & Hijmans, R. J. WorldClim 2: new 1-km spatial resolution climate surfaces for global land areas. Int. J. Climatol. 37, 4302–4315 (2017).
Salati, E. & Vose, P. B. Amazon Basin: A system in equilibrium. Science. 225, 129–138 (1984).
Melack, J. M. & Forsberg, B. R. In The Biogeochemistry of the Amazon Basin Vol. 1 (eds. MacCLain, M. E., Victoria, R. & Richey, J. E.). (Oxford Scholarship Online, 2001).
Junk, W. J., Bayley, P. B. & Sparks, R. E. The flood pulse concept in river-floodplain systems. Can. J. Fish. Aquat. Sci. 106, 110–127 (1989).
Ratter, J. A., Ribeiro, J. F. & Bridgewater, S. The Brazilian cerrado vegetation and threats to its biodiversity. Ann. Bot. 80, 223–230 (1997).
Haridasan, M. Nutritional adaptations of native plants of the cerrado biome in acid soils. Braz. J. Plant Physiol. 20, 183–195 (2008).
Vasconcelos, V., de Carvalho Júnior, O. A., de Souza Martins, É. & Couto Júnior, A. F. in World Geomorphological Landscapes. Vol. 1 (eds. Vieira, B., Salgado, A. & Santos, L.) (Springer, 2015).
Bichsel, D. et al. Water quality of rural ponds in the extensive agricultural landscape of the Cerrado (Brazil). Limnology 17, 239–246 (2016).
Cunha, D. G. F., Calijuri, M., do, C. & Lamparelli, M. C. A trophic state index for tropical/subtropical reservoirs (TSItsr). Ecol. Eng. 60, 126–134 (2013).
Morellato, L. P. C. & Haddad, C. F. B. Introduction: The Brazilian atlantic forest. Biotropica 32, 786–792 (2000).
Galindo-Leal, C. & Câmara, I. de G. The Atlantic Forest of South America: Biodiversity status, threats, and outlook 1st edn (Island Press, 2003).
Joly, C. A., Metzger, J. P. & Tabarelli, M. Experiences from the Brazilian Atlantic Forest: Ecological findings and conservation initiatives. New Phytologist 204, 459–473 (2014).
Caliman, A. et al. Temporal coherence among tropical coastal lagoons: A search for patterns and mechanisms. Brazilian J. Biol. 70, 803–814 (2010).
Junger, P. C. et al. Salinity Drives the Virioplankton Abundance but Not Production in Tropical Coastal Lagoons. Microb. Ecol. 75, 52–63 (2018).
Depetris, P. J., Probst, J. L., Pasquini, A. I. & Gaiero, D. M. The geochemical characteristics of the Paraná River suspended sediment load: An initial assessment. Hydrol. Process. 17, 1267–1277 (2003).
Orfeo, O. & Stevaux, J. Hydraulic and morphological characteristics of middle and upper reaches of the Paraná River (Argentina and Brazil). Geomorphology 44, 309–322 (2002).
Neiff, J. J. Large rivers of South America: toward the new approach. Verh. Internat. Verein. Limnol 26, 167–180 (1996).
Unrein, F. Changes in phytoplankton community along a transversal section of the Lower Paraná floodplain, Argentina. Hydrobiologia 468, 123–134 (2002).
Devercelli, M. Changes in phytoplankton morpho-functional groups induced by extreme hydroclimatic events in the Middle Paraná river (Argentina). Hydrobiologia 639, 5–19 (2010).
Huber, P. et al. Environmental heterogeneity determines the ecological processes that govern bacterial metacommunity assembly in a floodplain river system. ISME J. 14, 2951–2966 (2020).
Olson, D. M. et al. Terrestrial ecoregions of the world: A new map of life on Earth. Bioscience 51, 933–938 (2001).
Peel, M. C., Finlayson, B. L. & McMahon, T. A. Updated world map of the Köppen-Geiger climate classification. Hydrol. Earth Syst. Sci. 11, 1633–1644 (2007).
Conde, D., Arocena, R. & Recursos, R.-G. L. acuáticos superficiales de Uruguay: ambientes, algunas problemáticas y desafios para la gestión. Ambios 10, 1–7 (2003).
Martin, L. & Suguio, K. Variation of coastal dynamics during the last 7000 years recorded in beach-ridge plains associated with river mouths: example from the central Brazilian coast. Palaeogeogr. Palaeoclimatol. Palaeoecol. 99, 119–140 (1992).
Alonso, C. et al. Environmental dynamics as a structuring factor for microbial carbon utilization in a subtropical coastal lagoon. Front. Microbiol. 4, 1664–302X (2013).
Amaral, V., Graeber, D., Calliari, D. & Alonso, C. Strong linkages between DOM optical properties and main clades of aquatic bacteria. Limnol. Oceanogr. 61, 906–918 (2016).
Rennella, A. M. M., Quiro, R. & Quirós, R. The effects of hydrology on plankton biomass in shallow lakes of the Pampa Plain. Hydrobiologia 556, 181–191 (2006).
Diaz, M., Pedrozo, F. & Baccala, N. Summer classification of Southern Hemisphere temperate lakes (Patagonia, Argentina). Lakes Reserv. Res. Manag. 5, 213–229 (2000).
Izaguirre, I. et al. Influence of fish introduction and water level decrease on lakes of the arid Patagonian plateaus with importance for biodiversity conservation. Glob. Ecol. Conserv. 14, e00391 (2018).
Porcel, S., Saad, J. F., Sabio y García, C. A. & Izaguirre, I. Microbial planktonic communities in lakes from a Patagonian basaltic plateau: influence of the water level decrease. Aquat. Sci. 81, 51 (2019).
Bernal, M. C. et al. Spatial variation of picoplankton communities along a cascade reservoir system in Patagonia, Argentina. J. Limnol. 80, 84–99 (2021).
Leinonen, R. et al. The European nucleotide archive. Nucleic Acids Res. 39, 44–47 (2011).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA217932 (2013).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA302313 (2017).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA294718 (2022).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA309832 (2016).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA326475 (2016).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJEB48609 (2022).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA289691 (2015).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA414894 (2018).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA323673 (2016).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA356055 (2017).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA310230 (2017).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA390178 (2019).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA411849 (2017).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA725228 (2021).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA292014 (2015).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA310230 (2017).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA411849 (2017).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA316315 (2017).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA406945 (2017).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA515842 (2019).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA310230 (2017).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA321235 (2016).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:SAMN07998328 (2015).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:SAMN07998330 (2015).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJEB36116 (2020).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJEB29989 (2019).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA788397 (2021).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJEB48353 (2022).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJEB37379 (2020).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJEB46122 (2021).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJEB40710 (2020).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJEB40864 (2020).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJEB40854 (2020).
ENA European Nucleotide Archive https://identifiers.org/ena.embl:PRJNA268541 (2015).
Acknowledgements
We acknowledge the funding agencies that supported MicroSudaqua meetings in Uruguay (Centro Universitario Regional del Este -CURE-, International Society for Microbial Ecology -ISME-) and Argentina (National Agency for Scientific and Technological Promotion -ANPCyT-, National Council for Scientific and Technical Research - CONICET -, Municipality of Chascomús). We would like to thank all members of the MicroSudAqua collaborative network (listed by name on https://microsudaqua.netlify.app) for their suggestions on this study and collective effort to promote the research in Latin America, despite financial challenges and other political issues. Bioinformatics analyses were performed at the PIRAYU cluster (https://cimec.org.ar/c3/pirayu/index.php) via grants obtained from the Agencia Santafesina de Ciencia, Tecnología e Innovación (ASACTEI; Res N° 117/14) and we thank A. Dabin for informatic support. This work was funded by the São Paulo Research Foundation (Fundação de Amparo à Pesquisa do Estado de São Paulo, FAPESP process: 20/03716-0).
Author information
Authors and Affiliations
Contributions
S.M. and P.H. contributed equally. All authors contributed with 16SrRNA sequences and metadata to the database (directly or indirectly by acquiring funds, sampling, analyzing samples or processing pos-sequencing data), and participated in our meetings discussing the study design and results. S.M., P.H., E.M.B., M.M. and P.J. worked on the data compilation. S.M. and P.H. worked on the processing of sequencing data and validation. H.S., I.I., C.A., J.N.G., I.L.B., M.E.L. and M.D. wrote the first manuscript outline. V.A. and M.V.Q. provided the map of the ecoregion’s distribution. S.M., P.H. and H.S. wrote the final version of the manuscript. All authors reviewed, edited, and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Metz, S., Huber, P., Mateus-Barros, E. et al. A georeferenced rRNA amplicon database of aquatic microbiomes from South America. Sci Data 9, 565 (2022). https://doi.org/10.1038/s41597-022-01665-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-022-01665-z