Abstract
Circular RNA (circRNA) are a recently discovered class of RNA characterized by a covalently-bonded back-splice junction. As circRNAs are inherently more stable than other RNA species, they may be detected extracellularly in peripheral biofluids and provide novel biomarkers. While circRNA have been identified previously in peripheral biofluids, there are few datasets for circRNA junctions from healthy controls. We collected 134 plasma and 114 urine samples from 54 healthy, male college athlete volunteers, and used RNASeq to determine circRNA content. The intersection of six bioinformatic tools identified 965 high-confidence, characteristic circRNA junctions in plasma and 72 in urine. Highly-expressed circRNA junctions were validated by qRT-PCR. Longitudinal samples were collected from a subset, demonstrating circRNA expression was stable over time. Lastly, the ratio of circular to linear transcripts was higher in plasma than urine. This study provides a valuable resource for characterization of circRNA in plasma and urine from healthy volunteers, one that can be developed and reassessed as researchers probe the circRNA contents of biofluids across physiological changes and disease states.
Measurement(s) | transcriptome • RNA(circular) |
Technology Type(s) | RNA sequencing |
Factor Type(s) | biofluid |
Sample Characteristic - Organism | Homo sapiens |
Machine-accessible metadata file describing the reported data: https://doi.org/10.6084/m9.figshare.14991822
Similar content being viewed by others
Background and Summary
The advent of next-generation sequencing has spurred the discovery of a growing list of RNA biotypes, many of which are detectable across species, detected in numerous biofluids, and have biological function. While many studies have focused on microRNAs (miRNA), several other small RNA species (e.g. piwi-interacting RNAs (piRNA), tRNA fragments, and Y RNA fragments have been detected across a range of biofluids and are being developed as clinical biomarkers1,2,3,4. In addition to these linear RNAs, the discovery and detection of circular RNAs (circRNA), those with a covalently closed loop structure, have gained attention.
CircRNAs were initially discovered by electron microscopy, in the 1970s, as viroid molecules5. Nearly two decades later, circRNA were identified for a handful of mammalian genes6,7,8. Though initially thought to be rare splicing events, circRNAs have recently been identified as an abundant, endogenous RNA species in a number of organisms from Archaea to yeast, plants, worms, flies, fish, and mammals9,10,11. Additionally, circRNAs are abundantly expressed in a number of human tissues and cell types, and circRNA expression changes during development, and as a response to extrinsic factors such as stress, immune response, and hormonal stimuli12,13,14,15,16,17. These endogenous RNAs are characterized by their circular structures, which are formed by a back-splicing event that covalently links the 3′ “tail” splice donor with the upstream 5′ splice acceptor “head” of the transcript, forming a back-spliced, or “head-to-tail” junction. While circRNA function is still being elucidated, there are examples of circRNA inhibiting microRNA, regulating alternative splicing, and modulating the expression of parental genes18,19,20,21,22.
In comparison to their linear counterparts, circRNA transcripts can be more abundant and have greater stability as they are resistant to linear decay mechanisms and do not contain 5′-3′ polarity nor polyadenylated tails14,21,23,24, suggesting feasibility as stable biomarkers. CircRNA stability and detection in biofluids, saliva25, blood24,26,27,28,29,30, and urine31,32,33, comes, in part, from their being protected in extracellular vesicles28,34,35,36. Changes in circRNA expression is altered in multiple diseases, including preeclampsia, glioblastoma and colorectal cancer30,37,38. More recently, circRNAs in tumor tissues, as determined by next-generation sequencing, correlated with disease progression39,40. Urine circRNAs correlated with kidney rejection post-transplant31, while differentially expressed circRNAs have been determined in plasma exosomes of lung cancer patients versus controls41. Most studies of circRNA have small sample sizes or are based on targeted microarray data, rather than discovery-based methods. This dataset includes more than 100 samples from 54 volunteers from two easily accessible biofluids (plasma and urine). In some cases, multiple samples were collected from the same participant longitudinally, allowing us to assess the reliability of circRNA detection in biofluids.
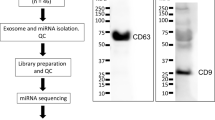
The stability and abundance of circRNAs led us to investigate detection in two easily accessed biofluids: plasma and urine. As the volunteers were part of a larger study elucidating concussion biomarkers in male, college athletes, the samples are derived from young (18–25), healthy, male volunteers as depicted in Table 1. The longitudinal sample collections of plasma and urine are depicted in Online-only Table 1, including the number of circRNAs identified in each biofluid and those circRNAs observed concurrently in the biofluids. We identified circRNA in plasma (n = 134) and urine (n = 114), using RNAseq data followed by one of six different bioinformatic tools (Fig. 1). The intersection of the 6 bioinformatic tools provides a catalog for circRNA in plasma (Fig. 2a) and urine (Fig. 2c).
CircRNAs were predicted from 134 plasma (a,b) and 114 urine (c,d) samples using 6 different bioinformatic tools. 965 circRNA were identified by all 6 tools in plasma (a; red bar), and 72 circRNA were identified by all 6 tools in urine (c; red bar). Genomic features located within predicted back-spliced junctions in plasma (b) and urine (d), respectively.
As there are few datasets with circular RNAs cataloged in clinically-relevant biofluids, we expect this data to contribute to the characterization of circRNAs in young, healthy males. While this might be a direct comparator for concussions, or other diseases more prevalent in young men, we also expect this dataset to help begin to fill out a broader assessment of circRNAs present in healthy populations.
Methods
Sample collection and participants
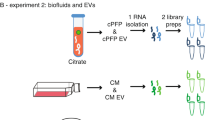
Samples were collected from healthy, male volunteers, ages 18–25, with consent and approval from the Western Institutional Review Board (WIRB) study ID #1307009395. All participants provided written consent prior to enrollment. We obtained plasma (n = 134) and urine (n = 114) samples from 54 healthy male volunteers. In 71.4% of participants, both biofluid types were collected from the same individual. Blood samples were collected in EDTA tubes, and urine was collected in sterile cups. After collection, samples were placed in a cooler with ice packs and transported from Arizona State University to the Translational Genomics Research Institute, within 2–3 hours of collection. Blood samples were spun down at 1320 x G for 10 minutes at 4 °C, and 1 mL aliquots of plasma were collected in RNase/DNase free microcentrifuge tubes (VWR) and stored at −80 C. Urine samples were spun at 1900 x G for 10 minutes at 4 °C and 15 mL aliquots were collected in 50 mL conical tubes for storage at 80 °C.
RNA isolation, library preparation, and sequencing
For plasma samples, total RNA was isolated from 1 mL plasma using the mirVana PARIS RNA and Native Protein Purification Kit (Thermo Fisher, Cat. No.: AM1556) as in Burgos et al.42, treated with the DNA-free DNA Removal Kit (Thermo Fisher, Cat. No.: AM1906), and purified and concentrated with RNA Clean & Concentrator – 5 columns (Zymo Research, Cat. No.: R1016) by following Appendix C in the kit’s protocol. For urine samples, total RNA was isolated from 15 mL urine using Norgen’s Urine Total RNA Purification Maxi Kit (Slurry Format) (Norgen, Cat. No.: 29600), treated with the RNase-Free DNase Set (Qiagen, Cat. No.: 79254), and concentrated with the speed vacuum. The isolated RNA was quantitated with Quant-iT Ribogreen RNA Assay (Thermo Fisher, Cat. No.: R11490). Samples were not ribo-depleted, double-stranded cDNA was synthesized from 10 ng total RNA with the SMARTer Universal Low Input RNA Kit for Sequencing (Clontech, Cat. No.: 634940) using thirteen PCR cycles. The double-stranded cDNA was quantitated with the Qubit dsDNA HS Assay Kit (Thermo Fisher, Cat. No.: Q32854). For each healthy control sample, Illumina-compatible libraries were synthesized from 2 ng double-stranded cDNA with Clontech’s Low Input Library Prep Kit (Clontech, Cat. No.: 634947) using four mandatory PCR cycles plus ten additional cycles. Each library was measured for size via Agilent’s High Sensitivity D1000 Screen Tape and reagents (Agilent, Cat. No.: 5067–5602 & 5067–5585) and measured for concentration via the KAPA SYBR FAST Universal qPCR Kit (Kapa Biosystems, Cat. No.: KK4824). Libraries were then combined into equimolar pools, and each pool was measured for size and concentration. Pools were clustered onto a paired-end flowcell (Illumina, Cat. No.: PE-401–3001) with a 20% v/v PhiX v3 spike-in (Illumina, Cat. No.: FC-110-3001) and sequenced on Illumina’s HiSeq. 2500 with TruSeq v3 chemistry (Illumina, Cat. No.: FC-401-3002). The first and second reads were each 83 bases.
CircRNA prediction
Samples were demultiplexed and raw fastqs generated using CASAVA (v1.8.2, Illumina). Raw fastqs were trimmed using cutadapt (v1.9) with a quality score cutoff of 30 and a minimum length of 30 bp43. For each sample, 6 different algorithms (Table 2) were used to predict circRNA: KNIFE v1.444, find_circ21, MapSplice245, CIRCexplorer46, CIRI247, and DCC48. Indices of the GRCh37/hg19 genome were created using bwa and STAR v2.4.0j using default parameters49,50; bowtie and bowtie2 genome indices were downloaded with the KNIFE package51,52. Reads were mapped to the genome with the recommended aligner and alignment parameters for each program: STAR v2.4.0j for DCC and CIRCexplorer, bowtie2 v2.2.1 for find_circ and KNIFE, bowtie v0.12.9 for MapSplice2, and bwa v0.7.13 for CIRI2. CircRNA prediction was then completed with the suggested parameters for each program, with the exception of incorporating a minimum 18nt overlap on either side of the junction. CircRNAs were kept for downstream analysis if they 1) had 2 or more junction counts and 2) were identified in at least 5 samples for each respective program.
Analysis of predicted circRNA
The version of CIRCexplorer used here does not support paired-end data; therefore, circRNA prediction was performed on each pair separately and then combined for analysis. For each program, BED files containing count expression data were created from the output data. CIRCexplorer, KNIFE, and find_circ output files all produce output files with 0-based coordinates while CIRI2, MapSplice, and DCC output files have 1-based coordinates; therefore, all coordinates were converted to a 0-based system for comparison. BED12 GRCh37 RefSeq gene annotation files were obtained from UCSC (http://genome.ucsc.edu/cgi-bin/hgTables), and bedtools v2.26.0 was used to infer genes from reported backsplice junction genome locations53. Data were analyzed using the R v3.3.2 statistical package (https://cran.r-project.org). UpSet plots were generated using the UpSetR v1.3.3 package54.
Quantification of circRNA expression
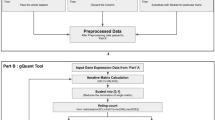
CircRNA count expression data was obtained from each respective bioinformatic program. Junction reads per million (JRPM) were calculated according to the total number of junction reads found in each sample as identified by STAR (both canonical and chimeric); therefore, JRPM = (circRNA count/junction reads) * 1,000,000. The circular-to-linear ratio (CLR) for each circRNA was calculated as described previously13,27, by counting the linear spliced reads identified by STAR on the 5′ and 3′ flanks of each circRNA junction, and dividing the back-spliced read count by the flank with the highest count; therefore, CLR = circRNA count/max (5′ linear junction count, 3′ linear junction count). In order to avoid division by zero, if no linearly spliced reads were detected, a pseudo count of 1 was added to the denominator. The number of reads assigned to the transcriptome was calculated using featureCounts (subread v1.5.1) with the Ensembl75 gene annotation55. Differential expression analysis was performed using DESeq. 2 v1.14.156, after filtering to select samples which had detected at least 300 circRNA/sample as well as exclusion of circRNA that were expressed in less than 50% of samples.
DNA isolation and qRT-PCR
After centrifugation of blood samples, DNA was isolated from the buffy coat using the DNeasy Kit (Qiagen, Cat. No.: 69504). Previously isolated RNA from samples matching those used for library prep were selected for cDNA synthesis. cDNA was synthesized with random hexamers using the SuperScript III First-Strand Synthesis System for RT-PCR following manufacturer’s protocols (Invitrogen, Cat. No.: 18080-051) with three nanograms of total RNA as input, and stored at −20 °C. Inward-facing (crossing the back-splice junction) custom primers were designed with Primer3 and LabReady primers (100 µM in IDTE pH 8.0) were ordered from Integrative DNA Technologies with Standard Desalting Purification57,58. Real-time qRT-PCR was performed with SYBR Select Master Mix (Thermo Fisher, Cat. No.: 4472919) on the QuantStudio 7 (Applied Biosystems), with 0.2 µM of primer and 0.2 µL of cDNA template or 2 ng of gDNA template per 10 µL reaction. U6 was used as a positive control and no template controls (NTCs) were used as a negative control. All results are expressed as the mean of three independent reactions, with a standard deviation less than 0.5. The ReadqPCR v1.20.0 and NormqPCR v1.20.0 Bioconductor v3.4 packages were used for qRT-PCR data analysis59.
Data Records
Raw FASTQ files for the RNAseq libraries were deposited into dbGap (accession # phs001258.v2.p1) (https://identifiers.org/dbgap:phs001258.v2.p1)60. Data (circRNAs identified across all informatic tools and raw cirRNA expression) are also provided in figshare: https://doi.org/10.6084/m9.figshare.c.542083261.
Technical Validation
CircRNA set size and genomic alignment
The set size (all circRNA in any sample by one tool) ranges from 1,835 to 7,462 and 163 to 1,349 in plasma and urine, respectively (Table 3). 965 and 72 circRNA were detected across all six tools in plasma and urine, respectively (Fig. 2a,c, red bars; Table 4; full list in figshare File 1 and 261). KNIFE predicted the most circRNA per sample in plasma and urine, while MapSplice predicted the fewest (Table 3). Table 5 displays the correlations between all of the tools, CIRCexplorer and DCC had the highest correlation. 85% (61 of the 72) of the circRNAs found in urine were also detected in plasma (Table 4). Figure 2b(plasma) and 2d (urine) display the number of detected circRNAs and the number that span introns, exons, and UTRs for both plasma and urine. The majority of circRNA identified in plasma and urine contain at least two exons and span an intron; 671 in plasma and 52 in urine; green bars (Fig. 2b, plasma and 2d, urine). A small number of circRNA are transcribed from a single exon (15 in plasma and 2 in urine).
Highly expressed, back-spliced junctions were validated by qRT-PCR
In order to validate predicted back-spliced junctions by qRT-PCR, we designed inward-facing primers for the 15 most highly expressed circRNA in each biofluid and tested each primer pair in samples from 10 different individuals, using the same source RNA for cDNA synthesis that was used for RNAseq (Fig. 3a,b). Figure 3a shows that the 15 circRNAs are detected in most of the 10 plasma samples. The numbers of samples are described in Table 6, and compared with the RNASeq detection for those circRNAs in the same samples. 13 primer pairs were validated in urine. Detection in urine samples was sparse, with fewer samples positive for each circRNA than for plasma (Fig. 3b and Table 6). For the two back-spliced junctions detected in RNASeq data, but not validated by qRT-PCR in urine (circMYO5B and circPHC3), it is possible that the circRNA primers did not work, or there were qPCR inhibitors in the sample, or the circRNA was not present. Two of the samples did not have enough assigned reads via RNASeq to be included, so the total number of samples was 8. In order to rule out chimeric junctions that might be present in DNA or resemble artifacts introduced during library preparation, we also used genomic DNA (gDNA) from each individual as a negative control. All 15 primer pairs used in the plasma and urine samples were not detected in gDNA (data not shown). Table 7 describes the rank from highest to lowest expression for each of the circRNA validated by qRT-PCR, and compares it with the expression detected with sequencing. Their ranks do not correlate well between the two platforms.
(a,b) Highly-expressed, predicted back-spliced junctions were validated by qRT-PCR. qRT-PCR validation of the 15 most highly expressed circRNA found in plasma (a) and urine (b), respectively. Each circRNA was examined in 10 cDNA samples from the same source RNA as sequenced samples. (c,d) Circular-linear ratios are higher in plasma than urine. Linear splice junction expression plotted against circular splice junction expression in plasma (c) and urine (d). Points representing circRNA between 1-fold and 5-fold higher than their linear counterparts are blue; 5x or higher are red.
Circular-to-linear RNA ratios
While the overall expression of most circRNAs is low compared to their linear counterparts, there are a number of circular RNA transcripts that have been described as more abundant than their linear host, cellularly as well as extracellularly23,27,62,63. We examined the circular-to-linear ratio (CLR) of circRNA transcripts found in plasma and urine as described previously; by taking the ratio of the circular, back-spliced junction counts compared to the linear count of the nearest 5′ or 3′ splice junction13,24,27. On average, 28.5% of circRNA transcripts in plasma and 21.5% of circRNA transcripts in urine have higher expression than their linear host gene (Fig. 3c, plasma and 3d, urine). Extracellular RNA is often fragmented and may have a 3′ bias64. Before examining the expression of circular RNA in relation to their host genes, we calculated the overall 5′ to 3′ coverage of linear transcripts and did not find a bias in our samples.
Participants sequenced 5 or more times have less inter-sample variation
A notable feature of this dataset is that many participants were sampled longitudinally, allowing for analysis of circRNA stability in individuals versus the entire dataset. Figure 4a,b show longitudinal circRNA expression in the same participants in plasma and urine, respectively. Broadly speaking, the heatmaps demonstrate similar expression patterners in the same participant over time. In order to assess variability within individuals, we calculated the coefficient of variation (CV) of circRNA expression, normalized to junction reads per million (JRPM). Here, we focus on participants sampled on 5 or more occasions over approximately one year. In both plasma and urine, the CV for each individual participant is displayed along with the CV for all participant samples. The data indicate that individuals have a statistically-significant consistency in circRNA expression pattern over time (Fig. 4c, plasma and 4d, urine).
Participants sequenced five or more times have less inter-sample variation. CircRNA populations identified in plasma (a,c) and urine (b,d) from participants sampled five or more times. (a,b) Heatmaps showing the log-normalized JRPM expression of plasma (a) and urine (b) samples taken longitudinally from the same participant. The coefficient of variation (CV) of circRNA expression is significantly lower across individual participant samples when compared to the entire dataset (c, plasma; and d, urine). ****p < = 0.0001.
Usage Notes
As the approach to detecting circRNA from RNA-Seq data differ with available tools, we employed 6 different bioinformatic tools: CIRCexplorer, CIRI2, DCC, KNIFE, find_circ, and MapSplice, in two clinically relevant biofluids, plasma and urine, using 134 and 114 samples, respectively. Most of these circRNA pipelines use an external aligner, such as bowtie, STAR, or bwa, to align reads to the genome and/or transcriptome (Table 2). After alignment, reads that contiguously align to the genome and/or transcriptome are filtered out, and the remaining unmapped reads are further filtered to identify back-spliced junctions. Differences in circRNA identification algorithms include: 1) how paired-end reads and gene annotations are used, if at all, 2) the amount of overlap over the junction that a read must contain, 3) the types of junctions considered, and 4) various filtering steps (Table 1)65. We sought to generate a high confidence set of circRNA expressed in plasma and urine with the following requirements for each circRNA: 1) detection in at least 5 samples for each respective biofluid, 2) a minimum 18 nt overlap on either side of the junction, 3) at least two reads spanning the back-spliced junction, and 4) identification by all 6 tested bioinformatic tools as identification can vary widely between tools66,67,68.
We tested alignment parameters and their influence on the detection rate of circRNA and found that the number of input reads, genome mapped reads, and junction reads did not correlate well with the number of circRNA detected per sample; rather the number of reads assigned to the transcriptome had the greatest correlation with the number of circRNA (R2 = 0.805; data not shown).
Code availability
Code used for circRNA identification is available in the Supplemental data. Software versions used for analysis are as follows:
STAR v2.4.0j for DCC and CIRCexplorer
bowtie2 v2.2.1 for find_circ and KNIFE
bowtie v0.12.9 for MapSplice2
bwa v0.7.13 for CIRI2
bedtools v2.26.0
Data were analyzed using the R v3.3.2 statistical package (https://cran.r-project.org). UpSet plots were generated using the UpSetR v1.3.3 package54. The ReadqPCR v1.20.0 and NormqPCR v1.20.0 Bioconductor v3.4 packages were used for qRT-PCR data analysis59.
References
Byron, S. A., Van Keuren-Jensen, K. R., Engelthaler, D. M., Carpten, J. D. & Craig, D. W. Translating RNA sequencing into clinical diagnostics: opportunities and challenges. Nat Rev Genet 17, 257–271, https://doi.org/10.1038/nrg.2016.10 (2016).
Stępień, E. et al. The circulating non-coding RNA landscape for biomarker research: lessons and prospects from cardiovascular diseases. Acta Pharmacol Sin 39, 1085–1099, https://doi.org/10.1038/aps.2018.35 (2018).
Murillo, O. D. et al. exRNA Atlas Analysis Reveals Distinct Extracellular RNA Cargo Types and Their Carriers Present across Human Biofluids. Cell 177, 463–477 e415, https://doi.org/10.1016/j.cell.2019.02.018 (2019).
Pardini, B., Sabo, A. A., Birolo, G. & Calin, G. A. Noncoding RNAs in Extracellular Fluids as Cancer Biomarkers: The New Frontier of Liquid Biopsies. Cancers (Basel) 11, https://doi.org/10.3390/cancers11081170 (2019).
Sanger, H. L., Klotz, G., Riesner, D., Gross, H. J. & Kleinschmidt, A. K. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc Natl Acad Sci USA 73, 3852–3856, https://doi.org/10.1073/pnas.73.11.3852 (1976).
Nigro, J. M. et al. Scrambled exons. Cell 64, 607–613 (1991).
Capel, B. et al. Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell 73, 1019–1030 (1993).
Cocquerelle, C., Mascrez, B., Hetuin, D. & Bailleul, B. Mis-splicing yields circular RNA molecules. FASEB J 7, 155–160, https://doi.org/10.1096/fasebj.7.1.7678559 (1993).
Danan, M., Schwartz, S., Edelheit, S. & Sorek, R. Transcriptome-wide discovery of circular RNAs in Archaea. Nucleic Acids Res 40, 3131–3142, https://doi.org/10.1093/nar/gkr1009 (2012).
Lu, T. et al. Transcriptome-wide investigation of circular RNAs in rice. RNA 21, 2076–2087, https://doi.org/10.1261/rna.052282.115 (2015).
Wang, P. L. et al. Circular RNA is expressed across the eukaryotic tree of life. PLoS One 9, e90859, https://doi.org/10.1371/journal.pone.0090859 (2014).
Dang, Y. et al. Tracing the expression of circular RNAs in human pre-implantation embryos. Genome Biol 17, 130, https://doi.org/10.1186/s13059-016-0991-3 (2016).
Rybak-Wolf, A. et al. Circular RNAs in the Mammalian Brain Are Highly Abundant, Conserved, and Dynamically Expressed. Mol Cell 58, 870–885, https://doi.org/10.1016/j.molcel.2015.03.027 (2015).
Salzman, J., Gawad, C., Wang, P. L., Lacayo, N. & Brown, P. O. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS One 7, e30733, https://doi.org/10.1371/journal.pone.0030733 (2012).
Tan, W. L. et al. A landscape of circular RNA expression in the human heart. Cardiovasc Res 113, 298–309, https://doi.org/10.1093/cvr/cvw250 (2017).
Veno, M. T. et al. Spatio-temporal regulation of circular RNA expression during porcine embryonic brain development. Genome Biol 16, 245, https://doi.org/10.1186/s13059-015-0801-3 (2015).
You, X. et al. Neural circular RNAs are derived from synaptic genes and regulated by development and plasticity. Nat Neurosci 18, 603–610, https://doi.org/10.1038/nn.3975 (2015).
Ashwal-Fluss, R. et al. circRNA biogenesis competes with pre-mRNA splicing. Mol Cell 56, 55–66, https://doi.org/10.1016/j.molcel.2014.08.019 (2014).
Hansen, T. B. et al. Natural RNA circles function as efficient microRNA sponges. Nature 495, 384–388, https://doi.org/10.1038/nature11993 (2013).
Li, Z. et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol 22, 256–264, https://doi.org/10.1038/nsmb.2959 (2015).
Memczak, S. et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495, 333–338, https://doi.org/10.1038/nature11928 (2013).
Zhang, Y. et al. Circular intronic long noncoding RNAs. Mol Cell 51, 792–806, https://doi.org/10.1016/j.molcel.2013.08.017 (2013).
Jeck, W. R. et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19, 141–157, https://doi.org/10.1261/rna.035667.112 (2013).
Maass, P. G. et al. A map of human circular RNAs in clinically relevant tissues. J Mol Med (Berl) 95, 1179–1189, https://doi.org/10.1007/s00109-017-1582-9 (2017).
Bahn, J. H. et al. The landscape of microRNA, Piwi-interacting RNA, and circular RNA in human saliva. Clin Chem 61, 221–230, https://doi.org/10.1373/clinchem.2014.230433 (2015).
Alhasan, A. A. et al. Circular RNA enrichment in platelets is a signature of transcriptome degradation. Blood 127, e1–e11, https://doi.org/10.1182/blood-2015-06-649434 (2016).
Memczak, S., Papavasileiou, P., Peters, O. & Rajewsky, N. Identification and Characterization of Circular RNAs As a New Class of Putative Biomarkers in Human Blood. PLoS One 10, e0141214, https://doi.org/10.1371/journal.pone.0141214 (2015).
Preusser, C. et al. Selective release of circRNAs in platelet-derived extracellular vesicles. J Extracell Vesicles 7, 1424473, https://doi.org/10.1080/20013078.2018.1424473 (2018).
Savelyeva, A. V. et al. Variety of RNAs in Peripheral Blood Cells, Plasma, and Plasma Fractions. Biomed Res Int 2017, 7404912, https://doi.org/10.1155/2017/7404912 (2017).
Zhang, Y. G., Yang, H. L., Long, Y. & Li, W. L. Circular RNA in blood corpuscles combined with plasma protein factor for early prediction of pre-eclampsia. BJOG 123, 2113–2118, https://doi.org/10.1111/1471-0528.13897 (2016).
Kolling, M. et al. Circular RNAs in Urine of Kidney Transplant Patients with Acute T Cell-Mediated Allograft Rejection. Clin Chem 65, 1287–1294, https://doi.org/10.1373/clinchem.2019.305854 (2019).
Liu, B. et al. Characterization of tissue-specific biomarkers with the expression of circRNAs in forensically relevant body fluids. Int J Legal Med 133, 1321–1331, https://doi.org/10.1007/s00414-019-02027-y (2019).
Ma, H. et al. Differential expression study of circular RNAs in exosomes from serum and urine in patients with idiopathic membranous nephropathy. Arch Med Sci 15, 738–753, https://doi.org/10.5114/aoms.2019.84690 (2019).
Dou, Y. et al. Circular RNAs are down-regulated in KRAS mutant colon cancer cells and can be transferred to exosomes. Sci Rep 6, 37982, https://doi.org/10.1038/srep37982 (2016).
Lasda, E. & Parker, R. Circular RNAs Co-Precipitate with Extracellular Vesicles: A Possible Mechanism for circRNA Clearance. PLoS One 11, e0148407, https://doi.org/10.1371/journal.pone.0148407 (2016).
Li, Y. et al. Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Res 25, 981–984, https://doi.org/10.1038/cr.2015.82 (2015).
Bachmayr-Heyda, A. et al. Correlation of circular RNA abundance with proliferation–exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Sci Rep 5, 8057, https://doi.org/10.1038/srep08057 (2015).
Song, X. et al. Circular RNA profile in gliomas revealed by identification tool UROBORUS. Nucleic Acids Res 44, e87, https://doi.org/10.1093/nar/gkw075 (2016).
Chen, S. et al. Widespread and Functional RNA Circularization in Localized Prostate Cancer. Cell 176, 831–843 e822, https://doi.org/10.1016/j.cell.2019.01.025 (2019).
Ding, X. et al. Profiling expression of coding genes, long noncoding RNA, and circular RNA in lung adenocarcinoma by ribosomal RNA-depleted RNA sequencing. FEBS Open Bio 8, 544–555, https://doi.org/10.1002/2211-5463.12397 (2018).
Chen, F. et al. Circular RNAs expression profiles in plasma exosomes from early-stage lung adenocarcinoma and the potential biomarkers. J Cell Biochem 121, 2525–2533, https://doi.org/10.1002/jcb.29475 (2020).
Burgos, K. L. et al. Identification of extracellular miRNA in human cerebrospinal fluid by next-generation sequencing. RNA 19, 712–722, https://doi.org/10.1261/rna.036863.112 (2013).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12, https://doi.org/10.14806/ej.17.1.200 (2011).
Szabo, L. et al. Statistically based splicing detection reveals neural enrichment and tissue-specific induction of circular RNA during human fetal development. Genome Biol 16, 126, https://doi.org/10.1186/s13059-015-0690-5 (2015).
Wang, K. et al. MapSplice: accurate mapping of RNA-seq reads for splice junction discovery. Nucleic Acids Res 38, e178, https://doi.org/10.1093/nar/gkq622 (2010).
Zhang, X. O. et al. Complementary sequence-mediated exon circularization. Cell 159, 134–147, https://doi.org/10.1016/j.cell.2014.09.001 (2014).
Gao, Y., Wang, J. & Zhao, F. CIRI: an efficient and unbiased algorithm for de novo circular RNA identification. Genome Biol 16, 4, https://doi.org/10.1186/s13059-014-0571-3 (2015).
Cheng, J., Metge, F. & Dieterich, C. Specific identification and quantification of circular RNAs from sequencing data. Bioinformatics 32, 1094–1096, https://doi.org/10.1093/bioinformatics/btv656 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21, https://doi.org/10.1093/bioinformatics/bts635 (2013).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760, https://doi.org/10.1093/bioinformatics/btp324 (2009).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat Methods 9, 357–359, https://doi.org/10.1038/nmeth.1923 (2012).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10, R25, https://doi.org/10.1186/gb-2009-10-3-r25 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842, https://doi.org/10.1093/bioinformatics/btq033 (2010).
Conway, J. R., Lex, A. & Gehlenborg, N. UpSetR: an R package for the visualization of intersecting sets and their properties. Bioinformatics 33, 2938–2940, https://doi.org/10.1093/bioinformatics/btx364 (2017).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930, https://doi.org/10.1093/bioinformatics/btt656 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq 2. Genome Biol 15, 550, https://doi.org/10.1186/s13059-014-0550-8 (2014).
Koressaar, T. & Remm, M. Enhancements and modifications of primer design program Primer3. Bioinformatics 23, 1289–1291, https://doi.org/10.1093/bioinformatics/btm091 (2007).
Untergasser, A. et al. Primer3–new capabilities and interfaces. Nucleic Acids Res 40, e115, https://doi.org/10.1093/nar/gks596 (2012).
Perkins, J. R. et al. ReadqPCR and NormqPCR: R packages for the reading, quality checking and normalisation of RT-qPCR quantification cycle (Cq) data. BMC Genomics 13, 296, https://doi.org/10.1186/1471-2164-13-296 (2012).
Van Keuren-Jensen, K. & Huentelman, M. dbGaP https://identifiers.org/dbgap:phs001258.v2.p1 (2020).
Van Keuren-Jensen, K. et al. figshare https://doi.org/10.6084/m9.figshare.c.5420832 (2021).
Guo, J. U., Agarwal, V., Guo, H. & Bartel, D. P. Expanded identification and characterization of mammalian circular RNAs. Genome Biol 15, 409, https://doi.org/10.1186/s13059-014-0409-z (2014).
Salzman, J., Chen, R. E., Olsen, M. N., Wang, P. L. & Brown, P. O. Cell-type specific features of circular RNA expression. PLoS Genet 9, e1003777, https://doi.org/10.1371/journal.pgen.1003777 (2013).
Batagov, A. O. & Kurochkin, I. V. Exosomes secreted by human cells transport largely mRNA fragments that are enriched in the 3′-untranslated regions. Biol Direct 8, 12, https://doi.org/10.1186/1745-6150-8-12 (2013).
Szabo, L. & Salzman, J. Detecting circular RNAs: bioinformatic and experimental challenges. Nat Rev Genet 17, 679–692, https://doi.org/10.1038/nrg.2016.114 (2016).
Chen, I., Chen, C. Y. & Chuang, T. J. Biogenesis, identification, and function of exonic circular RNAs. Wiley Interdiscip Rev RNA 6, 563–579, https://doi.org/10.1002/wrna.1294 (2015).
Hansen, T. B., Veno, M. T., Damgaard, C. K. & Kjems, J. Comparison of circular RNA prediction tools. Nucleic Acids Res 44, e58, https://doi.org/10.1093/nar/gkv1458 (2016).
Zeng, X., Lin, W., Guo, M. & Zou, Q. A comprehensive overview and evaluation of circular RNA detection tools. PLoS Comput Biol 13, e1005420, https://doi.org/10.1371/journal.pcbi.1005420 (2017).
Acknowledgements
This work was funded by support from the Flinn Foundation (Grant Award #1994 and #2037) and by NIH grant UH2TR000891. We would like to thank Terry Lee, Dan Arment, Thad Ide, Dan Vooletich, Erin Griffin, Taylor Hanohano and Brian Roche from Riddell for their significant input, time, effort, and financial support. There were a large number of individuals that made the collection of these samples possible. We would like to thank the staff at Arizona State University: Todd Graham, Ray Anderson, Jean Boyd, Tim Cassidy, Anikar Chhabra, and Jerry Neilly. We would also like to thank Ann Marie Bothwell (Desert Testing), April Allen, Yana Gadev, Stephanie Buchholtz, Cassandra Lucas, Therese de la Torre, Brian Anderson, Stephanie Althoff, and Brian Churas, Sean Allen, Ryan Bruhns, Ashley Suiter, Brandon Chaves, Mari Turk, Khalouk Shahbander, Michael Schmalle, Kirk Ryden, and Alex Starr for assistance with sample collection.
Author information
Authors and Affiliations
Contributions
Conceptualization, E.H. and K.V.K.J.; Methodology, E.H. and K.V.K.J.; Validation, E.H. and J.W.; Formal Analysis, E.H.; Investigation, E.H., R.R., J.W., R.R., T.B., E.C., A.J., A.S., C.B., J.A.; Resources, M.A., R.M. Y.K.; Writing - Original Draft, E.H. and K.V.K.J.; Writing, E.H., T.G.W. and K.V.K.J.; Review & Editing, E.H., T.G.W., Y.K., M.J.H., K.V.K.J. Funding Acquisition, M.H., Y.K. and K.V.K.J.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Online-only Table
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files associated with this article.
About this article
Cite this article
Hutchins, E., Reiman, R., Winarta, J. et al. Extracellular circular RNA profiles in plasma and urine of healthy, male college athletes. Sci Data 8, 276 (2021). https://doi.org/10.1038/s41597-021-01056-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-021-01056-w