Abstract
Toxicogenomics (TGx) approaches are increasingly applied to gain insight into the possible toxicity mechanisms of engineered nanomaterials (ENMs). Omics data can be valuable to elucidate the mechanism of action of chemicals and to develop predictive models in toxicology. While vast amounts of transcriptomics data from ENM exposures have already been accumulated, a unified, easily accessible and reusable collection of transcriptomics data for ENMs is currently lacking. In an attempt to improve the FAIRness of already existing transcriptomics data for ENMs, we curated a collection of homogenized transcriptomics data from human, mouse and rat ENM exposures in vitro and in vivo including the physicochemical characteristics of the ENMs used in each study.
Measurement(s) | microarray data • transcriptome • RNA • Toxicogenomics |
Technology Type(s) | digital curation |
Factor Type(s) | exposure to engineered nanomaterials |
Sample Characteristic - Organism | Homo sapiens • Mus musculus • Rattus |
Machine-accessible metadata file describing the reported data: https://doi.org/10.6084/m9.figshare.13154381
Similar content being viewed by others
Background & Summary
Engineered nanomaterials (ENMs) are an emerging class of chemicals with great technological and societal impact. Their unique physicochemical properties have already inspired multitudes of applications, ranging from medicine to industry and consumer products. While these unique properties make ENMs attractive for endless applications, they can also be responsible for potentially harmful effects on human health and the environment. ENMs can be synthesized in various sizes, shapes and chemistries with the smallest differences in the composition leading to novel properties and effects that need to be considered. Rigorous risk assessment is needed to ensure the safety of ENMs. Toxicogenomics (TGx) has emerged as a complementary approach to traditional toxicology with the potential to facilitate faster and cheaper hazard assessment of ENMs1,2. The large-scale profiling of exposure-induced molecular alterations sets the stage for mechanistic toxicology and expedites the development of predictive models. Furthermore, the application of TGx data to nanosafety can provide novel possibilities of grouping and classifying ENMs based on the similarity of molecular alterations in biological systems and furthermore can help to derive biomarkers to identify nano-specific signatures.
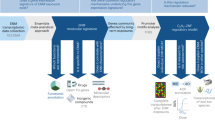
Transcriptomics technologies are the frontline of TGx. Vast amounts of transcriptomics data for multiple ENMs have already been generated offering a valuable resource for future studies and applications. However, the data are scattered across public repositories, and their FAIRness is currently hampered by their heterogeneous nature and lack of standardization in the preprocessing and analysis. The FAIR principles for scientific data were defined in 2016 and have since been the guide for more Findable, Accessible, Interoperable, and Reusable data3. The FAIRness of ENM-relevant databases, including ArrayExpress, the Gene Expression Omnibus (GEO), eNanoMapper and NanoCommons have recently been evaluated, and while the six datasets extracted from these met the majority of the criteria defined by the FAIR maturity indicators, areas identified for improvement included the use of standard schema for metadata and the presence of specific attributes in registries of repositories that would increase the FAIRness of datasets4. In order to unleash the full potential of already existing transcriptomics data on ENM exposures, which are lacking the metadata related to the exposure conditions and ENM characteristics, we created a unified collection of 101 manually curated and preprocessed data sets, covering a range of ENMs, organisms, and exposure setups, using the approach represented in Fig. 1.
The overarching aim of this study was to manually curate a comprehensive collection of transcriptomics data in the field of nanosafety, thereby increasing the degree of FAIRness of the original data sets. In particular, our collection is characterized by a higher degree of FAIRness as compared to the individual original data sets composing it.
Methods
Data set identification and collection
The first step in compiling the collection was to identify relevant data sets across public repositories. The search was limited to human, mouse, and rat data. We queried the Gene Expression Omnibus (GEO) and ArrayExpress databases with the following search terms: “engineered nanomaterial”, “nanomaterial” and “nanoparticle”. The initial collection yielded 124 unique entries, which went through manual assessment. Raw, non-normalized data for each microarray-based entry was downloaded from the series entry page, while for RNA-Seq data sets raw sequencing data in .fastq format were retrieved from the European Nucleotide Archive (ENA) (https://www.ebi.ac.uk/ena/browser/home).
Metadata curation
Next, supporting information (metadata) for each entry in the initial collection was downloaded and manually curated on R (version 3.5.2). Metadata gives context to the data by mapping each sample to biological variables, such as dose and time point, as well as technical variables crucial for the preprocessing of the data.
Metadata were obtained from the sample records of GEO entries by using the function getGEO from the R package GEOquery5. For data sets available only on ArrayExpress, the sample information for each entry was downloaded. These data were then manually curated to produce a homogenized file for each data set consisting of the following variables: GSE (a unique identifier for each data set), GSM (sample id), treatment (exposure; i.e. ENM or control), group (experimental group; combination of a unique exposure, dose, and time point), organism, biological system, dose, dose unit, time point, time point unit, slide, array, dye and platform. Although some of these variables are not relevant for RNA-Seq data, all the columns were included for all the data to ensure convenient data usability. The nomenclature was unified to an extent that could be reached based on the information provided in the original metadata. Each sample was then mapped to its corresponding raw data file (column filenames) or annotated later to the fastq-files based on the sample names (GSM). If one or more predefined technical variables were missing, the column was left empty (NA). However, if biological variables were missing or ambiguous, the data set was discarded. Lastly, for entries containing human primary cells, the donor was further included in the metadata as an additional column donor.
ENM physicochemical characteristics curation
The majority of the datasets were associated with a published article describing the study and including some details of the materials used and their physico-chemical characteristics. In some cases, the information provided was the nominal size information from the ENM manufacturer, while others provided more detailed characterization of the ENM in the exposure medium. Newer studies tended to provide more detailed characterization information than older ones, as the community knowledge regarding minimum characterization needs and properties influencing ENM toxicity increased6,7. Several of the studies utilized ENMs already used in previous studies and referred to the characterization provided in those earlier studies, in which case the information was manually extracted from the earlier papers. The curated information for the ENMs includes information on the supplier (including batch and lot information where available), the purity / impurities, the nominal size and surface area, as well as characterization data such as the core particle size (shape) as determined by Transmission Electron Microscopy (TEM) size, the hydrodynamic size and zeta potential (surface charge) in water and/or the exposure medium determined by Dynamic Light Scattering (DLS), information on the presence of endotoxin contamination (where provided) and a link to the commercial providers material specification sheet where relevant. As many of the studies utilized several different ENMs, or several variants (e.g. sizes, capping agents, polymeric coatings etc.) each individual ENM within each study is described in a separate row of the ENM characteristics datasheet.
Manual quality assessment
The quality of transcriptomics data is highly dependent on the experimental design2. Low number of replicates results in weak statistics, while transcriptomics technologies themselves are often prone to technical bias. In order to ensure the quality and usability of each individual data set, evaluation was carried out based on the availability of raw data and supporting information as well as technical aspects of the experimental setup. The experiment was considered inappropriate for the collection if the experimental groups consisted of less than three biological replicates or if the experimental design introduced an unmanageable batch effect. Such batch effects were commonly introduced by consistently labeling different experimental groups with separate dyes in a two-color microarray experiment (i.e. lack of dye swapping). Furthermore, data sets representing non-commercial/custom or marginally represented platforms, for instance microarrays specific for miRNA or lncRNA, were excluded. As a result, only commercial gene expression microarrays from Agilent, Affymetrix, and Illumina were included alongside Illumina RNA-Seq platforms. The manual quality assessment of the collection is further described in the section Technical Validation.
Data preprocessing
Preprocessing of transcriptomics data must be performed prior to any further analysis. The current standard preprocessing pipeline for microarray data includes steps for sample quality checking, probe filtering, data normalization, batch effect assessment and correction as well as probe annotation8. Similarly, the state-of-the-art preprocessing of RNA-Seq data includes quality control, read alignment, read count extraction, filtering low counts, normalization, and batch effect assessment8. Here, each data set was preprocessed and analyzed individually. Data sets consisting of several cell lines or tissues were further separated by the biological system to better focus on the transcriptional differences between the exposures.
Preprocessing was performed in the R programming language (R version 3.5.2) following standard preprocessing pipelines suitable for each platform. For Agilent and Affymetrix microarrays, the preprocessing was implemented in the software eUTOPIA9. For Illumina BeadChips, a similar approach was applied following the suggested workflow of the R Bioconductor package lumi10. The preprocessing workflow applied to each platform is summarized in Fig. 2.
Preprocessing workflow applied to Agilent, Affymetrix, and Illumina microarrays and Illumina RNA-sequencing. Boxes with a blue background represent preprocessing steps and methods applied for each platform while boxes outlined with a dashed line represent the output obtained for each data set. The lack of a white box indicates that the step was not applied for the platform.
Quality check
Omics data are prone to technical errors that can arise from sample handling as well as the intrinsic characteristics of the platforms8. For this, an important step prior to any manipulation of the data is the quality check (QC) that allows the assessment of the gene expression distributions across samples revealing outliers and poor-quality samples. We applied a platform specific QC on each data set to evaluate the quality of the samples as well as the prevalence of outliers in the data.
For Agilent microarrays, the R package arrayQualityMetrics11 was used, while the QC for Affymetrix was performed using the R packages affyQCreport12 and yaqcaffy13. Outliers were further assessed based on the visual representation in the form of density plots, bar plots, dendrograms, and multi-dimensional scaling (MDS) plots, which were also the primary method of outlier detection for Illumina arrays. Outliers were removed from subsequent preprocessing and analysis.
Quality checking of the RNA sequencing data was performed using FastQC v0.11.7 (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/).
Read alignment
RNA sequencing reads of mouse samples were aligned against the mouse reference genome assembly GRCm38, while sequencing reads of human samples were aligned against the human reference genome assembly GRCh38. The alignment was performed using the HISAT2 algorithm14,15 employing the genome indexes built for usage with HISAT2 (retrieved from https://ccb.jhu.edu/software/hisat2/manual.shtml). Sequencing file format conversions, such as.sam to.bam, sorting and extraction of uniquely mapped reads were performed using SAMtools (version 1.8-27-g0896262)16.
Read counts extraction
Raw read counts for the RNA-Seq data were computed using the R package Rsubread (v2.2.3)17. The human Gencode version 35 annotation was applied for read counts extraction of human samples, while for mouse samples the mouse Gencode version M25 was employed. Both of the annotations were downloaded from https://www.gencodegenes.org.
Low counts filtering
In order to filter out the transcripts with low expression levels in the samples of each RNA-Seq dataset, the proportion test was used as implemented in the Bioconductor NOISeq package (v2.31.0)18.
Probe filtering
For microarray experiments, probe filtering is commonly applied to remove probes showing low variance in the intensity range similar to the background8. These low-intensity probes were removed prior to data normalization. For Agilent microarrays, filtering was based on estimating the robustness of the probe signal intensities against the background (negative control probes) and applying a quantile-based method for eliminating probes with low signals. Individual thresholds based on the data and the number of experimental groups and replicates were determined for Agilent. For Illumina gene expression microarrays, probe filtering was performed after normalization based on the detection p-values10 provided in the raw data. Only probes with a detection p-value < 0.01 in at least one sample were considered for further analysis.
Normalization
Normalization of transcriptomics data is crucial for robust comparisons of gene expression. Here, the normalization of the expression signal distribution in the samples was performed on the log2 transformed signal intensities using the quantile normalization from the R package limma19 for Agilent, and the function justRMA from the package affy20 for Affymetrix microarrays, respectively. For Illumina microarrays, quantile normalization was performed with the function lumiN from the lumi R package10, while for Illumina RNA-Seq data, normalization was performed using the Bioconductor DESeq. 2 package21. In detail, the filtered raw counts underwent normalization by median of ratios method implemented in the package (for details see DESeq. 2 documentation).
Batch effect assessment and correction
Microarray experiments are susceptible to technical variation arising from the experimental setup, sample preparation, and the equipment, for example. This type of variation can lead to decreased quality and incorrect results. Thus, reducing the variation associated with technical variables (batch effect), while maintaining biological variation, improves the robustness of the results. Here, batch effects were evaluated by inspecting the results of principal component analysis, hierarchical clustering and multi-dimensional scaling9. Technical variation arising from unknown batches were evaluated with the function sva from the R package sva22. If variation associated to known technical variables or any of the surrogate variables was observed, its correlation with biological variables of interest was assessed via a confounding plot23. Batches that were not confounded with any of the variables of interest were corrected using the ComBat24 function from the R package sva22.
Probe annotation
Lastly, it is meaningful to map the probes to genes. For Agilent, the latest version of the annotation file for the specific microarray design was downloaded from the Agilent eArray website (https://earray.chem.agilent.com/earray/, 2020), and the probes were mapped to the Ensembl transcript IDs25. For Affymetrix gene expression arrays, the latest available alternative CDF files with Ensembl gene ID mappings were downloaded from Brainarray (http://brainarray.mbni.med.umich.edu/Brainarray/Database/CustomCDF/CDF_download.asp, 2020), while for Illumina BeadChips, the platform specific R annotation packages (illuminaHumanv3.db26, illuminaHumanv4.db27, illuminaRatv1.db28 or illuminaMousev2.db29) were used.
Multiple probes mapped onto the same gene ID were summarized by their median values. Agilent probes that were initially annotated to Ensembl transcripts were further mapped to the corresponding Ensembl gene IDs. If multiple transcripts were mapped to the same gene, the one with the highest absolute score, as calculated by the -log(p-value) x log2(fold change) for each exposure vs. control pairwise comparison, was selected.
Differential expression analysis
Transcriptomics analysis aims at identifying gene expression differences between biological conditions. Here, we performed a differential expression analysis on each microarray data set using the R package limma19. Comparisons were made between each specific experimental group consisting of a single exposure, dose, and time point and its corresponding control samples. Batch corrected variables were included as covariates of the linear model. In case the biological material was obtained from human donors, the donor was included as a covariate for the analysis. For RNA-Seq based data sets similar comparisons were made using the Bioconductor DESeq. 2 package21.
As a result of the differential expression analysis, we provide full lists of genes with their specific fold changes and statistics as well as the results filtered to only contain significantly differentially expressed genes with the threshold of |logFC| > 0.58 and Benjamini & Hochberg adjusted p-value < 0.05. Due to the implementation of DESeq. 2 independent filtering (for details see DESeq. 2 documentation), we also computed the adjusted p-values for RNA-Seq data externally from DESeq. 2 to obtain the full list of adjusted p-values with no missing values. These values are included in the unfiltered result files of the differential expression analysis under the column “adj.P.Val.no.ind.filt”.
FAIRness optimization
To further assist accessibility, interoperability and reusability, the data sets have been curated, imported and made publicly available from the NanoPharos database (https://db.nanopharos.eu/), which has been developed under the Horizon 2020 (H2020) NanoSolveIT30 (https://www.nanosolveit.eu) and NanoCommons projects (https://nanocommons.eu/). The NanoPharos database has been primarily developed to include computationally derived data based on simulations for ENMs at different levels of accuracy. The database was then further extended to include ENM characterization data and biological effects. With the inclusion of omics data, the NanoPharos database is now covering, in a ready for modelling format, the full spectrum of data needed to initiate a computational workflow for in silico exploitation of the data. The data set was checked for inconsistencies in the data structure and harmonized where needed. The ENM physico-chemical characterization data have been enriched, where applicable, with molecular (e.g. atomic/ionic radii, electronegativity, energy band gap) and structural (e.g. crystallographic space group, unit cell dimensions and angles). Each ENM has been linked to the respective transcriptomics data set to facilitate querying and user study. The datasets can be queried and grouped, among others, based on the ENM core material, ENM batch, exposure time and dose, biological information, experiment type, analysis platform etc. (Supplementary File 1).
The NanoPharos database has been designed under the FAIR data principles3 to offer users with high-quality, ready-for-modelling data sets, while allowing further development, adaptation and expansion. The FAIR data principles are meant to help database managers to improve data accessibility and reusability from the wider community in a way resembling Library Science31. To achieve this, data digitization in the NanoPharos database is being optimized to be machine readable to allow the seamless data comparison, transformation and, where possible, combination, providing the user with bigger and more complete data sets. On top of that, the NanoPharos database goes beyond the technical character of the FAIR data principles and is implementing the scientific FAIR data principles (SFAIR) as defined recently by Papadiamantis et al.31, providing users with the necessary scientific context and background information for them to be able to reuse the data with the highest possible confidence. Furthermore, NanoPharos is readily accessible via Representational State Transfer (REST) application programming interface (API) and is able to interact with external databases (e.g. NanoSolveIT Cloud) and modelling tools through API programmatic access. The available datasets can be accessed through: https://db.nanopharos.eu/Queries.
Data Records
The data collection32 generated here is freely available on Zenodo at https://doi.org/10.5281/zenodo.4146981. The collection comprises 85 preprocessed microarray-based data sets totaling 506 unique ENM vs. control comparisons and 16 RNA-Seq based data sets representing 23 ENM vs. control comparisons. Additionally, 24 comparisons of non-nanoparticle compounds used as positive/negative controls in the original experiments are included for the microarray data sets and 7 additional compounds are included for the RNA-Seq data. All of the data sets and their descriptions are available in Online-only Table 1, while the physico-chemical characteristics of the tested ENMs are available in Online-only Table 2, respectively.
In order to facilitate the selection of data suitable for different applications and modelling approaches, we classified the data into four categories based on the experimental design as follows:
I – Multiple doses, multiple time points.
II – Multiple doses, one time point.
III – One dose, multiple time points.
IV – One dose, one time point.
The proportion of each data class in the collection is visualized in Fig. 3a. Each class contains data obtained both in vivo and in vitro with at least two organisms represented (Fig. 3b). The collection covers a range of ENM compositions, as well as variants in size, shape, surface capping/coating etc. within a specific composition, in multiple biological systems in these organisms (Fig. 3c,d).
The data collection comprises of various experimental setups and exposures of multiple ENM compositions. (a) The total of 101 data sets were divided into four classes based on the experimental setup. The pie chart represents the distribution of data sets by class. (b) Bars representing the proportion of data sets in each organism divided by the four classes. In vivo and in vitro exposures are separated. (c) Horizontal bars represent the number of data sets with the specific ENM core material or material type. Grey bars represent in vivo exposures and pink bars in vitro exposures, respectively. (d) Bars represent the biological system used in the experiment. In vitro exposures are represented by pink bars and in vivo exposures by grey bars.
Files available for each data set
Each data set contains a homogenized metadata file, normalized and batch corrected expression matrices as well as complete and filtered results of the differential expression analysis (Table 1).
Technical Validation
The quality of transcriptomics data is a product of careful design of the experiment, technical execution as well as reporting of the data. The results of each downstream analysis substantially rely on the quality of the data. For this, we ensured that the collection contains high-quality data sets and defined a selection of criteria for data sets to be included:
-
Three or more biological replicates are included for statistical robustness
-
Microarray platform is a commercial gene expression microarray produced by Agilent, Affymetrix or Illumina
-
The labelling of 2-color microarrays has been done considering dye swapping
-
Non-normalized raw data is available
-
Supporting information reports all variables required for preprocessing
-
Untreated control samples are included
Each entry was evaluated based on the criteria, and either removed from the collection or selected for further preprocessing and analysis. The number of entries discarded for each of the listed reasons is represented in Table 2. Out of the 124 original entries 84 passed the quality assessment and were further divided into a total of 101 data sets (85 microarray and 16 RNA-Seq) based on the biological systems as specified in Data preprocessing.
Usage Notes
Here we provide the biggest homogenized collection of transcriptomics data sets in the field of nanosafety supplemented with metadata and ENM physico-chemical characteristics. The collection offers a valuable source for multiple analysis and modeling approaches33. For instance, the mechanism of action of each ENM can be characterized by investigating the provided lists of differentially expressed genes, and may be linked to specific physico-chemical characteristics such as size, surface capping or coating which can guide redesign of ENMs that are safer and may support grouping into sets of nanoforms in accordance with REACH regulation (https://echa.europa.eu/documents/10162/13655/how_to_register_nano_en.pdf/f8c046ec-f60b-4349-492b-e915fd9e3ca0), for example. Moreover, pathway enrichment analysis can be performed to annotate these genes onto biological functions34. ENMs can be further compared and grouped based on the similarities between their molecular alteration profiles.
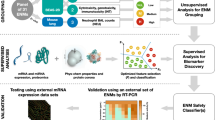
Due to the homogenized preprocessing and manual curation of the metadata, this collection is a relevant resource for identification of toxicity biomarkers. This can be addressed by using multiple feature selection approaches35,36 or more advanced data modelling techniques37,38,39. Biomarkers could also be detected by means of gene co-expression network analysis, under the assumption that central network genes play a key role in the adaptation to the exposure40,41.
The availability of data for multiple organisms or tissues can contribute to the development of more accurate adverse outcome pathways by linking ENM-specific molecular initiating events with cascades of relevant biological processes leading to an adverse outcome42,43. In addition, our data collection can be easily integrated with other transcriptomics data in the context of a read-across analysis to identify similarities in the molecular alterations induced by the ENMs with other phenotypic entities such as chemicals, drugs, and diseases44. Moreover, the data sets that we denoted as class I and II, where exposure at multiple doses are available, can be further analyzed to identify dose-dependent molecular alterations45,46,47,48.
Our manually curated transcriptomics data collection with supporting ENM descriptions will have a high impact on the nanosafety community and can aid the development of new methodologies for nanomaterial safety assessment2,8,30,33,43.
Code availability
Preprocessing of the data was performed on R version 3.5.2. The preprocessing of Agilent and Affymetrix expression data was performed using eUTOPIA9, an R shiny software freely available on https://github.com/Greco-Lab/eUTOPIA. Custom scripts used for preprocessing of Illumina BeadChip and RNA sequencing data are available on GitHub on https://github.com/grecolab/Public_Nano.
References
Liu, Z., Huang, R., Roberts, R. & Tong, W. Toxicogenomics: A 2020 vision. Trends Pharmacol. Sci. 40, 92–103 (2019).
Kinaret, P. A. S. et al. Transcriptomics in toxicogenomics, part I: experimental design, technologies, publicly available data, and regulatory aspects. Nanomaterials (Basel) 10 (2020).
Wilkinson, M. D. et al. The FAIR Guiding Principles for scientific data management and stewardship. Sci. Data 3, 160018 (2016).
Ammar, A. et al. A Semi-Automated Workflow for FAIR Maturity Indicators in the Life Sciences. Nanomaterials 10, 2068 (2020).
Davis, S. & Meltzer, P. S. GEOquery: a bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics 23, 1846–1847 (2007).
Rasmussen, K. et al. Physico-chemical properties of manufactured nanomaterials - Characterisation and relevant methods. An outlook based on the OECD Testing Programme. Regul Toxicol Pharmacol 92, 8–28 (2018).
Stefaniak, A. B. et al. Nanoscale reference materials for environmental, health and safety measurements: needs, gaps and opportunities. Nanotoxicology 7, 1325–1337 (2013).
Federico, A. et al. Transcriptomics in toxicogenomics, part II: preprocessing and differential expression analysis for high quality data. Nanomaterials (Basel) 10 (2020).
Marwah, V. S. et al. eUTOPIA: solUTion for Omics data PreprocessIng and Analysis. Source Code Biol. Med. 14, 1 (2019).
Du, P., Kibbe, W. A. & Lin, S. M. lumi: a pipeline for processing Illumina microarray. Bioinformatics 24, 1547–1548 (2008).
Kauffmann, A., Gentleman, R. & Huber, W. arrayQualityMetrics–a bioconductor package for quality assessment of microarray data. Bioinformatics 25, 415–416 (2009).
Parman, C., Halling, C. & Gentleman, R. affyQCReport: QC Report Generation for affyBatch objects. R package version 1.66.0. (2020).
Gatto, L. yaqcaffy: Affymetrix expression data quality control and reproducibility analysis. R package version 1.48.0. (2020).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Liao, Y., Smyth, G. K. & Shi, W. The R package Rsubread is easier, faster, cheaper and better for alignment and quantification of RNA sequencing reads. Nucleic Acids Res. 47, e47 (2019).
Tarazona, S. et al. Data quality aware analysis of differential expression in RNA-seq with NOISeq R/Bioc package. Nucleic Acids Res. 43, e140 (2015).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Gautier, L., Cope, L., Bolstad, B. M. & Irizarry, R. A. affy–analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20, 307–315 (2004).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq. 2. Genome Biol. 15, 550 (2014).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Lauss, M. swamp: Visualization, Analysis and Adjustment of High-Dimensional Data in Respect to Sample Annotations. R package version 1.4.2. (2018).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Hunt, S. E. et al. Ensembl variation resources. Database (Oxford) 2018 (2018).
Dunning, M., Lynch, A. & Eldridge, M. illuminaHumanv3.db: Illumina HumanHT12v3 annotation data (chip illuminaHumanv3). R package version 1.26.0. (2015).
Dunning, M., Lynch, A. & Eldridge, M. illuminaHumanv4.db: Illumina HumanHT12v4 annotation data (chip illuminaHumanv4). R package version 1.26.0. (2015).
Dunning, M., Lynch, A. & Eldridge, M. illuminaRatv1.db: Illumina Ratv1 annotation data (chip illuminaRatv1). R package version 1.26.0. (2015).
Dunning, M., Lynch, A. & Eldridge, M. illuminaMousev2.db: Illumina MouseWG6v2 annotation data (chip illuminaMousev2). R package version 1.26.0. (2015).
Afantitis, A. et al. NanoSolveIT Project: Driving nanoinformatics research to develop innovative and integrated tools for in silico nanosafety assessment. Comput Struct Biotechnol J 18, 583–602 (2020).
Papadiamantis, A. G. et al. Metadata Stewardship in Nanosafety Research: Community-Driven Organisation of Metadata Schemas to Support FAIR Nanoscience Data. Nanomaterials (Basel) 10 (2020).
Saarimäki, L. S. et al. Manually curated transcriptomics data collection for toxicogenomic assessment of engineered nanomaterials. Zenodo https://doi.org/10.5281/zenodo.4146981 (2020).
Serra, A. et al. Transcriptomics in toxicogenomics, part III: data modelling for risk assessment. Nanomaterials (Basel) 10 (2020).
Scala, G., Serra, A., Marwah, V. S., Saarimäki, L. A. & Greco, D. FunMappOne: a tool to hierarchically organize and visually navigate functional gene annotations in multiple experiments. BMC Bioinformatics 20, 79 (2019).
Fortino, V., Kinaret, P., Fyhrquist, N., Alenius, H. & Greco, D. A robust and accurate method for feature selection and prioritization from multi-class OMICs data. PLoS ONE 9, e107801 (2014).
Díaz-Uriarte, R. & Alvarez de Andrés, S. Gene selection and classification of microarray data using random forest. BMC Bioinformatics 7, 3 (2006).
Fortino, V., Scala, G. & Greco, D. Feature set optimization in biomarker discovery from genome scale data. Bioinformatics https://doi.org/10.1093/bioinformatics/btaa144 (2020).
Singh, A. et al. DIABLO: an integrative approach for identifying key molecular drivers from multi-omics assays. Bioinformatics 35, 3055–3062 (2019).
Serra, A., Önlü, S., Festa, P., Fortino, V. & Greco, D. MaNGA: a novel multi-objective multi-niche genetic algorithm for QSAR modelling. Bioinformatics https://doi.org/10.1093/bioinformatics/btz521 (2019).
Marwah, V. S. et al. Inform: inference of network response modules. Bioinformatics 34, 2136–2138 (2018).
Kinaret, P. et al. Network analysis reveals similar transcriptomic responses to intrinsic properties of carbon nanomaterials in vitro and in vivo. ACS Nano 11, 3786–3796 (2017).
Nymark, P. et al. A data fusion pipeline for generating and enriching adverse outcome pathway descriptions. Toxicol. Sci. 162, 264–275 (2018).
Nymark, P. et al. Toward rigorous materials production: new approach methodologies have extensive potential to improve current safety assessment practices. Small e1904749 https://doi.org/10.1002/smll.201904749 (2020).
Serra, A. et al. INSIdE NANO: a systems biology framework to contextualize the mechanism-of-action of engineered nanomaterials. Sci. Rep. 9, 179 (2019).
Serra, A., Saarimäki, L. A., Fratello, M., Marwah, V. S. & Greco, D. BMDx: a graphical Shiny application to perform Benchmark Dose analysis for transcriptomics data. Bioinformatics https://doi.org/10.1093/bioinformatics/btaa030 (2020).
Phillips, J. R. et al. BMDExpress 2: enhanced transcriptomic dose-response analysis workflow. Bioinformatics 35, 1780–1782 (2019).
Serra, A. et al. TinderMIX: Time-dose integrated modelling of toxicogenomics data. Gigascience 9 (2020).
Schüttler, A. et al. Map and model-moving from observation to prediction in toxicogenomics. Gigascience 8 (2019).
Snyder-Talkington, B. N. et al. Multiwalled carbon nanotube-induced pulmonary inflammatory and fibrotic responses and genomic changes following aspiration exposure in mice: A 1-year postexposure study. J Toxicol Environ Health Part A 79, 352–366 (2016).
Dymacek, J. & Guo, N. L. Systems Approach to Identifying Relevant Pathways from Phenotype Information in Dose-Dependent Time Series Microarray Data. Proceedings (IEEE Int Conf Bioinformatics Biomed) 2011, 290–293 (2011).
Bourdon, J. A. et al. Hepatic and pulmonary toxicogenomic profiles in mice intratracheally instilled with carbon black nanoparticles reveal pulmonary inflammation, acute phase response, and alterations in lipid homeostasis. Toxicol. Sci. 127, 474–484 (2012).
Husain, M. et al. Pulmonary instillation of low doses of titanium dioxide nanoparticles in mice leads to particle retention and gene expression changes in the absence of inflammation. Toxicol. Appl. Pharmacol. 269, 250–262 (2013).
Tilton, S. C. et al. Three human cell types respond to multi-walled carbon nanotubes and titanium dioxide nanobelts with cell-specific transcriptomic and proteomic expression patterns. Nanotoxicology 8, 533–548 (2014).
Ronzani, C., Safar, R., Le Faou, A., Rihn, B. H. & Joubert, O. Comment on: S-nitrosoglutathione (GSNO) is cytotoxic to intracellular amastigotes and promotes healing of topically treated Leishmania major or Leishmania braziliensis skin lesions. J. Antimicrob. Chemother. 69, 2300–2302 (2014).
Poulsen, S. S. et al. MWCNTs of different physicochemical properties cause similar inflammatory responses, but differences in transcriptional and histological markers of fibrosis in mouse lungs. Toxicol. Appl. Pharmacol. 284, 16–32 (2015).
Bajak, E. et al. Changes in Caco-2 cells transcriptome profiles upon exposure to gold nanoparticles. Toxicol. Lett. 233, 187–199 (2015).
Halappanavar, S. et al. Transcriptional profiling identifies physicochemical properties of nanomaterials that are determinants of the in vivo pulmonary response. Environ Mol Mutagen 56, 245–264 (2015).
Nymark, P. et al. Extensive temporal transcriptome and microRNA analyses identify molecular mechanisms underlying mitochondrial dysfunction induced by multi-walled carbon nanotubes in human lung cells. Nanotoxicology 9, 624–635 (2015).
Pisani, C. et al. High-throughput, quantitative assessment of the effects of low-dose silica nanoparticles on lung cells: grasping complex toxicity with a great depth of field. BMC Genomics 16, 315 (2015).
Rahman, L., Wu, D., Johnston, M., William, A. & Halappanavar, S. Toxicogenomics analysis of mouse lung responses following exposure to titanium dioxide nanomaterials reveal their disease potential at high doses. Mutagenesis 32, 59–76 (2017).
Pisani, C. et al. Biocompatibility assessment of functionalized magnetic mesoporous silica nanoparticles in human HepaRG cells. Nanotoxicology 11, 871–890 (2017).
Ilves, M. et al. Surface PEGylation suppresses pulmonary effects of CuO in allergen-induced lung inflammation. Part Fibre Toxicol 16, 28 (2019).
Kooter, I. et al. Molecular Signature of Asthma-Enhanced Sensitivity to CuO Nanoparticle Aerosols from 3D Cell Model. ACS Nano 13, 6932–6946 (2019).
Thai, S. et al. Differential Genomic Effects of Six Different Nanomaterials on Human Liver HepG2 Cells. NCBI GEO (2018).
Sellamuthu, R., Umbright, C., Li, S., Kashon, M. & Joseph, P. Mechanisms of crystalline silica-induced pulmonary toxicity revealed by global gene expression profiling. Inhal Toxicol 23, 927–937 (2011).
Søs Poulsen, S. et al. Transcriptomic analysis reveals novel mechanistic insight into murine biological responses to multi-walled carbon nanotubes in lungs and cultured lung epithelial cells. PLoS One 8, e80452 (2013).
Böhmert, L., Niemann, B., Lichtenstein, D., Juling, S. & Lampen, A. Molecular mechanism of silver nanoparticles in human intestinal cells. Nanotoxicology 9, 852–860 (2015).
Perkins, T. N. et al. Indications for distinct pathogenic mechanisms of asbestos and silica through gene expression profiling of the response of lung epithelial cells. Hum. Mol. Genet. 24, 1374–1389 (2015).
Rahman, L. et al. Multi-walled carbon nanotube-induced genotoxic, inflammatory and pro-fibrotic responses in mice: Investigating the mechanisms of pulmonary carcinogenesis. Mutat. Res. 823, 28–44 (2017).
Busch, W., Kühnel, D., Schirmer, K. & Scholz, S. Tungsten carbide cobalt nanoparticles exert hypoxia-like effects on the gene expression level in human keratinocytes. BMC Genomics 11, 65 (2010).
Tuomela, S. et al. Gene expression profiling of immune-competent human cells exposed to engineered zinc oxide or titanium dioxide nanoparticles. PLoS One 8, e68415 (2013).
Martin, S. L. et al. Effect of MWCNT on Lung Fibroblast gene expression. NCBI GEO (2013).
Osmond-McLeod, M. J. et al. Surface coatings of ZnO nanoparticles mitigate differentially a host of transcriptional, protein and signalling responses in primary human olfactory cells. Part Fibre Toxicol 10, 54 (2013).
Teeguarden, J. G. et al. Comparative iron oxide nanoparticle cellular dosimetry and response in mice by the inhalation and liquid cell culture exposure routes. Part Fibre Toxicol 11, 46 (2014).
Aday, S. et al. Inflammatory modulation of stem cells by Magnetic Resonance Imaging (MRI)-detectable nanoparticles. RSC Adv. 4, 31706–31709 (2014).
Fede, C. et al. Altered gene transcription in human cells treated with Ludox® silica nanoparticles. Int. J. Environ. Res. Public Health 11, 8867–8890 (2014).
Husain, M. et al. Intratracheally instilled titanium dioxide nanoparticles translocate to heart and liver and activate complement cascade in the heart of C57BL/6 mice. Nanotoxicology 9, 1013–1022 (2015).
Poulsen, S. S. et al. Changes in cholesterol homeostasis and acute phase response link pulmonary exposure to multi-walled carbon nanotubes to risk of cardiovascular disease. Toxicol. Appl. Pharmacol. 283, 210–222 (2015).
Husain, M. et al. Carbon black nanoparticles induce biphasic gene expression changes associated with inflammatory responses in the lungs of C57BL/6 mice following a single intratracheal instillation. Toxicol. Appl. Pharmacol. 289, 573–588 (2015).
van der Zande, M. et al. Different responses of Caco-2 and MCF-7 cells to silver nanoparticles are based on highly similar mechanisms of action. Nanotoxicology 10, 1431–1441 (2016).
Kinaret, P. et al. Network analysis reveals similar transcriptomic responses to intrinsic properties of carbon nanomaterials in vitro and in vivo. ACS Nano 11, 3786–3796 (2017).
Lastra, R. O. et al. Protein binding effects of dopamine coated titanium dioxide shell nanoparticles. prnano 2, 393–438 (2019).
Scala, G. et al. Multi-omics analysis of ten carbon nanomaterials effects highlights cell type specific patterns of molecular regulation and adaptation. NanoImpact 11, 99–108 (2018).
Mohan, A. et al. Transcriptional survey of alveolar macrophages in a murine model of chronic granulomatous inflammation reveals common themes with human sarcoidosis. Am. J. Physiol. Lung Cell Mol. Physiol. 314, L617–L625 (2018).
Jang, J. & Choi, I. Increased interleukin-11 and stress genes in human endothelial and human bronchial epithelial cell lines by silver nanoparticles. NCBI GEO https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE103101 (2017).
Vis, B. et al. Non-Functionalized Ultrasmall Silica Nanoparticles Directly and Size-Selectively Activate T Cells. ACS Nano 12, 10843–10854 (2018).
Kawata, K., Osawa, M. & Okabe, S. In vitro toxicity of silver nanoparticles at noncytotoxic doses to HepG2 human hepatoma cells. Environ. Sci. Technol. 43, 6046–6051 (2009).
Peng, L. et al. Whole genome expression analysis reveals differential effects of TiO2 nanotubes on vascular cells. Nano Lett. 10, 143–148 (2010).
Halappanavar, S. et al. Pulmonary response to surface-coated nanotitanium dioxide particles includes induction of acute phase response genes, inflammatory cascades, and changes in microRNAs: a toxicogenomic study. Environ Mol Mutagen 52, 425–439 (2011).
Eom, H.-J., Chatterjee, N., Lee, J. & Choi, J. Integrated mRNA and micro RNA profiling reveals epigenetic mechanism of differential sensitivity of Jurkat T cells to AgNPs and Ag ions. Toxicol. Lett. 229, 311–318 (2014).
Fabbro, A. et al. Adhesion to carbon nanotube conductive scaffolds forces action-potential appearance in immature rat spinal neurons. PLoS One 8, e73621 (2013).
Langley, R. J. et al. Fibrogenic and redox-related but not proinflammatory genes are upregulated in Lewis rat model of chronic silicosis. J Toxicol Environ Health Part A 74, 1261–1279 (2011).
Kodali, V. et al. Dysregulation of macrophage activation profiles by engineered nanoparticles. ACS Nano 7, 6997–7010 (2013).
Hussien, R. et al. Unique growth pattern of human mammary epithelial cells induced by polymeric nanoparticles. Physiol. Rep. 1, e00027 (2013).
Karoly, E. D., Li, Z., Dailey, L. A., Hyseni, X. & Huang, Y.-C. T. Up-regulation of tissue factor in human pulmonary artery endothelial cells after ultrafine particle exposure. Environ. Health Perspect. 115, 535–540 (2007).
Chorley, B., Ward, W., Simmons, S. O., Vallanat, B. & Veronesi, B. The cellular and genomic response of rat dopaminergic neurons (N27) to coated nanosilver. Neurotoxicology 45, 12–21 (2014).
Rydman, E. M. et al. Inhalation of rod-like carbon nanotubes causes unconventional allergic airway inflammation. Part Fibre Toxicol 11, 48 (2014).
Chernova, T. et al. Long-Fiber Carbon Nanotubes Replicate Asbestos-Induced Mesothelioma with Disruption of the Tumor Suppressor Gene Cdkn2a (Ink4a/Arf). Curr. Biol. 27, 3302–3314.e6 (2017).
Huang, Y.-C. T. et al. Comparison of gene expression profiles induced by coarse, fine, and ultrafine particulate matter. J Toxicol Environ Health Part A 74, 296–312 (2011).
Gao, X., Topping, V. D., Keltner, Z., Sprando, R. L. & Yourick, J. J. Toxicity of nano- and ionic silver to embryonic stem cells: a comparative toxicogenomic study. J. Nanobiotechnology 15, 31 (2017).
Guo, C. et al. Amorphous silica nanoparticles induce malignant transformation and tumorigenesis of human lung epithelial cells via P53 signaling. Nanotoxicology 11, 1176–1194 (2017).
Kinaret, P. et al. Inhalation and Oropharyngeal Aspiration Exposure to Rod-Like Carbon Nanotubes Induce Similar Airway Inflammation and Biological Responses in Mouse Lungs. ACS Nano 11, 291–303 (2017).
Proquin, H. et al. Time course gene expression data in colon of mice after exposure to food-grade E171. Data Brief 16, 531–600 (2018).
Kalmodia, S. et al. Characterization and Molecular Mechanism of Peptide-Conjugated Gold Nanoparticle Inhibiting p53-HDM2 Interaction in Retinoblastoma. Mol. Ther. Nucleic Acids 9, 349–364 (2017).
Li, M. et al. Hydroxylated-Graphene Quantum Dots Induce DNA Damage and Disrupt Microtubule Structure in Human Esophageal Epithelial Cells. Toxicol. Sci. 164, 339–352 (2018).
Orecchioni, M. et al. Single-cell mass cytometry and transcriptome profiling reveal the impact of graphene on human immune cells. Nat. Commun. 8, 1109 (2017).
Kang, M. H. et al. Near-infrared-emitting nanoparticles activate collagen synthesis via TGFβ signaling. Sci. Rep. 10, 13309 (2020).
House, J. S. et al. Low-Dose Silver Nanoparticle Surface Chemistry and Temporal Effects on Gene Expression in Human Liver Cells. Small 16, e2000299 (2020).
Yazdimamaghani, M., Moos, P. J. & Ghandehari, H. Global gene expression analysis of macrophage response induced by nonporous and porous silica nanoparticles. Nanomedicine 14, 533–545 (2018).
Wu, B. et al. Nano genome altas (NGA) of body wide organ responses. Biomaterials 205, 38–49 (2019).
Yang, D. & Landry, M. P. Transcriptomic response of SIM-A9 microglia to neuro-probes. NCBI GEO https://identifiers.org/geo:GSE153419 (2020).
Frank, E. A. et al. Genetic susceptibility to toxicologic lung responses among inbred mouse strains following exposure to carbon nanotubes and profiling of underlying gene networks. Toxicol. Appl. Pharmacol. 327, 59–70 (2017).
Acknowledgements
We thank the partners in the NanoSolveIT consortium for their valuable help in identifying data sets for the collection. This work was supported by the EU H2020 project NanoSolveIT (grant agreement 814572) as well as the Academy of Finland (grant agreement 322761).
Author information
Authors and Affiliations
Contributions
L.A.S. collected and organized the data, preprocessed and analyzed the microarray data and co-authored the manuscript. A.F. preprocessed and analyzed the RNA-Seq data and co-authored the manuscript. I.L. collected the ENM characteristics data and co-authored the manuscript. A.G.P. co-authored the manuscript and curated the data for the NanoPharos database. A.T. co-authored the manuscript and further developed the NanoPharos database to cover omics data. G.M. & A.A. co-authored the manuscript, organized the curation of the data and the development of NanoPharos database to cover omics data. A.S. defined the methodology, contributed to supervising the study and co-authored the manuscript. D.G. conceived and supervised the study and co-authored the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Online-only Tables
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files associated with this article.
About this article
Cite this article
Saarimäki, L.A., Federico, A., Lynch, I. et al. Manually curated transcriptomics data collection for toxicogenomic assessment of engineered nanomaterials. Sci Data 8, 49 (2021). https://doi.org/10.1038/s41597-021-00808-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-021-00808-y
This article is cited by
-
PROTEOMAS: a workflow enabling harmonized proteomic meta-analysis and proteomic signature mapping
Journal of Cheminformatics (2023)
-
An ancestral molecular response to nanomaterial particulates
Nature Nanotechnology (2023)
-
Prospects and challenges for FAIR toxicogenomics data
Nature Nanotechnology (2022)
-
Reply to: Prospects and challenges for FAIR toxicogenomics data
Nature Nanotechnology (2022)