Abstract
CaliPopGen is a database of population genetic data for native and naturalized eukaryotic species in California, USA. It summarizes the published literature (1985–2020) for 5,453 unique populations with genetic data from more than 187,394 individuals and 448 species (513 species plus subspecies) across molecular markers including allozymes, RFLPs, mtDNA, microsatellites, nDNA, and SNPs. Terrestrial habitats accounted for the majority (46.4%) of the genetic data. Taxonomic groups with the greatest representation were Magnoliophyta (20.31%), Insecta (13.4%), and Actinopterygii (12.85%). CaliPopGen also reports life-history data for most included species to enable analyses of the drivers of genetic diversity across the state. The large number of populations and wide taxonomic breadth will facilitate explorations of ecological patterns and processes across the varied geography of California. CaliPopGen covers all terrestrial and marine ecoregions of California and has a greater density of species and georeferenced populations than any previously published population genetic database. It is thus uniquely suited to inform conservation management at the regional and state levels across taxonomic groups.
Measurement(s) | genetic variation |
Technology Type(s) | DNA sequencing |
Factor Type(s) | Kingdom • Phylum • TaxonGroup • MarkerType • SampleSize • GeneTarget • NumMarkers • YearStart • YearEnd • PopName • LongitudeDD • LatitudeDD • CoordError • HabitatType • Lifespan • Fecundity • LifetimeReprodOutput • AgeSexMat • NumBreedingEvents • ReprodMode • BodyLength • AdultMass • CANativeStatus • CESAStatus • SSCStatus • ESAStatus • LifeCycle • AdultHeight • SelfCompatibility • MonoeciousDioecious • Asexual • PollinationMode • SeedDispMode • MassPerSeed • CAEndemicStatus |
Sample Characteristic - Organism | eukaryota |
Sample Characteristic - Location | California |
Similar content being viewed by others
Background & Summary
The CaliPopGen database consists of four datasets that contain estimates of population genetic diversity, differentiation, and life history traits for 448 eukaryotic species sampled across California, USA. The state has exceptionally high plant and animal biodiversity, and a correspondingly large number of endangered taxa1. It is often divided into 19 terrestrial and three marine ecoregions, reflecting its tremendous geologic and ecological diversity2,3, including the highest and lowest elevations in the contiguous U.S., extreme deserts and temperate rainforests, and mean annual precipitation ranging from 150 mm–1200 mm4. California is the most populous state in the USA, accommodating roughly 12% of the nation’s human population; the third largest state geographically, encompassing 5% of the country’s continental land area; and is a major agricultural producer. This combination of high species richness and human-mediated pressures constitute a persistent threat to the short- and long-term persistence of biodiversity, and has led to California’s inclusion as one of only two global biodiversity hotspots in the USA5,6. Perhaps unsurprisingly, California has the greatest number of documented and possibly extinct species of vascular plants7, and more than twice as many federally protected species (total of 287) as any other state8. It has also been the focus of more population genetic studies, including states with similarly high numbers of threatened/endangered species like Florida and Hawaii1. However, this wealth of genetic information has never been adequately summarized or made publicly available. The few broadly comparative analyses for California have largely been based on inferences derived from fewer than 10 species9,10,11,12, with the exceptions of one review13, and one empirical study14, both of which were restricted to marine taxa. California is a perennial leader in biodiversity management, and our compilation of genetic data for the state aligns with the administrative level at which environmental legislation and biodiversity management is implemented, increasing the likelihood that the CaliPopGen database will inform conservation actions.
In compiling CaliPopGen, we examined 4,942 published studies identified by our search criteria in the Web of Science, of which 450 met our final inclusion criteria and are included in the database. The majority of genetic samples represented in this database were collected from 1995–2015 (ranging from 1888–2019), and all studies were published between 1983–2020. CaliPopGen contains information on more than 187,394 individuals from 5,453 unique populations, of which 5,276 are spatially georeferenced. These populations include terrestrial (46.6%), marine (21.9%), freshwater (14.1%), amphibious (9.7%), and diadromous (7.7%) populations of fungi (<2% of unique species), chromists (<2%), plants (23%), and animals (73%; Fig. 1). CaliPopGen includes population level data with broader taxonomic coverage than recent, more global compilations, which have focused on freshwater and marine fishes15, mammals16, mammals and amphibians17, vertebrates18, and birds, fishes, insects and mammals19. Its focus at the regional (state) level is unique. The CaliPopGen database also includes a wider range of molecular markers (Fig. 2), populations and species than these previous population genetic compilations. Molecular markers in our database include RFLPs, AFLPs, allozymes and isozymes, microsatellites, mitochondrial, and other nuclear markers, whereas previously published datasets frequently focussed on one or a few loci (e.g.15,16,17,19) or single marker types (e.g.18,). Our inclusion of all available marker types both reflects the change in methodological approaches through time (for example, the temporal replacement of allozymes with microsatellites in the early 2000’s), and presents opportunities for quantitative comparisons among different marker types.
Expressed as a proportion of the study area, CaliPopGen contains at least an order of magnitude more species (0.83/1,000 km²), populations (9.59/1,000 km²), and individuals (284.04/1,000 km2) than the aforementioned studies and databases. This high spatial density of samples across the full ecological scope of California (Fig. 3) should facilitate future analyses of ecological trends at the population level where biological processes actually occur, and is well suited to help identify important mechanisms shaping genetic diversity, connectivity and fragmentation. CaliPopGen should also serve as a point of departure for future studies, providing a genetic baseline against which researchers can contrast and quantify future population genetic impacts resulting from changes in climate or land use. As such, CaliPopGen is an historical antecedent to ongoing genomic initiatives to study the diversity and distribution of California’s flora and fauna, including the California Conservation Genomics Project20, and other projects using landscape genomic approaches.
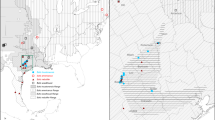
Maps of data contained in the CaliPopGen databases. (A) All unique sampling locations of both the population genetic (Dataset 121) and pairwise comparison (Dataset 221) data. The inset shows the location of California within the contiguous USA. (B) The number of unique populations in CaliPopGen per California ecoregion. Note the relative under-representation of inland desert regions (yellow) and over-representation of coastal ecoregions (purple-blue). (C) The number of unique populations of the populations genetic Dataset 121 per 20km raster cell. (D) The number of straight-line pairwise comparisons of Dataset 221 per 20km raster cell.
To supplement the genetic data in CaliPopGen, we also compiled datasets containing life history information for all plant and animal species in the database, including adult body size, lifespan, reproductive and dispersal traits, and conservation status.
Methods
Population genetic data collection from primary data sources
Figure 4 describes the overall data collection workflow for the four datasets that comprise CaliPopGen. We first identified literature potentially containing population genetic data for California by querying the Web of Science Core Collection (https://webofknowledge.com/) for relevant literature from 1900 to 2020 with the terms: topic = (California*) AND topic = (genetic* OR genomic*) AND topic = (species OR taxa* OR population*). We included only empirical peer-reviewed literature and excluded unreviewed preprints. In using these search terms, our goal was to broadly identify genetic papers focused on California with population or species-level analyses, while avoiding purely phylogenetic studies or those focused on agricultural or model species. This resulted in 4,942 unique records.
We next screened titles and abstracts to retain articles that: (1) provided data on populations of species which are self-sustaining without anthropogenic involvement; (2) included at least some eukaryote species; (3) included population(s) sampled within California; (4) mentioned measures of genetic diversity or differentiation; and (5) were not reviews (thus restricting our search to only primary literature). We retained 1869 studies after this first pass of literature screening (see Technical Validation for estimate of inter- and intra-screener bias).
Our second, more in-depth screening pass involved reading the full text of these 1869 studies. We had two goals. First, we confirmed that retained papers fully met all five of our inclusion criteria (the first screen was very liberal with respect to these criteria, and many papers failed to meet at least one criterion after close reading). Second, we eliminated papers where the data were not presented in a way that allowed us to extract population-level information. For example, many of the more systematics-focused studies pooled samples from large, somewhat ill-defined regions (“Sierra Nevada” or “Southern California”); if such regions were larger than 50 km in a linear dimension, we deemed them unusable for making geographically-informative inferences. Other studies presented summaries of population data, often in the form of phylogenetic networks or trees, but did not include information on actual population genetic parameters and therefore were not relevant to our database. We retained 528 publications after this second pass.
From this set of papers, we extracted species, locality, and genetic data for each California population or sampling locality described in each study (Fig. 3A). This included Latin binomial/trinomial, English common name, population identifiers, and geographic coordinates of sampling sites. We also noted population/sampling localities that were interpreted as comprised of interspecific hybrids, and listed both parental species. We collected population genetic diversity and differentiation statistics for each unique genetic marker for each population/sampling locality; as a result, a sampling locality may have multiple entry rows, one for each locus or marker type. Parameters extracted for each population/marker combination include sample size, genetic marker type, gene targets, number of loci, years of sampling, and reported values for effective population size (Ne), expected (HE) and observed (HO,) heterozygosity, nucleotide diversity (π, pi), alleles-per-locus (APL), allelic richness (AR), percent polymorphic loci (PPL), haplotype diversity (HDIV), inbreeding coefficient (e.g. FIS, FIT, GIS), and pairwise population genetic comparison parameters (FST, GST, DST, Nei’s D, Jost’s D, or phi). We note that while there are technical differences between allelic richness and alleles-per-locus, source literature often used the terms interchangeably, and we include the parameters and their values as named in the source. We define marker type as the general category of genetic marker used (e.g., “microsatellite” or “nuclear”), while gene targets are the specific locus/loci (e.g., “COI”). We present these data in two separate datasets, one containing all population-level genetic summary statistics (Dataset 121, see Fig. 3C and detailed description in Table 1) and a second for estimates of pairwise genetic differentiation (Dataset 221, see Fig. 3D and detailed description in Table 2).
All genetic data were extracted directly from the source literature. However, we also updated or added to the metadata for these population genetic values in several ways. We included kingdom, phylum, and a lower-level taxonomic grouping for each species (usually class), and updated scientific and common names based on the currently accepted taxonomy of the Global Biodiversity Information Facility22. When geographic coordinates were not provided for a sampling locality, as was frequently the case in the older literature, we used Google Maps (https://www.google.com/maps) to georeference localities based on either in-text descriptions or embedded figure maps guided by permanent landmarks like a bend in a river or administrative boundaries. Because this can only yield approximate coordinates, we recorded estimated accuracy as the radius of our best estimate of possible error in kilometers. If coordinates were provided in degree/minute/seconds, we used Google Maps to translate them to decimal degrees. In cases where coordinates were not provided and locality descriptions were too vague to determine coordinates with less than 50 km estimated coordinate error, we did not attempt to extract coordinates but still provide the genetic data. All coordinates are provided in the web Mercator projection (EPSG:3857). We excluded studies that reported genetic parameter values only for samples aggregated regionally (“Southern California” or “Sierra Nevada”). If marker type was not explicitly included, we classified marker type based on the gene targets reported, if provided.
Life history trait data collection
To increase the utility of CaliPopGen, we also assembled data on life history traits for all animal (Dataset 321) and plant (Dataset 421) species contained in Datasets 121 and 221. We assembled trait data that have previously been shown to correlate with genetic diversity, including those related to reproduction, life cycle, and body size, as well as conservation status (e.g.23,24,25,26,). Life history data were compiled by first referencing large online repositories, often specific to taxonomic groups, like the TRY plant trait database27, and the Royal Botanic Gardens Kew Seed Information Database28. If trait data for species of interest were unavailable from these compilations, we conducted keyword literature searches for each combination of species and life history trait, and extracted data from the primary literature. When data were not available for the subspecies or species for which we had genetic data, we report values for the next closest taxonomic level, up to and including family, as available in the literature.
For both animals and plants, we defined habitat types as marine, freshwater, diadromous, amphibious, or terrestrial. Marine species include those that are found in brackish or wetland-marine habitats, as well as bird species that primarily reside in marine habitats. Freshwater species include those that are found in wetland-freshwater habitats, as well as species that primarily reside in freshwater. The diadromous category includes fish species that are catadromous or anadromous. We considered species to be amphibious if they have an obligatory aquatic stage in their life cycle, but also spend a significant portion of their life cycle on land. Terrestrial species were defined as those that spend most of their life cycle on land and are not aquatic for any portion of their life cycle. In a few cases (e.g., waterbirds that are both freshwater and marine, semi-aquatic reptiles), a species could reasonably be placed in more than one category, and we did our best to identify the primary life history category for such taxa. If the taxonomic identity of an entry was hybrid between species or subspecies, this was noted in the speciesID column and no life history data were reported.
The CaliPopGen Animal Life History Traits Dataset 321 (description of dataset in Table 3) includes habitat type, lifespan, fecundity, lifetime reproductive success, age at sexual maturity, number of breeding events per year, mode of reproduction, adult length and mass, California native status, listing status under the US Endangered Species Act (ESA), listing status under the California Endangered Species Act (CESA), and status as a California Species of Special Concern (SSC). For some traits, value ranges were recorded–for example, minimum to maximum lifespan. In other cases, we recorded single values and, when available, a definition of this single value, (for example, minimum, average, or maximum lifespan). We report either the range of the age of sexual maturity (minimum to maximum), or a single value, depending on the available literature. For sexually dimorphic species, we report female adult length and weight when available, because female body size often correlates with fecundity. Across animal taxonomic groups, different measures of body size and length measurements are often used, reflecting community consensus on how to measure size. Given this variation, we report the type of length measurement, if available, as Standard Length (SL), Fork Length (FL), Total Length (TL), Snout-to-Vent Length (SVL), Straight-Line Carapace (SLC), or Wingspan (WS).
The CaliPopGen Plant Life History Traits Dataset 421 (description of dataset in Table 4) includes habitat type, lifespan, life cycle, adult height, self-compatibility, monoecious or dioecious, mode of reproduction, pollination and seed dispersal modes, mass per seed, California native status, NatureServe29 element ranks (global and state ranks, see Table 5 for definitions), listing status under the Federal Endangered Species Act (ESA), and listing status under the California Endangered Species Act (CESA). In contrast to most animal species, plant lifespan was typically reported as a single value. We define life cycles as the following: Annual: completes full life cycle in one year; Biennial: completes full life cycle in two years; Perennial: completes full life cycle in more than two years; Perennial-Evergreen: perennial and retains functional leaves throughout the year; Perennial-Deciduous: perennial and loses all leaves synchronously for part of the year. Some species are variable (for example, have annual and biennial individuals), and in those cases we attempted to characterize the most common modality.
Because of the paucity of data available for chromists and fungi, we did not extract life history trait data for the relatively few species in these taxonomic groups.
Data visualization and summary
We used the R-package raster (v3.1–5) to visualize the spatial extent of the data in CaliPopGen in Fig. 3. Panel (A) shows a summary plot of all unique populations of both the Population Genetic Diversity in Dataset 121 and the Pairwise Population Differentiation in Dataset 221. Panel (B) shows the total number of unique populations in each California terrestrial ecoregion. Panel (C) depicts all data entries of Population Genetic Diversity Dataset 121, summed for each 20x20 km grid cell. Panel (D) shows the density of pairwise straight lines drawn between pairs of localities in the Pairwise Population Differentiation Dataset 221, depicted as the total number of lines per 20x20 km grid cell. The number of populations and species of both Datasets 121 & 221 are summarized for each marine and terrestrial ecoregion in Table 6.
Data Records
The CaliPopGen database comprises four datasets, which are hosted at Figshare and can be downloaded as XLSX, TSV and CSV files. For convenience, the life history trait data for both animals (Dataset 321) and plants (Dataset 421) have also been included in Dataset 121 and Dataset 221. We combined the genetic and life history data under the assumption that potential users may want to examine correlations between these two classes of data. Dataset 121: The Population Genetic Diversity dataset consists of 101 columns, described in Table 1, and is comprised of data from 401 studies on 446 (sub-)species and 4,697 unique species-population-marker type combinations, with the latter equaling the number of rows in the dataset. The first 31 columns summarize taxonomic, population, marker type, and genetic data, while the remaining 70 columns contain data on animal and plant life history (Dataset 321 and Dataset 421, respectively, see below).
Dataset 221: The Pairwise Population Differentiation dataset consists of 106 columns, described in Table 2, and is comprised of data from 199 studies on 197 (sub-)species and 14,703 pairwise population comparisons, with the latter equaling the number of rows. The first 36 columns summarize taxonomic, population, marker type, and pairwise population comparison data, while the remaining 70 columns contain data on animal and plant life history (Dataset 321 and Dataset 421, respectively, see below).
Dataset 321: The Animal Life History Traits dataset consists of 45 columns, containing data for 432 species and subspecies, and is described in Table 3. The first 29 columns describe the life history of species and subspecies, and give details on their conservation status, while columns 30–45 provide information on the sources of these data.
Dataset 421: The Plant Life History Traits dataset consists of 37 columns containing data for 177 species and is described in Table 4. The first 23 columns describe the life-history of species and subspecies, and give details on their conservation status, while columns 24–37 provide sources of data. Total species numbers of Dataset 321 & 421 are higher than the number of species of Dataset 121 & 221 because we left species in these datasets even though their genetic entries may have been excluded based on the criteria set out in the Methods.
Technical Validation
Article classification
During the first step in our screening protocol based on titles and abstracts (see Fig. 4), we examined the repeatability (intra-individual variation), and reproducibility (inter-individual variation) of article classification. Given that multiple individuals were doing the article screening, we recognize that understanding variation at both of these levels is important. During this first screening, six screeners assigned a non-overlapping set of articles into three broad categories (“reject”, “include”, or “possibly include”), based on our five screening criteria (see Methods); we used “possibly include” if it was unclear from the title and abstract if a paper contained appropriate data. Each screener independently evaluated 777–782 articles (total screened = 4,942). To quantify the repeatability of our screeners, all of whom were UCLA undergraduates, each individual re-screened a subset of their original set of articles. 54 randomly selected papers were re-screened by the same person (6 screeners, range 6–13 papers per person, mean = 10.8 papers re-screened/screener). We allowed 10 weeks between the initial and re-screening procedures, which all screeners felt was a sufficiently long time that they would not remember their initial classification, and papers were randomly chosen by the senior authors. To quantify the reproducibility of the screening process across individuals, 421 papers were re-screened by a different individual than the original screener (8 re-screeners, range 46–60 per person, mean = 50.33 re-screened/screener). Each of the 421 papers was re-screened by exactly one new person. This procedure included JB and EMT in addition to the original six undergraduates.
As might be expected, intra-individual repeatability (agreement between the initial and re-screened classification of a paper screened by the same person) was higher than inter-individual reproducibility (agreement between the initial and re-screened classification of a paper screened by two different people): 92.6% (50/54) of papers re-screened by the same individual received an identical score whereas 74.8% (315/421) of papers re-screened by a different individual received an identical score. Across both of these exercises, 17.5% of articles that were re-screened by either the same or different individual (total = 475) were assigned to different categories between the first and second screening. For the inter-individual analyses, 27.7% of “possibly include” articles changed status when screened by different individuals, while only 16.0% of “reject” and 16.3% of “include” decisions changed. However, when we subsequently attempted to extract data during the Data Collection Phase, we did so from both “include” and “possibly include” papers, so the relatively low change of “reject” status makes us comfortable that screener variability and its potential bias had at most a very limited impact in our decision pipeline.
Data validation
To identify and correct potential recording errors in the datasets after the initial round of data extraction, we flagged numerical outliers and values outside of theoretical expectations for all genetic parameters and life history traits. Both outliers and values outside theoretical bounds may represent values as reported in the original publication, or they may be transcription errors as we compiled datasets. To increase the likelihood of identifying errors via outlier analysis, we examined each genetic parameter distribution separately for each marker type and taxonomic group (for example, HO of microsatellite markers in Aves was examined separately from HO of mitochondrial markers in Reptilia), and we examined life history trait distributions separately for each taxonomic group. In all cases, we identified outliers as values greater or less than the upper or lower quartiles +/− 1.5 * IQR (IQR = inter-quartile range), using the function boxplot.stats in the R-package grDevices. For all identified outliers we returned to the original source publication to confirm that values were as reported, or corrected them if they were a data-entry error. Correctly transcribed values falling outside of their theoretical bounds (HE, HO, π, FST, GST, DST are constrained between zero and 1, FIS is bounded by −1 and 1, Ne must be greater than zero) were left unaltered, which users of the CaliPopGen databases should consider carefully in using these results.
Code availability
The code used to generate figures is available at https://github.com/jbeninde/CaliPopGen. As the data was taken from the published literature manually, no additional code was used to extract data.
References
Dobson, A. P., Rodriguez, J. P., Roberts, W. M. & Wilcove, D. S. Geographic Distribution of Endangered Species in the United States. Science (New York, N.Y.) 275, 550–553, https://doi.org/10.1126/science.275.5299.550 (1997).
Griffith, G. E. et al. Ecoregions of California (poster): U.S. Geological Survey Open-File Report 2016-1021, with map, scale 1:1,100,000. Available at https://doi.org/10.3133/ofr20161021 (2016).
Spalding, M. D. et al. Marine Ecoregions of the World: A Bioregionalization of Coastal and Shelf Areas. BioScience 57, 573–583, https://doi.org/10.1641/B570707 (2007).
He, M. & Gautam, M. Variability and Trends in Precipitation, Temperature and Drought Indices in the State of California. Hydrology 3, 14, https://doi.org/10.3390/hydrology3020014 (2016).
Myers, N., Mittermeier, R. A., Mittermeier, C. G., da Fonseca, G. A. & Kent, J. Biodiversity hotspots for conservation priorities. Nature 403, 853–858, https://doi.org/10.1038/35002501 (2000).
Mittermeier, R. A., Turner, W. R., Larsen, F. W., Brooks, T. M. & Gascon, C. Global Biodiversity Conservation: The Critical Role of Hotspots. In Biodiversity Hotspots: Distribution and Protection of Conservation Priority Areas, edited by Zachos, F. E. & Habel, J. C., pp. 3–22 (Springer Berlin Heidelberg, Berlin, Heidelberg, 2011).
Kartesz, J. T. The biota of North America Program (BONAP). North American Plant Atlas. Available at http://bonap.net/napa (2015).
U.S. Fish & Wildlife Service. ECOS Environmental Conservation Online System. Listed species believed to or known to occur in each State. Available at https://ecos.fws.gov/ecp/report/species-listings-by-state-totals?statusCategory=Listed (2021).
Dawson, M. N., Louie, K. D., Barlow, M., Jacobs, D. K. & Swift, C. C. Comparative phylogeography of sympatric sister species, Clevelandia ios and Eucyclogobius newberryi (Teleostei, Gobiidae), across the California Transition Zone. Molecular ecology 11 (2002).
Rissler, L. J., Hijmans, R. J., Graham, C. H., Moritz, C. & Wake, D. B. Phylogeographic Lineages and Species Comparisons in Conservation Analyses: A Case Study of California Herpetofauna. Am. Nat. 167, 655, https://doi.org/10.2307/3844773 (2006).
Feldman, C. R. & Spicer, G. S. Comparative phylogeography of woodland reptiles in California: repeated patterns of cladogenesis and population expansion. Molecular ecology 15, 2201–2222, https://doi.org/10.1111/j.1365-294X.2006.02930.x (2006).
Thomassen, H. A. et al. Determining the drivers of population structure in a highly urbanized landscape to inform conservation planning. Conserv. Biol. 32, 148–158, https://doi.org/10.1111/cobi.12969 (2018).
Dawson, M. N. Phylogeography in coastal marine animals: a solution from California? Journal of Biogeography 28 (2001).
Kelly, R. P. & Palumbi, S. R. Genetic structure among 50 species of the northeastern Pacific rocky intertidal community. PLoS ONE 5, e8594, https://doi.org/10.1371/journal.pone.0008594 (2010).
Manel, S. et al. Global determinants of freshwater and marine fish genetic diversity. Nature communications 11, 692, https://doi.org/10.1038/s41467-020-14409-7 (2020).
Theodoridis, S. et al. Evolutionary history and past climate change shape the distribution of genetic diversity in terrestrial mammals. Nature communications 11, 2557, https://doi.org/10.1038/s41467-020-16449-5 (2020).
Miraldo, A. et al. An Anthropocene map of genetic diversity. Science (New York, N.Y.) 353, 1532–1535, https://doi.org/10.1126/science.aaf4381 (2016).
Lawrence, E. R. et al. Geo-referenced population-specific microsatellite data across American continents, the MacroPopGen Database. Sci Data 6, 14, https://doi.org/10.1038/s41597-019-0024-7 (2019).
Millette, K. L. et al. No consistent effects of humans on animal genetic diversity worldwide. Ecology letters 23, 55–67, https://doi.org/10.1111/ele.13394 (2020).
Shaffer, H. B. et al. Landscape genomics to enable conservation actions: The California Conservation Genomics Project. Journal of Heredity, esac020 (2022).
Beninde, J. CaliPopGen Dataset: Population Genetic Diversity, figshare, https://doi.org/10.6084/m9.figshare.c.5518389.v1 (2022).
GBIF Secretariat. GBIF Backbone Taxonomy, 2021.
Willoughby, J. R. et al. The reduction of genetic diversity in threatened vertebrates and new recommendations regarding IUCN conservation rankings. Biological Conservation 191, 495–503, https://doi.org/10.1016/j.biocon.2015.07.025 (2015).
Romiguier, J. et al. Comparative population genomics in animals uncovers the determinants of genetic diversity. Nature 515, 261–263, https://doi.org/10.1038/nature13685 (2014).
Robinson, J. A. et al. Genomic Flatlining in the Endangered Island Fox. Current biology: CB 26, 1183–1189, https://doi.org/10.1016/j.cub.2016.02.062 (2016).
Gallego-García, N., Caballero, S. & Shaffer, H. B. Are genomic updates of well-studied species worth the investment for conservation? A case study of the Critically Endangered Magdalena River turtle. Journal of Heredity 112, 575–589 (2021).
Kattge, J. et al. TRY plant trait database - enhanced coverage and open access. Glob. Change Biol. 26, 119–188, https://doi.org/10.1111/gcb.14904 (2020).
Royal Botanic Gardens Kew. Seed Information Database (SID). Version 7.1. Available at http://data.kew.org/sid/ (2021).
NatureServe. NatureServe Explorer [web application]. Available at https://explorer.natureserve.org/ (2021).
Acknowledgements
We thank Pauline Ordonez for her contribution to the first screening of articles. David Ackerley provided access to the unpublished Baker Seed Herbarium (BSH) Database. Many researchers helped us with their expertise: in particular, we thank Bruce Baldwin & Jason Alexander (UC Berkeley); Felipe Zapata, Lawren Sack & Nathan Kraft (UCLA); Nick Jensen (CNPS); Jeb McKay Bjerke & Kristi Lazar (CDFW).
Author information
Authors and Affiliations
Contributions
J.B., E.M.T. and H.B.S. conceptualized the study and supervised data extraction, curation, and validation, which were conducted by A.A., C.N., M.S., A.S., M.P., H.V.P., J.L.P., J.P.B., G.G. M.S. curated the animal life history dataset; M.P. curated the plant life history dataset. J.B. and E.M.T. wrote an initial draft of the manuscript, with assistance from C.N. and A.S. All authors contributed to review and editing of the manuscript, and J.B. and E.M.T. finalized the manuscript. Funding for J.B. and E.M.T. was provided by the UCLA La Kretz Center for California Conservation Science and the California Conservation Genomics Project. J.B. also received funding by the German Science Foundation (DFG: BE 6887/1-1).
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Beninde, J., Toffelmier, E.M., Andreas, A. et al. CaliPopGen: A genetic and life history database for the fauna and flora of California. Sci Data 9, 380 (2022). https://doi.org/10.1038/s41597-022-01479-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-022-01479-z
This article is cited by
-
A dataset of genetic diversity studies in the China Seas
Scientific Data (2024)
-
Best practices for genetic and genomic data archiving
Nature Ecology & Evolution (2024)
-
Citizen science helps in the study of fungal diversity in New Jersey
Scientific Data (2023)