Abstract
Plant traits are critical to plant form and function —including growth, survival and reproduction— and therefore shape fundamental aspects of population and ecosystem dynamics as well as ecosystem services. Here, we present a global species-level compilation of key functional traits for palms (Arecaceae), a plant family with keystone importance in tropical and subtropical ecosystems. We derived measurements of essential functional traits for all (>2500) palm species from key sources such as monographs, books, other scientific publications, as well as herbarium collections. This includes traits related to growth form, stems, armature, leaves and fruits. Although many species are still lacking trait information, the standardized and global coverage of the data set will be important for supporting future studies in tropical ecology, rainforest evolution, paleoecology, biogeography, macroecology, macroevolution, global change biology and conservation. Potential uses are comparative eco-evolutionary studies, ecological research on community dynamics, plant-animal interactions and ecosystem functioning, studies on plant-based ecosystem services, as well as conservation science concerned with the loss and restoration of functional diversity in a changing world.
Measurement(s) | fruit length • stem length • plant height • petiole length • fruit color • leaf length • plant trait • fruit morphology trait |
Technology Type(s) | digital curation |
Factor Type(s) | species |
Sample Characteristic - Organism | Arecaceae |
Sample Characteristic - Environment | tropical lowland evergreen broadleaf rain forest • tropical moist broadleaf forest biome • tropical deciduous broadleaf forest • tropical woodland biome • subtropical woodland biome |
Sample Characteristic - Location | global |
Machine-accessible metadata file describing the reported data: https://doi.org/10.6084/m9.figshare.9766919
Similar content being viewed by others
Background & Summary
Most ecosystems are composed of a large number of species with different characteristics. These characteristics (i.e. traits) reflect morphological, reproductive, physiological, phenological, or behavioural measurements of species that are usually collected to study intraspecific trait variation (i.e. among individuals or populations of the same species) or interspecific trait variation (i.e. among species)1,2,3,4,5. Many traits have an important functional role for species and ecosystems and are therefore referred to as ‘functional traits’. For instance, functional traits such as plant morphological and physiological properties are often directly linked to ecosystem structure and ecosystem functioning6,7. Such functional traits are further important for the response of organisms to their environment (‘response traits’) and the effects of organisms on ecosystems and other species (‘effect traits’)2,6,8. Hence, functional traits are key to understanding ecosystem dynamics and the response of organisms to human-induced disturbances and changing environmental conditions such as climate change4,9,10, habitat fragmentation11 or harvesting pressure12.
Over the last few years, comprehensive trait databases with continental or global scope have become available, covering diverse taxa in the marine13,14 and freshwater realm15 as well as terrestrial taxa such as plants16 and vertebrates17,18,19,20. Despite these monumental efforts that have involved community contributions as well as advanced techniques in data mining and data integration, digitally available information on functional traits is still missing for the majority of taxa on Earth3,21. Even for well-studied organisms such as vascular plants, information remains taxonomically and geographically limited. For instance, the TRY plant trait database16 has achieved an impressive compilation of almost 12 million trait records for currently 280,000 plant species (TRY database version 5 released in March 2019, www.try-db.org), but often only a few trait records are available per species. Moreover, as for other ecological information such as species occurrences22, digitally accessible information on traits remains particularly scarce in the tropics where most biodiversity occurs16,23,24,25. This is a major bottleneck for ecological and evolutionary science because tropical ecosystems such as rainforests are one of Earth’s greatest biological treasures, a major source of ecosystem services for a large proportion of the global human population, and a key component of the Earth system26,27.
In the tropics, palms are an iconic plant family with keystone importance in many forest and savanna ecosystems28,29,30. The pantropical palm family (Arecaceae or Palmae) is species-rich and contains nearly 2600 species in 181 genera and 5 subfamilies31. The ecology and evolution of palms is strongly linked to interspecific variation in growth, reproduction and morphology of stems, leaves, inflorescences, fruits and seeds32. Palms are a major resource for herbivores, pollinators and fruit- as well as seed-eating animals in the tropics29,30,32,33,34, provide provisioning services such as food, construction material and medicine to people (especially in rural communities)35, and belong to one of the most economically important plant groups globally36. Moreover, palms can provide important insights into the evolution of tropical rainforests28,37,38,39, historical biogeography40,41,42, past climate change43,44,45 and the vulnerability and response of ecosystems to ongoing and future global change46,47,48. Despite this outstanding role of palms in tropical ecosystems and tropical biological science, studies using palm functional trait data across broad spatial scales remain scarce35,38,49,50.
Here, we introduce the PalmTraits 1.0 database, an extensive database containing functional traits for palm species globally. PalmTraits 1.0 releases information on error-checked and referenced traits to capture interspecific variation in growth forms, armature and the morphology of stems, leaves and fruits of palms. Species-level trait information was assembled from >130 sources including monographs and taxonomic revisions as well as credible online sources and two herbaria with extensive palm collections. By making these data available to the scientific community, we aim to advance the sharing and digitalization of ecological trait data and understanding of the global ecology, biogeography and evolution of palms and the tropical rainforests they inhabit.
Methods
The data collection of the PalmTraits 1.0 database involved three major steps (Fig. 1a–c): (1) the identification of data sources, (2) the digitalization and encoding of trait values, and (3) the harmonization of fruit size data. Overall, the database was designed to capture species-level (interspecific) trait variation of palms rather than individual-level (intraspecific) variability. Such aggregated data (e.g. average values of continuous traits) facilitate biodiversity data integration across large spatial, temporal, and taxonomic scales, but are limited in their capacity to resolve fine-grained ecological patterns51. The PalmTraits 1.0 database captures trait variation of palms in terms of growth forms, stems, armature, leaves, and fruits (Online-only Table 1). This represents a large variation of trait diversity in palms (Fig. 1d–f). Some fundamental traits like wood density, specific leaf area or N-content5 are not represented because these traits are not commonly measured by palm taxonomists and hence are not available from palm books, monographs, species descriptions or herbarium specimens. Nevertheless, some of the available traits reflect major dimensions of plant form and function5, including the size of whole plants (e.g. growth form and stem height) and their organs (e.g. blade length). Other traits also capture characteristics that are relevant for studying plant-animal interactions such as herbivory (e.g. stem and leaf armature)48 and frugivory and animal-mediated seed dispersal (e.g. fruit length and width, fruit shape, and fruit colour)37,50,52. Below we describe the data collection (Fig. 1a–c) in more detail.
Trait compilation and trait variation in palms. The workflow (a–c) illustrates key steps in the compilation of palm trait data whereas the images (d–f) represent examples of trait variation in palms. (a) Main data sources for extracting trait data for PalmTraits 1.0. (b) Digitalization of the original trait information through different ways of encoding. (c) Standardization and harmonization of fruit trait information (size, shape and colour). (d) Palm growth forms (from left to right): Pritchardia viscosa, an erect canopy palm from Hawaii. Two erect palms (Thrinax radiata and Coccothrinax argentata) growing under a canopy gap in Mexico. Licuala telifera, an understory palm from New Guinea. Dypsis acaulis, an acaulescent understory palm from Madagascar. Plectocomiopsis geminiflora, a climbing rattan palm from Borneo. (e) Stem and leaf variation (from left to right): Astrocaryum standleyanum from Colombia, armed with long black spines. Daemonorops didymophylla from Southeast Asia, a climbing rattan palm armed with spines on the petiole of the leaf. Ceroxylon quindiuense from Colombia, with about 60 m stem height the tallest non-climbing palm in the world. Marojejya darianii from Madagascar, a medium-sized tree palm with large leaves of up to 5 m bade length. Johannesteijsmannia magnifica from Malaysia, an acaulescent palm with up to 2 metres long leaf blades covered with fine white hairs. (f) Fruit variation (from left to right): Lemurophoenix halleuxii from Madagascar, a canopy palm with large (5 cm) chestnut-brown fruits that have corky warts. Ravenea dransfieldii from Madagascar, a mid-story palm with small (1.5–2 cm) orange fruits. Calyptrocalyx sp., representing a genus of predominantly understory palms that have mostly small (1–2 cm) bright red fruits. Hydriastele microspadix from New Guinea, a mid-story palm with small dark red fruits. Drymophloeus litigiosus from New Guinea, an understory palm with small (1 cm) yellow to red fruits. Areca ipot from the Philippines, with large (5 cm long) fruits that ripen from green through yellow to red. Image credits: J. Dransfield, H. Balslev, and W.J. Baker.
Data sources
The main data sources for extracting the palm trait data were books and monographs, scientific articles (e.g. taxonomic revisions and species descriptions), herbarium specimen and specialized websites (Fig. 1a). We first extracted trait data from books, monographs and taxonomic revisions because these contain trait descriptions in a standardized way and for major clades or specific regions. We started the trait data extraction by obtaining maximum values for stem height, stem diameter, leaf number and fruit diameter as well as binary information (yes/no) for acaulescence and stem clustering for about 850–1250 species from the appendix I of the palm ecology and evolution book of A. Henderson32. We then extracted additional information for continuous traits (minimum, maximum and average values) as well as binary or categorical traits from books that synthesised species-specific palm knowledge for particular countries or regions (e.g. Africa53,54,55,56, Americas57, Australia58, Brazil59, Colombia60, Costa Rica61, Ecuador62, Hawaii63, Indonesia64, Madagascar65,66,67, Malaysia68,69, Mascarene Islands70, New Caledonia71, Philippines72, Sabah73, Southern Asia74, Thailand75, Vietnam76). Additionally, we went through taxonomic revisions, monographs and other publications that provided trait data for specific taxonomic groups such as palm genera or tribes (e.g. Acrocomia77, Aiphanes78, Archontophoenix79, Areca80, Asterogyne81, Astrocaryum82,83, Attalea84, Bactris85,86, Balaka87, Borassodendron88, Butia89, Calyptrocalyx90, Calyptrogyne91, Calamus92,93,94, Caryota95,96, Chamaedorea97,98,99,100,101,102,103, Cyrtostachys104, Drymophloeus105, Eremospatha55,106, Geonoma107, Heterospathe108, Hydriastele109,110, Hyospathe111, Johannesteijsmannia112, Laccosperma55,106, Lanonia113, Licuala114,115,116, Linospadix90,117, Livistona118,119, Metroxylon120, Nenga121, Oncocalamus55,106, Orania122, Parajubaea123, Phoenix124, Pinanga125, Ptychosperma126, Pritchardia127, Rhapis128, Sabal129, Syagrus130,131,132,133,134, Veitchia135, Wallichia136). We further obtained raw data (i.e. individual-level trait measurements) from A. Henderson that were used in taxonomic revisions for a number of palm genera, including Calyptrognye91, Chuniophoenix137, Desmoncus138, Geonoma107, Hyospathe139,140, Leopoldinia141, Pholidostachys142, Rhapis143, Synechanthus144 and Welfia145. These raw data allowed us to add a few additional trait data (especially minimum, mean and maximum fruit sizes) for 139 species. We additionally used other scientific literature on palms146,147,148,149,150,151,152,153,154,155,156,157,158,159,160,161,162,163,164,165,166,167,168,169,170,171,172,173,174 as well as specialized palm websites175,176,177,178,179, and the book Genera Palmarum180 for traits that do not vary within single genera (e.g. some genera have only climbers). Finally, we visited two major herbaria (Aarhus University Herbarium, Denmark, and the Royal Botanic Gardens, Kew, UK) harbouring very large palm collections to fill gaps in the database by obtaining trait information from herbarium specimens (e.g. measuring fruit sizes or recording fruit colour from specimen descriptions). All sources are provided together with the trait dataset in DRYAD181.
Digitalization
The digitalization of trait data from the original data sources into a database required to encode the information as continuous, binary, categorical or as text descriptions (Fig. 1b). For most traits, trait information was encoded either as continuous or binary (Online-only Table 1). For continuous traits, we usually recorded maximum values (e.g. for stem and leaf size) or minimum, maximum and average values (e.g. for fruit size) as reported in monographs and taxonomic revisions (Online-only Table 1). Binary traits were encoded as 0/1 (e.g. presence/absence of climbing, acaulescent or erect growth form, stem clustering, armature) and additionally as 2 if populations of the same species showed intraspecific trait variation (Online-only Table 1). Three traits were encoded as categorical information. This included understory/canopy information (a derived trait based on whether maximum stem height is ≤5 m or >5 m, and/or whether species have an acaulescent growth form or not)50, small/large fruit sizes (a derived trait based on whether fruit length is <4 cm or ≥4 cm in length, i.e. classifying small vs. megafaunal fruits)50,182, and fruit shape (Online-only Table 1). Two other traits (fruit colour and fruit shape) were encoded with text descriptions (Online-only Table 1). For those, we extracted verbatim text descriptions from the literature and herbarium sheets (e.g. glossy black, bright orange, or reddish brown as examples of fruit colour information) and later standardized and harmonized the information (see below).
Harmonization
Since our research has a particular focus on palm-frugivore interactions37,50,52,183, we further standardized and harmonized trait information on fruit size, fruit shape, and fruit colour (Fig. 1c).
For fruit size, the PalmTraits 1.0 database provides information on average, minimum and maximum values for both fruit length and fruit width (Online-only Table 1). However, in some monographs, species descriptions and taxonomic revisions the original information on fruit size was reported as fruit diameter rather than fruit length and fruit width. This typically included palm species that tend to have roundish fruits. We initially recorded these fruit diameter measurements in a separate column, but then merged it into the fruit length and/or fruit width columns. There was a measurement difference between fruit diameter compared to fruit width and length estimates for 168 palm species. For 74 of those 168 species, this difference was ≤0.1 cm and we therefore ignored (deleted) fruit diameter information. For the remaining 94 species, we revisited the original sources and additionally checked available online sources. In 82 cases, fruit diameter/width/length did not differ much (0.1–0.5 cm), and we updated the fruit width information based on the fruit diameter measurements. In the 12 remaining cases, fruit diameter values were much smaller or larger than fruit length (difference >0.5 cm), and we decided to omit these fruit diameter values to avoid biases and outliers.
For fruit shape, we harmonized the original trait descriptions from the literature into seven categories (ellipsoid, elongate, fusiform, globose, ovoid, pyramidal, and rounded) (Online-only Table 1). We chose those categories as they were most widely used. Note that these fruit shape descriptions are not necessarily distinctively different because no quantitative formulas are used when taxonomists describe the fruits.
For fruit colour, we kept the extracted verbatim text descriptions from the literature (‘FruitColorDescription’ in Online-only Table 1), but we additionally aggregated and harmonized the verbatim text descriptions in two ways. First, we derived the main fruit colour(s) from the verbatim text descriptions (‘MainFruitColors’ in Online-only Table 1) and separated them by semicolons (e.g. ‘black; blue’, or ‘brown; orange; yellow’). This allowed to keep the main fruit colour descriptions, but simplified and reduced the verbatim text. Immature fruit colours were excluded in this step, and fruit colours described with a suffix -ish or –ey were usually reduced to the main fruit colours. Second, we classified fruit colours into ‘cryptic’ and ‘conspicuous’ colours (‘Conspicuousness’ in Online-only Table 1). This was done because fruit-eating animals can differ in their colour vision, for instance birds vs. bats or dichromatic vs. trichromatic primates33. We classified fruit colours as cryptic if their reflectance spectra are difficult to detect against a background of leaves, and as conspicuous if reflectance spectra appear to be in strong contrast to the background of leaves184. Consequently, orange, red, yellow, pink, crimson and scarlet fruits were classified as conspicuous, and brown, black, green, blue, cream, grey, ivory, straw-coloured, white and purple fruits as cryptic (following ref.185). When a fruit colour description contained a combination of cryptic and conspicuous colours (e.g. ‘green/yellow’, ‘yellow-brown’, ‘brown orange’), or when colours were described with a suffix -ish or –ey (indicating to have only a touch of that colour), we inferred that the cryptic colour is the dominant hue and the fruit colour was classified as cryptic. The classification of cryptic vs. conspicuous is here provided as an example to show how the verbatim text descriptions of fruit colours could be used for ecological or evolutionary analyses, e.g. when analysing the colour vision of primates in relation to the distribution of palms with conspicuous fruit colours. Other colour classifications can be developed from the colour verbatim text descriptions as originally extracted from the data sources (column ‘FruitColorDescription’, Online-only Table 1).
Taxonomy
To standardize the taxonomic names of palms, we followed the World Checklist of palms186, using a version download from July 2015. This included a total of 2557 accepted palm species names. Since the palm taxonomy is regularly updated by taxonomic experts from the Royal Botanic Gardens in Kew, we recommend to use their taxonomic resources to search for synonyms and currently accepted names. Two useful online resources are the World Checklist of Selected Plant Families (WCSP, https://wcsp.science.kew.org, searching for ‘Arecaceae’) and PalmWeb (http://www.palmweb.org/).
Data Records
The PalmTraits 1.0 database can be downloaded from the DRYAD data repository181 under the terms of a Creative Commons Zero (CC0) waiver. The CC0 waiver facilitates the discovery, re-use, and interoperability of the data by removing any legal barriers. We also provide the PalmTraits 1.0 database in the TRY Plant Trait Database (https://www.try-db.org/; TRY DatasetID 540) which uses a Creative Commons Attribution License (CC BY 4.0). Regardless of the database version used, we ask users to cite this data paper when these data are used in publications or other activities (e.g. teaching and education), and to also cite the actual version of the database used in accord with emerging standards for data citation.
Data coverage
The PalmTraits 1.0 database covers 24 traits and additional taxonomic information (Online-only Table 1). The species coverage of trait information is complete for growth form (100% coverage), and particularly high for armature (>95%), stem habit (>84%), maximum stem height and diameter (>73%), and for average fruit size and width (>77%). Other traits are less covered (30–70%, Online-only Table 1), reflecting a lower availability of these traits in the published literature. Nevertheless, the high species coverage of several traits translates into a high geographic completeness of traits within country-level palm assemblages worldwide (Fig. 2, left column). For instance, global coverage of trait information is (near) complete for growth form and stem armature (Fig. 2, left top two maps). Other traits (e.g. maximum stem height, maximum blade length and average fruit length) have lower sampling completeness in species-rich tropical areas such as parts of South America, the Caribbean, Central Africa and Southeast Asia (Fig. 2, left bottom three maps).
Geographic variation of trait information for palm assemblages worldwide. In (a), geographic completeness of various palm trait data is shown at the spatial resolution of ‘botanical countries’ (TDWG level 3 units) which are standardized areas defined by the International Working Group on Taxonomic Databases (TDWG) for recording plant distributions193. Global palm distribution data from the World Checklist of Palms are available as presence-absence data in TDWG level 3 units186 and can be used to analyse the global distribution and biogeography of palm assemblages37,38,43,206. Geographic completeness is represented here as the proportion of species having trait information, with yellow showing botanical countries with high completeness and dark blue showing botanical countries with low completeness. In (b), interspecific variation of traits is shown for palm assemblages in botanical countries. Trait variation is exemplified as the proportion of species having a specific growth form (e.g. proportion of climbers), as the species richness of palm species with a particular binary trait (e.g. stem armature), or by representing the mean value of a continuous trait (e.g. maximum stem height, maximum blade length, or average fruit length) across all palm species in a given botanical country. Yellow indicates botanical countries with high trait values and dark blue low trait values.
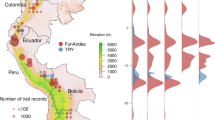
Mapping species-level trait information to a phylogeny allows visualizing the phylogenetic coverage of traits. Using a recently published all-evidence species-level supertree of palms187, we demonstrate that little phylogenetic bias exists in the coverage of key traits across the palm family (Fig. 3).
Phylogenetic distribution of exemplar palm traits. The five inner coloured circles represent species-level presence of trait information for key traits (growth form, maximum blade length, maximum stem height, average fruit length and stem armature). The outer coloured circle represents the five subfamilies of palms (Arecoideae, Ceroxyloideae, Coryphoideae, Nypoideae and Calamoideae). The time-calibrated phylogenetic tree illustrated here is a maximum clade credibility (MCC) tree50,52 derived from a recently published all-evidence species-level supertree of palms187.
Applications
The PalmTraits 1.0 allows the analysis of trait variation within palm species assemblages worldwide (Fig. 2, right column). This includes mapping the predominance (i.e. proportion) of particular growth forms (e.g. climbers), the species richness of palms with particular traits (e.g. stem armature), or the average size of stems, leaves or fruits across species that are present within botanical countries (Fig. 2, right column). Another avenue of application is to combine the species-level trait information with phylogenies (e.g. the recently published all-evidence species-level supertree of palms187) to perform macroevolutionary analyses such as trait-dependent models of speciation, extinction and transition rates50,52.
Technical Validation
All data were digitized by entering trait information from the original source (e.g. books, taxonomic revisions or specimen sheets) into an Excel spreadsheet, where each row represented a palm species and each column a single trait. Subsequent error detection and data quality control were done at three levels. First, trait information on growth forms (climbing, acaulescent, and erect) was carefully checked by a taxonomist (J.D.) with comprehensive experience with palms in the field and herbarium. Trait information of some specific palm genera was further checked by additional experts (see acknowledgements). Second, we sorted and filtered the Excel spreadsheet to search for erroneous entries (i.e. obvious errors in data entry) such as text or comma entries in columns with continuous data, or negative trait values and wrong values from unit conversion. These were corrected as much as possible. Third, we identified extreme values and detected outliers by looking at the most extreme (smallest and largest) values of each continuous trait across the whole family as well as within each tribe. These extreme values were checked for plausibility and reliability against external sources and our taxonomic and ecological knowledge of palms, and retained or corrected accordingly. For instance, several climbing palms (especially in the genus Calamus) have stem heights ≥100 m, with Calamus manan being the tallest climbing palm in the world (with 170 m stem height or more)188. Among erect palms, Ceroxylon quindiuense is with >60 m the tallest189. Fruit size is largest for Lodoicea maldivica (50 cm), the palm with the largest seed within the whole plant kingdom190. In contrast, the smallest fruit sizes are found in palm species in the genus Coccothrinax191. Palms also hold the record of the largest leaf of the plant kingdom, with Raphia regalis having a maximum blade length of 25 m192.
Usage Notes
We provide the data via the Dryad digital repository181 and via the TRY plant trait database (www.try-db.org; TRY DatasetID 540). The Dryad release181 contains three files related to the PalmTraits 1.0 database:
-
1.
A tab-delimited text file containing taxonomic information (species, genus, tribe, subfamily) together with all trait data
-
2.
A tab-delimited text file containing all references that have been used for each species.
-
3.
A tab-delimited text file containing the full details of all references that were used.
To facilitate integration with other datasets, we further provide the following files (also via the Dryad data repository181):
-
4.
An R script containing code that allows to combine the PalmTraits 1.0 database with species distribution and phylogenetic information
-
5.
A shape file with all botanical countries (TDWG level 3 units) worldwide
-
6.
Presence-absence data of palms at the resolution of botanical countries
-
7.
Phylogenetic information of palms represented as maximum clade credibility (MCC) tree
Tips for integrating the data records with other datasets
The R script that we provide contains guidance of how to integrate the PalmTraits 1.0 database with spatial and phylogenetic data and how to explore multi-variate trait variation181. We illustrate this by using the growth form information (climbing, acaulescent, and erect) from PalmTraits 1.0 (Fig. 4). We first load global species distribution data from the world checklist of palms186 and then combine them with the new palm growth form data and a polygon file that represents geographic units (‘botanical countries’, i.e. TDWG level 3 units) as defined by the International Working Group on Taxonomic Databases (TDWG), a geographic standard for recording plant distributions193. This allows plotting the proportion of growth forms in palm assemblages worldwide (Fig. 4a). We then map growth form information onto a species-level palm phylogeny187 using a Maximum Clade Credibility (MCC) phylogenetic tree as recently used in macroevolutionary analyses of palms50,52. This allows to explore growth form information in a phylogenetic context (Fig. 4b). Finally, the R script illustrates how continuous trait information (e.g. on stem height, leaf size and fruit size) can be combined with growth form information to explore the multi-dimensional nature of species traits (Fig. 4c).
Examples of combining palm trait data, species distribution and phylogenetic information. The global map in (a) shows the relative proportion of major palm growth forms within ‘botanical countries’ worldwide (i.e. geographic units as defined by the International Working Group on Taxonomic Databases, TDWG193) by combining growth form information (climbing, acaulescent, and erect) with global species distribution data from the world checklist of palms186. In (b), palm growth form information is linked to a species-level palm phylogeny187 using a Maximum Clade Credibility (MCC) phylogenetic tree50,52 to illustrate the phylogenetic distribution of climbing, acaulescence and erect growth forms in palms. Climbing dominates in the subfamily Calamoideae whereas erect palms are common in subfamilies Coryphoideae, Ceroxyloideae and Arecoideae. Acaulescent palms are scattered across the palm phylogenetic tree. In (c), the location of different palm growth forms (climbing, acaulescent, and erect) in a multivariate trait space is illustrated by the first two axes of a Principal Component Analysis (PCA) based on continuous trait information on stem height (logStemHeight), leaf size (logBladeLength and logRachisLength) and fruit size of palms (logFruitLength and logFruitWidth). The figure can be reproduced with data and an R script that integrates the PalmTraits 1.0 database with spatial and phylogenetic data181.
Imputation of missing trait data
As trait values are often not available for all species (Online-only Table 1), we recommend to explore data imputation methods to fill information for missing data. Data imputation might be especially important for analyses where complete trait-based representation of all palm species is crucial. For instance, metrics of functional diversity194 can be systematically biased when trait data coverage is incomplete195,196 and gap-filling may allow to reduce errors when interpreting functional diversity patterns197. Data imputation may be performed in a variety of ways, for example through the leveraging of phylogenetic comparative models198, taxonomic hierarchies199, or machine learning algorithms200. The relative performance and accuracy of the methods will depend on completeness and interspecific and intraspecific variation of traits. For instance, if correlations among traits are not strong, predictions based on observed covariation in existing trait data should be used with caution. We suggest that data imputation methods should be rigorously tested and accompanied with comprehensive sensitivity analyses to assess their performance201.
Semantic integration with other plant trait data
Plant trait data are measured in a multitude of ways202, and this heterogeneity together with a lack of standards for acquiring, organizing and describing trait data makes their integration often difficult3,203,204. Trait data of palms are usually described in a standardized and systematic way within taxonomic descriptions and revisions. This makes the extraction of palm trait data relatively straightforward. However, many of the palm trait terms and measurements are not directly captured in semantic descriptions of plant traits such as the global handbook for standardised measurement of plant functional traits205 or the thesaurus of plant characteristics (TOP)203. During the collection of palm trait data, we did not harmonize the terminology of palm trait definitions with other plant trait terminologies because they were internally consistent (i.e. within the palm family). However, after finalizing the data collection we mapped the palm trait definitions to the TOP (see Online-only Table 1). Several palm traits are currently not represented in the TOP. For instance, fruit colour is currently not represented within the dispersule trait category of the TOP. Similarly, maximum number of leaves as well as armature on leaves and stems are currently not captured by the TOP. This highlights the need for further development of the TOP and other semantic resources to facilitate the integration of trait data from multiple sources3. Such efforts will also allow better interoperability and effectiveness of automated data exchange among different sources. We therefore urge the research community to further develop and harmonize existing plant traits terminologies and semantic relations.
Code Availability
The original data collection was done by entering trait information into an Excel spreadsheet (Microsoft Office 2013). No code is available for this step. The final PalmTraits 1.0 dataset was saved as tab-delimited text file181. Scripts to load the PalmTraits 1.0 dataset into R, to plot multi-variate trait variation and to combine it with phylogenetic and species distribution data are available in the accompanying dataset181. The scripts were developed in R version 3.5.0, and using the associated libraries as indicated in the scripts. There are no restrictions to use the provided code.
References
Violle, C. et al. Let the concept of trait be functional! Oikos 116, 882–892, https://doi.org/10.1111/j.0030-1299.2007.15559.x (2007).
Díaz, S. et al. Functional traits, the phylogeny of function, and ecosystem service vulnerability. Ecology and Evolution 3, 2958–2975, https://doi.org/10.1002/ece3.601 (2013).
Kissling, W. D. et al. Towards global data products of Essential Biodiversity Variables on species traits. Nature Ecology & Evolution 2, 1531–1540, https://doi.org/10.1038/s41559-018-0667-3 (2018).
Bjorkman, A. D. et al. Plant functional trait change across a warming tundra biome. Nature 562, 57–62, https://doi.org/10.1038/s41586-018-0563-7 (2018).
Díaz, S. et al. The global spectrum of plant form and function. Nature 529, 167–171, https://doi.org/10.1038/nature16489 (2016).
Lavorel, S. & Garnier, E. Predicting changes in community composition and ecosystem functioning from plant traits: revisiting the Holy Grail. Functional Ecology 16, 545–556 (2002).
Schneider, F. D. et al. Mapping functional diversity from remotely sensed morphological and physiological forest traits. Nature Communications 8, 1441, https://doi.org/10.1038/s41467-017-01530-3 (2017).
Lavorel, S. et al. A novel framework for linking functional diversity of plants with other trophic levels for the quantification of ecosystem services. Journal of Vegetation Science 24, 942–948, https://doi.org/10.1111/jvs.12083 (2013).
Eskildsen, A. et al. Ecological specialization matters: long-term trends in butterfly species richness and assemblage composition depend on multiple functional traits. Diversity and Distributions 21, 792–802, https://doi.org/10.1111/ddi.12340 (2015).
Pacifici, M. et al. Species’ traits influenced their response to recent climate change. Nature Climate Change 7, 205–208, https://doi.org/10.1038/nclimate3223 (2017).
Hagen, M. et al. Biodiversity, species interactions and ecological networks in a fragmented world. Advances in Ecological Research 46, 89–210 (2012).
Genner, M. J. et al. Body size-dependent responses of a marine fish assemblage to climate change and fishing over a century-long scale. Global Change Biology 16, 517–527, https://doi.org/10.1111/j.1365-2486.2009.02027.x (2010).
Costello, M. J. et al. Biological and ecological traits of marine species. PeerJ 3, e1201 (2015).
Madin, J. S. et al. The Coral Trait Database, a curated database of trait information for coral species from the global oceans. Scientific. Data 3, 160017, https://doi.org/10.1038/sdata.2016.17 (2016).
Schmidt-Kloiber, A. & Hering, D. www.freshwaterecology.info – An online tool that unifies, standardises and codifies more than 20,000 European freshwater organisms and their ecological preferences. Ecological Indicators 53, 271–282, https://doi.org/10.1016/j.ecolind.2015.02.007 (2015).
Kattge, J. et al. TRY – a global database of plant traits. Global Change Biology 17, 2905–2935, https://doi.org/10.1111/j.1365-2486.2011.02451.x (2011).
Wilman, H. et al. EltonTraits 1.0: Species-level foraging attributes of the world’s birds and mammals. Ecology 95, 2027–2027, https://doi.org/10.1890/13-1917.1 (2014).
Kissling, W. D. et al. Establishing macroecological trait datasets: Digitalization, extrapolation, and validation of diet preferences in terrestrial mammals worldwide. Ecology and Evolution 4, 2913–2930, https://doi.org/10.1002/ece3.1136 (2014).
Oliveira, B. F., São-Pedro, V. A., Santos-Barrera, G., Penone, C. & Costa, G. C. AmphiBIO, a global database for amphibian ecological traits 4, 170123, https://doi.org/10.1038/sdata.2017.123 (2017).
Guralnick, R. P. et al. The importance of digitized biocollections as a source of trait data and a new VertNet resource. Database 2016, baw158, https://doi.org/10.1093/database/baw158 (2016).
Hortal, J. et al. Seven shortfalls that beset large-scale knowledge of biodiversity. Annual Review of Ecology, Evolution, and Systematics 46, 523–549, https://doi.org/10.1146/annurev-ecolsys-112414-054400 (2015).
Meyer, C., Kreft, H., Guralnick, R. & Jetz, W. Global priorities for an effective information basis of biodiversity distributions. Nature Communications 6, 8221, https://doi.org/10.1038/ncomms9221 (2015).
Bruelheide, H. et al. Global trait–environment relationships of plant communities. Nature Ecology & Evolution 2, 1906–1917, https://doi.org/10.1038/s41559-018-0699-8 (2018).
Salguero-Gómez, R. et al. COMADRE: a global data base of animal demography. Journal of Animal Ecology 85, 371–384, https://doi.org/10.1111/1365-2656.12482 (2016).
Salguero-Gómez, R. et al. The COMPADRE Plant Matrix Database: an open online repository for plant demography. Journal of Ecology 103, 202–218, https://doi.org/10.1111/1365-2745.12334 (2015).
Malhi, Y. et al. Climate change, deforestation, and the fate of the Amazon. Science 319, 169–172, https://doi.org/10.1126/science.1146961 (2008).
Richardson, J. E. & Pennington, R. T. Origin of tropical diversity: from clades to communities. (Frontiers Media, 2016).
Couvreur, T. L. P. & Baker, W. J. Tropical rain forest evolution: palms as a model group. BMC Biology 11, 48 (2013).
Eiserhardt, W. L., Svenning, J.-C., Kissling, W. D. & Balslev, H. Geographical ecology of the palms (Arecaceae): determinants of diversity and distributions across spatial scales. Annals of Botany 108, 1391–1416, https://doi.org/10.1093/aob/mcr146 (2011).
Zona, S. & Henderson, A. A review of animal mediated seed dispersal of palms. Selbyana 11, 6–21 (1989).
Baker, W. J. & Dransfield, J. Beyond Genera Palmarum: progress and prospects in palm systematics. Botanical Journal of the Linnean Society 182, 207–233, https://doi.org/10.1111/boj.12401 (2016).
Henderson, A. Evolution and ecology of palms. (The New York Botanical Garden Press, 2002).
Fleming, T. H. & Kress, W. J. The ornaments of life: coevolution and conservation in the tropics. (Chicago University Press, 2013).
Barfod, A. S., Hagen, M. & Borchsenius, F. Twenty-five years of progress in understanding pollination mechanisms in palms (Arecaceae). Annals of Botany 108, 1503–1516, https://doi.org/10.1093/aob/mcr192 (2011).
Cámara-Leret, R. et al. Fundamental species traits explain provisioning services of tropical American palms. Nature. Plants 3, 16220, https://doi.org/10.1038/nplants.2016.220 (2017).
Johnson, D. V. Tropical palms. (Food and Agriculture Organization of the United Nations, 2010).
Kissling, W. D. Has frugivory influenced the macroecology and diversification of a tropical keystone plant family? Research Ideas and Outcomes 3, e14944, https://doi.org/10.3897/rio.3.e14944 (2017).
Couvreur, T. L. P. et al. Global diversification of a tropical plant growth form: environmental correlates and historical contingencies in climbing palms. Frontiers in Genetics 5, 452, https://doi.org/10.3389/fgene.2014.00452 (2015).
Morley, R. J. Origin and evolution of tropical rain forests. (John Wiley & Sons, 2000).
Baker, W. J. & Couvreur, T. L. P. Global biogeography and diversification of palms sheds light on the evolution of tropical lineages. I. Historical biogeography. Journal of Biogeography 40, 274–285, https://doi.org/10.1111/j.1365-2699.2012.02795.x (2013).
Baker, W. J. & Couvreur, T. L. P. Global biogeography and diversification of palms sheds light on the evolution of tropical lineages. II. Diversification history and origin of regional assemblages. Journal of Biogeography 40, 286–298, https://doi.org/10.1111/j.1365-2699.2012.02794.x (2013).
Moore, H. E. The major groups of palms and their distribution. Gentes Herbarum 11, 27–140 (1973).
Kissling, W. D. et al. Quaternary and pre-Quaternary historical legacies in the global distribution of a major tropical plant lineage. Global Ecology and Biogeography 21, 909–921, https://doi.org/10.1111/j.1466-8238.2011.00728.x (2012).
Greenwood, D. R. & Wing, S. L. Eocene continental climates and latitudinal temperature gradients. Geology 23, 1044–1048 (1995).
Blach-Overgaard, A., Kissling, W. D., Dransfield, J., Balslev, H. & Svenning, J.-C. Multimillion-year climatic effects on palm species diversity in Africa. Ecology 94, 2426–2435, https://doi.org/10.1890/12-1577.1 (2013).
Blach-Overgaard, A., Balslev, H., Dransfield, J., Normand, S. & Svenning, J.-C. Global-change vulnerability of a key plant resource, the African palms. Scientific Reports 5, 12611, https://doi.org/10.1038/srep12611 (2015).
Galetti, M. et al. Functional extinction of birds drives rapid evolutionary changes in seed size. Science 340, 1086–1090, https://doi.org/10.1126/science.1233774 (2013).
Göldel, B., Araujo, A. C., Kissling, W. D. & Svenning, J.-C. Impacts of large herbivores on spinescence and abundance of palms in the Pantanal, Brazil. Botanical Journal of the Linnean Society 182, 465–479, https://doi.org/10.1111/boj.12420 (2016).
Göldel, B., Kissling, W. D. & Svenning, J.-C. Geographical variation and environmental correlates of functional trait distributions in palms (Arecaceae) across the New World. Botanical Journal of the Linnean Society 179, 602–617, https://doi.org/10.1111/boj.12349 (2015).
Onstein, R. E. et al. Frugivory-related traits promote speciation of tropical palms. Nature Ecology & Evolution 1, 1903–1911, https://doi.org/10.1038/s41559-017-0348-7 (2017).
König, C. et al. Biodiversity data integration–The significance of data resolution and domain. PLOS Biology 17, e3000183, https://doi.org/10.1371/journal.pbio.3000183 (2019).
Onstein, R. E. et al. To adapt or go extinct? The fate of megafaunal palm fruits under past global change. Proceedings of the Royal Society B: Biological Sciences 285, 20180882, https://doi.org/10.1098/rspb.2018.0882 (2018).
Dransfield, J. In Flora of tropical East Africa (ed. Polhill, R. M.) (A. A. Balkema, 1986).
Russell, T. A. In Flora of west tropical Africa Vol. 2 (ed. Hepper, F. N.) (Whitefriars Press, 1968).
Sunderland, T. C. H. Field guide to the rattans of Africa. (Royal Botanic Gardens, 2007).
Tuley, P. The palms of Africa. (The Trendrine Press, 1995).
Henderson, A., Galeano, G. & Bernal, R. Field guide to the palms of the Americas. (Princeton University Press, 1995).
Dowe, J. L. Australian palms: biogeography, ecology and systematics. (CSIRO Publishing, 2010).
Lorenzi, H. Brazilian Flora Arecaceae (Palms). (Instituto Plantarum, 2010).
Galeano, G. & Bernal, R. Palmas de Colombia: guia de campo. (Panamericana Formas e Impresos S.A., 2010).
Grayum, M. H. Nomenclatural and taxonomic notes on Costa Rican palms (Arecaceae), with five new species. Phytologia 84, 307–327 (1998).
Borchsenius, F., Pedersen, H. B. & Balslev, H. Manual to the palms of Ecuador. AAU Reports 37. (Aarhus University Press, 1998).
Read, R. & Hodel, D. R. In Manual of the flowering plants of Hawai’i. Bishop Museum Special Publication 83 (eds Wagner, W. L., Herbst, D. R. & Sohmer, S. H.) (B.P. Bishop Museum, 1990).
Mogea, J. P. In Palms for human needs in Asia (ed. Johnson, D.) (A.A. Balkema, 1991).
Dransfield, J. & Beentje, H. The palms of Madagascar. (Royal Botanic Gardens Kew and The International Palm Society, 1995).
Rakotoarinivo, M. & Dransfield, J. New species of Dypsis and Ravenea (Arecaceae) from Madagascar. Kew Bulletin 65, 279–303, https://doi.org/10.1007/s12225-010-9210-7 (2010).
Rakotoarinivo, M., Trudgen, M. S. & Baker, W. J. The palms of the Makira Protected Area, Madagascar. Palms 53, 125–146 (2009).
Dransfield, J. A manual of the rattans of the Malay Peninsula. Malayan Forest Records 29. (Forest Department, Ministry of Primary Industries, 1979).
Kiew, R. In Proceedings 1st International Sago Symposium (ed. Tan, K.) (Demajuan Kanji Sdn. Bhd., 1977).
Moore, H. E. & Guého, L. J. In Flore des Mascareignes (eds Bosser, J., Cadet, T., Guého, J. & Marais, W.) (The Sugar Industry Research Institute, l’Office de la Recherche Scientifique Outre-Mer & Royal Botanic Gardens, Kew, 1984).
Hodel, D. R. & Pintaud, J.-C. The palms of New Caledonia. (Kampon Tansacha, Nong Nooch Tropical Garden, 1998).
de Guzman, E. D. & Fernando, E. S. Guide to Philippine flora and fauna. (1986).
Dransfield, J. The rattans of Sabah. (Forest Department, 1984).
Henderson, A. Palms of Southern Asia. (Princeton University Press, 2009).
Hodel, D. R. The palms of Thailand. (Allen Press, 1998).
Henderson, A. & Dung, N. Q. Notes on rattans (Arecaceae) from Vietnam. Phytotaxa 8, 25–33 (2010).
Vianna, S. A., Berton, L. H. C., Pott, A., Guerreiro, S. M. C. & Colombo, C. A. Biometric characterization of fruits and morphoanatomy of the mesocarp of Acrocomia species (Arecaceae). International. Journal of Biology 9, 78–92, https://doi.org/10.5539/ijb.v9n3p78 (2017).
Borchsenius, F. & Bernal, R. Aiphanes (Palmae). Flora Neotropica 70, 1–94 (1996).
Dowe, J. L. & Hodel, D. R. A revision of Archontophoenix H.Wendl. & Drude (Arecaceae). Austrobaileya 4, 227–244 (1994).
Heatubun, C. D. Seven New Species of Areca (Arecaceae). Phytotaxa 28, 6–26 (2011).
Stauffer, F. W., Asmussen, C. B., Henderson, A. & Endress, P. K. A revision of Asterogyne (Arecaceae: Arecoideae: Geonomeae). Brittonia 55, 326–356, https://doi.org/10.1663/0007-196x(2003)055[0326:aroaaa]2.0.co;2 (2003).
Kahn, F. The genus Astrocaryum (Arecaceae). Revista Peruana de Biología 15, 31–48 (2008).
Kahn, E. & Ferreira, E. J. L. A new species of Astrocaryum (Palmae) from Acre, Brazil. Candollea 50, 321–328 (1995).
Pintaud, J.-C., Del Castillo, A. M. R., Ferreira, E. J. L. & R, M. M. Towards a revision of Attalea in Western Amazonia. Palms 60, 57–78 (2016).
Henderson, A. Bactris (Palmae). Flora Neotropica Monographs 79, 1–181 (2000).
Cascante, A. Additions to the Genus Bactris (Arecaceae) of Mesoamerica. Palms 44, 146–153 (2000).
Hodel, D. R. A synopsis of the genus Balaka. Palms 54, 161–188 (2010).
Dransfield, J. The genus Borassodendron in Malesia. Reinwardtia 8, 351–363 (1972).
Noblick, L. R. Butia: What we think we know about the genus. Journal of Oil Palm Research 208, 5–23 (2014).
Dowe, J. L. & Ferrero, M. D. Revision of Calyptrocalyx and the New Guinea species of Linospadix (Linospadicinae: Arecoideae: Arecaceae). Blumea - Biodiversity, Evolution and Biogeography of Plants 46, 207–251 (2001).
Henderson, A. A multivariate study of Calyptrogyne (Palmae). Systematic Botany 30, 60–83, https://doi.org/10.1600/0363644053661913 (2005).
Dransfield, J. & Baker, W. J. An account of the Papuasian species of Calamus (Arecaceae) with paired fruit. Kew Bulletin 58, 371–387, https://doi.org/10.2307/4120621 (2003).
Henderson, A., Ban, N. K. & Dung, N. Q. New species of Calamus (Palmae) from Vietnam. Palms 52, 187–197 (2008).
Jacob, J., Mohanan, N. & Kariyappa, K. C. A new species of Calamus L. (Arecaceae) from Silent Valley, the Western Ghats, India. Rheedea 18, 29–31 (2008).
Hahn, W. Biosystematics and evolution of the genus Caryota (Palmae: Arecoideae) PhD thesis, University of Wisconsin, (1993).
Jeanson, M. L., Yusuf, Z. & Labat, J.-N. A new species of Caryota (Arecaceae, Coryphoideae) from Central and North Sulawesi. Systematic Botany 36, 600–604 (2011).
Hodel, D. R. Chamaedorea palms: the species and their cultivation. (The International Palm Society, Allen Press, 1992).
Hodel, D. R. Two new species of Chamaedorea (Arecaceae). Novon 7, 35–37, https://doi.org/10.2307/3392071 (1997).
Hodel, D. R. Additons to Chamaedorea palms: new species from Mexico and Guatemala and miscellaneous notes. Principes 36, 188–202 (1992).
Hodel, D. R. Three new species of Chamaedorea from Panama. Principes 39, 14–20 (1995).
Hodel, D. R. Two new species of Chamaedorea from Costa Rica. Principes 40, 212–216 (1996).
Hodel, D. R., Herrera, G. & Cascante, A. A remarkable new species and additional novelties of Chamaedorea from Costa Rica and Panama. Palm. Journal 137, 32–44 (1997).
Hodel, D. R., Mont, J. J. C. & Zuniga, R. Two new species of Chamaedorea from Honduras. Principes 39, 183–189 (1995).
Heatubun, C. D. et al. A monograph of Cyrtostachys (Arecaceae). Kew Bulletin 64, 67–94, https://doi.org/10.1007/s12225-009-9096-4 (2009).
Zona, S. Revision of Drymophloeus (Areceacea: Arecoideae). Blumea - Biodiversity, Evolution and Biogeography of Plants 44, 1–24 (1999).
Sunderland, T. C. H. A taxonomic revision of the rattans of Africa (Arecaceae: Calamoideae). Phytotaxa 51, 1–76 (2012).
Henderson, A. A revision of Geonoma (Arecaceae). Phytotaxa 17, 1–271 (2011).
Trudgen, M. S. & Baker, W. J. A revision of the Heterospathe elegans (Arecaceae) complex in New Guinea. Kew Bulletin 63, 639–647 (2008).
Heatubun, C. D., Petoe, P. & Baker, W. J. A monograph of the Nengella group of Hydriastele (Arecaceae). Kew Bulletin 73, 18, https://doi.org/10.1007/s12225-018-9743-8 (2018).
Petoe, P., Cámara-Leret, R. & Baker, W. J. A monograph of the Hydriastele wendlandiana group (Arecaceae: Hydriastele). Kew Bulletin 73, 17, https://doi.org/10.1007/s12225-018-9736-7 (2018).
Skov, F. & Balslev, H. A revision of Hyospathe (Arecaceae). Nordic Journal of Botany 9, 189–202, https://doi.org/10.1111/j.1756-1051.1989.tb02114.x (1989).
Dransfield, J. The genus Johannesteijsmannia H.E. Moore Jr. Garden Bulletin Singapore 26, 63–83 (1972).
Henderson, A. J. & Bacon, C. D. Lanonia (Arecaceae: Palmae), a new genus from Asia, with a revision of the species. Systematic Botany 36, 883–895, https://doi.org/10.1600/036364411x604903 (2011).
Saw, L. G., Dransfield, J. & Keith-Lucas, D. M. Morphological diversity of the genus Licuala (Palmae). Telopea 10, 187–206 (2003).
Saw, L. G. A revision of Licuala (Palmae) in the Malay Peninsula. Sandakania 10, 1–95 (1997).
Saw, L. G. A revision of Licuala (Arecaceae, Coryphoideae) in Borneo. Kew Bulletin 67, 577–654, https://doi.org/10.1007/s12225-012-9414-0 (2012).
Dowe, J. L. & Irvine, A. K. A revision of Linospadix in Australia, with the description of a new species. Principes 41, 192–197 (1997).
Rodd, A. N. Revision of Livistona (Arecaceae) in Australia. Telopea 8, 49–153 (1998).
Dowe, J. L. A taxonomic account of Livistona R.Br. (Arecaceae). Garden’s Bulletin Singapore 60, 185–344 (2009).
Rauwerdink, J. B. An essay on Metroxylon, the sago palm. Principes 30, 165–180 (1986).
Fernando, E. S. A revision of the genus Nenga. Principes 27, 55–70 (1983).
Keim, A. P. & Dransfield, J. A monograph of the genus Orania (Arecaceae: Oranieae). Kew Bulletin 67, 127–190, https://doi.org/10.1007/s12225-012-9356-6 (2012).
Mónica, M. R. & Henderson, A. The genus Parajubaea (Palmae). Brittonia 42, 92–99, https://doi.org/10.2307/2807619 (1990).
Barrow, S. C. A monograph of Phoenix L. (Palmae: Coryphoideae). Kew Bulletin 53, 513–575, https://doi.org/10.2307/4110478 (1998).
Dransfield, J. Systematic notes on Pinanga (Palmae) in Borneo. Kew Bulletin 34, 769–788, https://doi.org/10.2307/4119070 (1980).
Essig, F. B. A revision of the genus Ptychosperma Labill. (Arecaceae). Allertonia 1, 415–478 (1978).
Hodel, D. R. A review of the genus Pritchardia. Palms 51, S1–S53 (2007).
Hastings, L. A revision of Rhapis, the lady palms. Palms 47, 62–78 (2003).
Zona, S. A monograph of Sabal (Arecaceae: Coryphoideae). Aliso 12, 583–666 (1990).
Glassman, S. F. Revisions of the palm genus Syagrus Mart. and other selected genera in the Cocos alliance. Illinois Biological Monographs 59, 1–414 (1987).
Noblick, L. R. A revision of the genus Syagrus (Arecaceae). Phytotaxa 294, 1–262 (2018).
Hodel, D. R. Hybrids in the genus Syagrus. Palms 55, 141–154 (2011).
Noblick, L. R. Syagrus × mirandana, a naturally occurring hybrid of S. coronata and S. microphylla. Palms 56, 57–60 (2012).
Noblick, L. R. & Lorenzi, H. New Syagrus species from Brazil. Palms 54, 18–42 (2010).
Moore, H. E. Synopses of various genera of Arecoideae: Veitchia. Gentes Herbarum 8, 483–536 (1957).
Henderson, A. A revision of Wallichia (Palmae). Taiwania 52, 1–11, https://doi.org/10.6165/tai.2007.52(1).1 (2007).
Henderson, A. A revision of Chuniophoenix (Arecaceae). Phytotaxa 218, 163–170 (2015).
Henderson, A. A revision of Desmoncus (Arecaceae). Phytotaxa 35, 1–88 (2011).
Henderson, A. J. A multivariate analysis of Hyospathe (Palmae). American Journal of Botany 91, 953–965 (2004).
Henderson, A. Hyospathe. Palms 48, 161–166 (2004).
Henderson, A. A revision of Leopoldinia (Arecaceae). Phytotaxa 32, 1–17 (2011).
Henderson, A. A revision of Pholidostachys (Arecaceae). Phytotaxa 43, 1–48, https://doi.org/10.11646/phytotaxa.43.1.1 (2012).
Henderson, A. A revision of Rhapis (Arecaceae). Phytotaxa 258, 137–152 (2016).
Henderson, A. & Ferreira, E. A morphometric study of Synechanthus (Palmae). Systematic Botany 27, 693–702 (2002).
Henderson, A. & Villalba, I. A revision of Welfia (Arecaceae). Phytotaxa 119, 33–44 (2013).
Aliaga-Rossel, E., Moraes, R. & Mamíferos, M. consumidores de frutas y semillas de la chonta (Astrocaryum gratum, Arecaceae) en bosques submontanos y aluviales de Bolivia. Ecología en Bolivia 49, 98–103 (2014).
Bacon, C. D. & Baker, W. J. Saribus resurrected. Palms 55, 109–116 (2011).
Balick, M. J., Anderson, A. B. & de Medeiros-Costa, J. T. Hybridization in the babassu palm complex. II. Attalea compta × Orbignya oleifera (Palmae). Brittonia 39, 26–36, https://doi.org/10.2307/2806969 (1987).
Barfod, A. S. A monographic study of the subfamily Phytelephantoideae (Arecaceae). Opera Botanica 105, 1–73 (1991).
Barfod, A. S. & Heatubun, C. D. Two new species of Licuala Thunb. (Arecaceae: Coryphoideae) from North Moluccas and Western New Guinea. Kew Bulletin 64, 553–557, https://doi.org/10.1007/s12225-009-9129-z (2009).
Bernal, R. Demography of the vegetable ivory palm Phytelephas seemannii in Colombia, and the impact of seed harvesting. Journal of Applied Ecology 35, 64–74, https://doi.org/10.1046/j.1365-2664.1998.00280.x (1998).
Bernal, R., Galeano, G. & Hodel, D. R. A new species of Chamaedorea from Columbia. Palms 48, 27–29 (2004).
Bernal, R. & Borchsenius, F. Taxonomic novelties in Aiphanes (Palmae) from Colombia and Venezuela. Caldasia 32, 117–127 (2010).
Dowe, J. L. & Ferrero, M. D. A new species of rheophytic palm from New Guinea. Palms 44, 194–197 (2000).
Dransfield, J. In Tropical trees as living systems (eds P. B. Tomlinson & M. H. Zimmermann) 232–246 (Cambridge University Press, 1978).
Dupuyoo, J.-M. Two palms with surprising qualities. Palms 50, 179–183 (2006).
Essig, F. B. & Hernandez, N. A systematic histological study of palm fruits. V. Subtribe Archontophoenicinae (Arecaceae). Brittonia 54, 65–71 (2002).
Essig, F. B. & Litten, L. A systematic histological analysis of palm fruits VII. The Cyrtostachydinae (Arecaceae). Brittonia 56, 375–379 (2004).
Essig, F. B. A systematic histological study of palm fruits. VI. Subtribe Linospadicinae (Arecaceae). Brittonia 54, 196–201 (2002).
Essig, F. B. A systematic histological study of palm fruits. VIII. Subtribe Dypsidinae (Arecaceae). Brittonia 60, 82–92 (2008).
Essig, F. B., Bussard, L. & Hernandez, N. A systematic histological study of palm fruits. IV. Subtribe Oncospermatinae (Arecaceae). Brittonia 53, 466–471 (2001).
Essig, F. B., Manka, T. J. & Bussard, L. A systematic histological study of palm fruits. III. Subtribe Iguanurinae (Arecaceae). Brittonia 51, 307–325 (1999).
Evans, T. & Sengdala, K. The Indochinese rattan Calamus acanthophyllus: a fire-loving palm. Palms 45, 25–28 (2001).
Fong, F. Studies on the population structure, growth dynamics and resource importance of nipa palm (Nypa fruticans Wurmb.) Ph.D. thesis, University of Malaya (1986).
Gibbons, M. & Spanner, T. W. Medemia argun lives! Principes 40, 65–74 (1996).
Henderson, A., Ban, N. K. & Thanh, B. V. New species of Areca, Pinanga, and Licuala (Arecaceae) from Vietnam. Phytotaxa 8, 34–40 (2010).
Henderson, A. Arecaceae part I. Introduction and the Iriarteinae. Flora Neotropica 53, 1–101 (1990).
Jones, D. L. Palms throughout the world. (Smithsonian Institution Press, 1995).
Keat, L. C. Notes on recent palm species and records from Peninsular Thailand. Principes 42, 110–119 (1998).
Kumagai, L. & Hanazaki, N. Ethnobotanical and ethnoecological study of Butia catarinensis Noblick & Lorenzi: contributions to the conservation of an endangered area in southern Brazil. Acta Botanica Brasilica 27, 13–20 (2013).
McClatchey, W. C. Phylogenetic analysis of morphological characters of Metroxylon section Coelococcus (Palmae) and resulting implications for studies of other Calamoideae genera. Memoirs of the New York Botanical Garden 83, 285–306 (1999).
Moraes, R. M. & Pintaud, J.-C. Attalea blepharopus Mart. (Arecaceae) from Bolivia revisited since Martius. Candollea 71, 27–32 (2016).
Skov, F. Geonoma polyandra (Arecaceae), a new species from Ecuador. Nordic Journal of Botany 14, 39–41, https://doi.org/10.1111/j.1756-1051.1994.tb00567.x (1994).
Soares, K. P., Longhi, S. J., Witeck Neto, L. & Assis, L. C. D. Palmeiras (Arecaceae) no Rio Grande do Sul, Brasil. Rodriguésia 65, 113–139 (2014).
Palmweb. Palmweb - palms of the world online, http://www.palmweb.org/ (2019).
Palmpedia. Palmpedia - palm grower’s guide, http://www.palmpedia.net (2019).
Fern, K. The useful tropical plants database, http://tropical.theferns.info/ (2014).
Couvreur, T. L. P. Palms of Africa, http://palms.myspecies.info/ (2019).
Virtual Herbarium. The Fairchild tropical botanic garden virtual herbarium (FTG), http://www.virtualherbarium.org/ (2011).
Dransfield, J. et al. Genera palmarum - the evolution and classification of palms. (Royal Botanical Gardens, 2008).
Kissling, W. D. et al. Data from: PalmTraits 1.0, a species-level functional trait database of palms worldwide. Dryad Digital Repository. https://doi.org/10.5061/dryad.ts45225 (2019).
Guimarães, P. R. Jr., Galetti, M. & Jordano, P. Seed dispersal anachronisms: rethinking the fruits extinct megafauna ate. PLoS ONE 3, e1745 (2008).
Muñoz, G., Trøjelsgaard, K. & Kissling, W. D. A synthesis of animal-mediated seed dispersal of palms reveals distinct biogeographical differences in species interactions. Journal of Biogeography 46, 466–484, https://doi.org/10.1111/jbi.13493 (2019).
Regan, B. C. et al. Fruits, foliage and the evolution of primate colour vision. Philosophical Transactions of the Royal Society of London. Series B: Biological Sciences 356, 229–283, https://doi.org/10.1098/rstb.2000.0773 (2001).
Dominy, N. J., Svenning, J. C. & Li, W. H. Historical contingency in the evolution of primate color vision. Journal of Human Evolution 44, 25–45, https://doi.org/10.1016/s0047-2484(02)00167-7 (2003).
Govaerts, R. & Dransfield, J. World checklist of palms. (Royal Botanic Gardens Kew, 2005).
Faurby, S., Eiserhardt, W. L., Baker, W. J. & Svenning, J.-C. An all-evidence species-level supertree for the palms (Arecaceae). Molecular Phylogenetics and Evolution 100, 57–69, https://doi.org/10.1016/j.ympev.2016.03.002 (2016).
Burkill, I. H. A dictionary of the economic products of the Malay Peninsula. (Ministry of Agriculture and Cooperatives, 1966).
Bernal, R., Martinez, B. & Sanin, M. J. The World’s tallest palms. Palms 62, 5–16 (2018).
Bailey, L. H. Palms of the Seychelles Islands. Gentes Herbarum 6, 3–48 (1942).
Craft, P. The palms of Cuba. (Palm Nut Pages, 2018).
Hallé, F. The longest leaf in palms? Principes 21, 18 (1977).
Brummitt, R. K. World Geographical Scheme for Recording Plant Distributions. 2nd edn, (Hunt Institute for Botanical Documentation Carnegie Mellon University, 2001).
Laliberté, E. & Legendre, P. A distance-based framework for measuring functional diversity from multiple traits. Ecology 91, 299–305, https://doi.org/10.1890/08-2244.1 (2010).
Pakeman, R. J. Functional trait metrics are sensitive to the completeness of the species’ trait data? Methods in Ecology and Evolution 5, 9–15, https://doi.org/10.1111/2041-210x.12136 (2014).
van der Plas, F., van Klink, R., Manning, P., Olff, H. & Fischer, M. Sensitivity of functional diversity metrics to sampling intensity. Methods in Ecology and Evolution 8, 1072–1080, https://doi.org/10.1111/2041-210x.12728 (2017).
Kim, S. W., Blomberg, S. P. & Pandolfi, J. M. Transcending data gaps: a framework to reduce inferential errors in ecological analyses. Ecology Letters 21, 1200–1210, https://doi.org/10.1111/ele.13089 (2018).
Swenson, N. G. Phylogenetic imputation of plant functional trait databases. Ecography 37, 105–110, https://doi.org/10.1111/j.1600-0587.2013.00528.x (2014).
Schrodt, F. et al. BHPMF – a hierarchical Bayesian approach to gap-filling and trait prediction for macroecology and functional biogeography. Global Ecology and Biogeography 24, 1510–1521, https://doi.org/10.1111/geb.12335 (2015).
Stekhoven, D. J. & Bühlmann, P. MissForest—non-parametric missing value imputation for mixed-type data. Bioinformatics 28, 112–118, https://doi.org/10.1093/bioinformatics/btr597 (2011).
Penone, C. et al. Imputation of missing data in life-history trait datasets: which approach performs the best? Methods in Ecology and Evolution 5, 961–970, https://doi.org/10.1111/2041-210x.12232 (2014).
Cornelissen, J. H. C. et al. A handbook of protocols for standardised and easy measurement of plant functional traits worldwide. Australian Journal of Botany 51, 335–380, https://doi.org/10.1071/BT02124 (2003).
Garnier, E. et al. Towards a thesaurus of plant characteristics: an ecological contribution. Journal of Ecology 105, 298–309, https://doi.org/10.1111/1365-2745.12698 (2017).
Parr, C. S. et al. TraitBank: Practical semantics for organism attribute data. Semantic Web 7, 577–588, https://doi.org/10.3233/SW-150190 (2016).
Pérez-Harguindeguy, N. et al. New handbook for standardised measurement of plant functional traits worldwide. Australian Journal of Botany 61, 167–234, https://doi.org/10.1071/BT12225 (2013).
Kissling, W. D. et al. Cenozoic imprints on the phylogenetic structure of palm species assemblages worldwide. Proceedings of the National Academy of Sciences of the United States of America 109, 7379–7384, https://doi.org/10.1073/pnas.1120467109 (2012).
Acknowledgements
We thank Birgitte Bergmann (Aarhus University) for help with data entry, and Andrew Henderson (New York Botanical Garden) for providing individual trait measurements for species from several palm genera (Calyptrognye, Chuniophoenix, Desmoncus, Geonoma, Hyospathe, Leopoldinia, Pholidostachys, Rhapis, Synechanthus, Welfia). We acknowledge additional cross-checking of trait information for specific palm genera by Lauren Gardiner (Heterospathe), Finn Borchsenius (Aiphanes, Bactris, Desmoncus, Geonoma), Anders Barfod (Licuala), Rodrigo Bernal (Syagrus hybrids) and Donald Hodel (some Chamaedorea species). W.D.K. was supported by the University of Amsterdam (starting grant), the Danish Council for Independent Research–Natural Sciences (grant 11-106163) and the Netherlands Organisation for Scientific Research (Grant 824.15.007), H.B. by the European Union 7th Framework Programme (FP7-PALMS-Contract no. 212631) and by the Danish National Science Research Council (272-06-0476), W.J.B. by a grant from the Garfield Weston Foundation to the Global Tree Seed Bank Project at the Royal Botanic Gardens, Kew, and J.C.S. by the European Research Council (ERC-2012-StG-310886-HISTFUNC). J.C.S. also considers this work a contribution to his VILLUM Investigator project ‘Biodiversity Dynamics in a Changing World’ funded by VILLUM FONDEN. R.E.O. acknowledges the support of the German Centre for Integrative Biodiversity Research (iDiv) Halle‐Jena‐Leipzig funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation)—FZT 118. R.E.O. further acknowledges SYNTHESYS grant GB-TAF-6695.
Author information
Authors and Affiliations
Contributions
W.D.K., H.B. and J.C.S. conceived the idea; W.D.K. designed the study; W.D.K., B.G., R.E.O., J.D., W.J.B. collected data and/or facilitated data collection; W.D.K., R.E.O. and J.Y.L. assembled the dataset and prepared figures and code; W.D.K. wrote the manuscript; all authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Online-only Table
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files associated with this article.
About this article
Cite this article
Kissling, W.D., Balslev, H., Baker, W.J. et al. PalmTraits 1.0, a species-level functional trait database of palms worldwide. Sci Data 6, 178 (2019). https://doi.org/10.1038/s41597-019-0189-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-019-0189-0
This article is cited by
-
Current insights into palm fungi with emphasis on taxonomy and phylogeny
Fungal Diversity (2024)
-
The AusTraits plant dictionary
Scientific Data (2024)
-
Specific leaf area (SLA) serves as a proxy to predict total carbon content in understory individuals of the neotropical canopy palm Socratea exorrhiza
Trees (2023)
-
The likely extinction of hundreds of palm species threatens their contributions to people and ecosystems
Nature Ecology & Evolution (2022)
-
Climate change impacts on the Copernicia alba and Copernicia prunifera (Arecaceae) distribution in South America
Brazilian Journal of Botany (2022)