Abstract
The nucleolus is a compartmentalized organelle in eukaryotic cells known to form during embryogenesis, yet how its layered architecture is transformed from homogenous precursor bodies is unclear, and any impacts of this formation on embryonic cell fate determination remain unknown. Here, we demonstrate that lncRNA LoNA tethers granular-component-enriched NPM1 to dense-fibrillar-component-enriched FBL and drives the formation of compartmentalized nucleolus via facilitating liquid–liquid phase separation of those two nucleolar proteins. Phenotypically, LoNA-deficient embryos show developmental arrest at the two-cell (2C) stage. Mechanistically, we demonstrate that LoNA deficiency leads to nucleolar formation failure, resulting in mislocalization and acetylation of NPM1 in the nucleoplasm. Acetylated NPM1 recruits and guides PRC2 complex to 2C genes, where PRC2 complex trimethylates H3K27, leading to transcriptional repression of these genes. Collectively, our findings reveal that lncRNA is required for the establishment of nucleolar structure, and this process has an impact on two-cell embryonic development via 2C transcriptional activation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
RNA-seq and ChIP–seq data generated in the present study have been deposited at the GEO under accession no. GSE166142. Source data are provided with this paper.
References
Koné, M. C. et al. Three-dimensional distribution of UBF and Nopp140 in relationship to ribosomal DNA transcription during mouse preimplantation development. Biol. Reprod. 94, 95 (2016).
Biggiogera, M. et al. Nucleolar distribution of proteins B23 and nucleolin in mouse preimplantation embryos as visualized by immunoelectron microscopy. Development 110, 1263–1270 (1990).
Kresoja-Rakic, J. & Santoro, R. Nucleolus and rRNA gene chromatin in early embryo development. Trends Genet. 35, 868–879 (2019).
Fulka, H. & Aoki, F. Nucleolus precursor bodies and ribosome biogenesis in early mammalian embryos: old theories and new discoveries. Biol. Reprod. 94, 143 (2016).
Bouckenheimer, J. et al. Long non-coding RNAs in human early embryonic development and their potential in ART. Hum. Reprod. Update 23, 19–40 (2016).
Carlevaro-Fita, J. & Johnson, R. Global positioning system: understanding long noncoding RNAs through subcellular localization. Mol. Cell 73, 869–883 (2019).
Yan, L. et al. Single-cell RNA-seq profiling of human preimplantation embryos and embryonic stem cells. Nat. Struct. Mol. Biol. 20, 1131–1139 (2013).
Li, D. F. et al. Activity dependent LoNA regulates translation by coordinating rRNA transcription and methylation. Nat. Commun. 9, 1726 (2018).
Macfarlan, T. S. et al. Embryonic stem cell potency fluctuates with endogenous retrovirus activity. Nature 487, 57–63 (2012).
Ishiuchi, T. et al. Early embryonic-like cells are induced by downregulating replication-dependent chromatin assembly. Nat. Struct. Mol. Biol. 22, 662–671 (2015).
Nachmani, D. et al. Germline NPM1 mutations lead to altered rRNA 2’-O-methylation and cause dyskeratosis congenita. Nat. Genet. 51, 1518–1529 (2019).
Frottin, F. et al. The nucleolus functions as a phase-separated protein quality control compartment. Science 365, 342–347 (2019).
Feric, M. et al. Coexisting liquid phases underlie nucleolar subcompartments. Cell 165, 1686–1697 (2016).
Leung, A. K. et al. Quantitative kinetic analysis of nucleolar breakdown and reassembly during mitosis in live human cells. J. Cell Biol. 166, 787–800 (2004).
Percharde, M. et al. A LINE1-nucleolin partnership regulates early development and ESC identity. Cell 174, 391–405 (2018).
Xia, W. et al. Resetting histone modifications during human parental-to-zygotic transition. Science 365, 353–360 (2019).
Shandilya, J. et al. Acetylated NPM1 localizes in the nucleoplasm and regulates transcriptional activation of genes implicated in oral cancer manifestation. Mol. Cell Biol. 29, 5115–5127 (2009).
Swaminathan, V., Kishore, A. H., Febitha, K. K. & Kundu, T. K. Human histone chaperone nucleophosmin enhances acetylation-dependent chromatin transcription. Mol. Cell Biol. 25, 7534–7545 (2005).
Federici, L. et al. Nucleophosmin C-terminal leukemia-associated domain interacts with G-rich quadruplex forming DNA. J. Biol. Chem. 285, 37138–37149 (2010).
Laugesen, A., Højfeldt, J. W. & Helin, K. Molecular mechanisms directing PRC2 recruitment and H3K27 methylation. Mol. Cell 74, 8–18 (2019).
Liu, X. et al. Distinct features of H3K4me3 and H3K27me3 chromatin domains in pre-implantation embryos. Nature 537, 558–562 (2016).
Vastenhouw, N. L. & Schier, A. F. Bivalent histone modifications in early embryogenesis. Curr. Opin. Cell Biol. 24, 374–386 (2012).
Margueron, R. & Reinberg, D. The Polycomb complex PRC2 and its mark in life. Nature 469, 343–349 (2011).
Cela, I., Di Matteo, A. & Federici, L. Nucleophosmin in Its Interaction with ligands. Int. J. Mol. Sci. 21, 4885 (2020).
Iarovaia, O. V. et al. Nucleolus: a central hub for nuclear functions. Trends Cell Biol. 29, 647–659 (2019).
Sauvageau, M. et al. Multiple knockout mouse models reveal lincRNAs are required for life and brain development. eLife 2, e01749 (2013).
Andersen, R. E. et al. The long noncoding RNA Pnky is a trans-acting regulator of cortical development in vivo. Dev. Cell 49, 632–642 (2019).
Grote, P. et al. The tissue-specific lncRNA Fendrr is an essential regulator of heart and body wall development in the mouse. Dev. Cell 24, 206–214 (2013).
Li, X. & Fu, X. D. Chromatin-associated RNAs as facilitators of functional genomic interactions. Nat. Rev. Genet. 20, 503–519 (2019).
Brangwynne, C. P., Mitchison, T. J. & Hyman, A. A. Active liquid-like behavior of nucleoli determines their size and shape in Xenopus laevis oocytes. Proc. Natl Acad. Sci. USA 108, 4334–4339 (2011).
Mitrea, D. M. et al. Nucleophosmin integrates within the nucleolus via multi-modal interactions with proteins displaying R-rich linear motifs and rRNA. eLife 5, e13571 (2016).
Mao, Y. S., Zhang, B. & Spector, D. L. Biogenesis and function of nuclear bodies. Trends Genet. 27, 295–306 (2011).
Chujo, T., Yamazaki, T. & Hirose, T. Architectural RNAs (arcRNAs): a class of long noncoding RNAs that function as the scaffold of nuclear bodies. Biochim. Biophys. Acta 1859, 139–146 (2016).
Quinn, J. J. & Chang, H. Y. Unique features of long non-coding RNA biogenesis and function. Nat. Rev. Genet. 17, 47–62 (2016).
Tripathi, V. et al. The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol. Cell 39, 925–938 (2010).
Chujo, T. & Hirose, T. Nuclear bodies built on architectural long noncoding RNAs: unifying principles of their construction and function. Mol. Cells 40, 889–896 (2017).
Xie, S. Q. et al. Nucleolar-based Dux repression is essential for embryonic two-cell stage exit. Genes Dev. 36, 331–347 (2022).
Yu, H. et al. rRNA biogenesis regulates mouse 2C-like state by 3D structure reorganization of peri-nucleolar heterochromatin. Nat. Commun. 12, 6365 (2021).
Li, X. et al. Astrocytic ApoE reprograms neuronal cholesterol metabolism and histone-acetylation-mediated memory. Neuron 109, 957–970 (2021).
Yin, Q. F. et al. SnoVectors for nuclear expression of RNA. Nucleic Acids Res. 43, e5 (2015).
Sztein, J. M., Farley, J. S. & Mobraaten, L. E. In vitro fertilization with cryopreserved inbred mouse sperm. Biol. Reprod. 63, 1774–1780 (2000).
Tang, A. H. et al. A trans-synaptic nanocolumn aligns neurotransmitter release to receptors. Nature 536, 210–214 (2016).
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M. & Weissman, J. S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protoc. 7, 1534–1550 (2012).
Zhang, Q. et al. Nuclear speckle specific hnRNP D-like prevents age- and AD-related cognitive decline by modulating RNA splicing. Mol. Neurodegener. 16, 66 (2021).
Gromava, J. E. in Cell Biology: A Laboratory Handbook 3rd edn (eds Celis, J. E. et al.) 421–429 (Academic Press, 2005).
Zhang, W., Xie, M., Shu, M. D., Steitz, J. A. & DiMaio, D. A proximity-dependent assay for specific RNA protein interactions in intact cells. RNA 22, 1785–1792 (2016).
Beischlag, T. V., Prefontaine, G. G. & Hankinson, O. ChIP–re-ChIP: co-occupancy analysis by sequential chromatin immunoprecipitation. Methods Mol. Biol. 1689, 103–112 (2018).
Lambrecht, E. et al. Transmission electron microscopy sample preparation protocols for the ultrastructural study of cysts of free-living protozoa. BioTechniques 58, 181–188 (2015).
Acknowledgements
This research was supported by the National Natural Science Foundation of China (82125009, 31871082, 91849101, 32121002, 82071185, 32100794 and 92149303), the National Key R&D Program of China (2020YFA0509300, 2021YFA0804900), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB39000000), CAS Project for Young Scientists in Basic Research (YSBR-013), the Key Research Program of Frontier Sciences of the Chinese Academy of Sciences (QYZDB-SSW-SMC035), Collaborative Innovation Program of Hefei Science Center, CAS (2022HSC-CIP003), the Anhui Provincial Natural Science Foundation (2008085QC117), the Fundamental Research Funds for the Central Universities (YD2070002011, WK2070000168), and Institute of Health and Medicine, Hefei Comprehensive National Science Center (QYZD20220003).
Author information
Authors and Affiliations
Contributions
Q. Liu, J.Z., and D.L. designed the study and wrote the manuscript, Q. Liu supervised the work. D.L. carried out most of the experiments, conducted data analysis, and designed the figures. Q. Li conducted IVF. A.T. and R.C. performed STORM imaging and data analysis. Y.Y. purified recombinant proteins.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Sara Osman was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 LoNA binds to NPM1 through three binding sites and this binding is required for nucleolar formation.
a, Schematic drawing of initiation of NPB maturation and LoNA’s expression pattern during transition from zygote to two-cell embryo. b, Upper: Representation of the 2C-EGFP reporter. Lower: Representative immunofluorescence images of OCT4 (red), EGFP (green), and DAPI (blue) in mESCs electroporated with 2C-EGFP reporter plasmid. Indicated EGFP positive and OCT4 negative cells were selected as 2C-like cells. Scale bar:10 μm. c, Representative images of 2C-like cells and quantification of EGFP positive and negative cells. Scale bar: 50 μm. d, LoNA levels in 2C-like cells and mESCs, by qPCR analysis (n = 3); two-sided paired t-test. e, Representative bright filed and fluorescence images of EGFP in control and LoNA KD 2C-like cells. Scale bar: 50 μm. f, Distribution of EGFP positive and negative cells in control and LoNA KD 2C-like cells. g, Silver staining of pull-down proteins by biotinylated LoNA antisense or control sense probes. h, Upper: Schematic drawing of full-length NPM1 and its three truncation fragments, with HA tagged on their C-terminus. Lower: HA tagged full-length and truncated NPM1s were electroporated to 2C-like cells, followed by immunoblotting with an anti-HA antibody. i, Binding affinities of LoNA to full-length NPM1 or NPM1 truncation fragments. j, Upper: Schematic drawing of LoNA-WT and its three NPM1 binding sites, LoNA mutants with point mutation(s) on one/two/three of the NPM1 binding sites. Lower: Binding affinities of LoNA mutants and NPM1. k, Densiometric analysis for (j) (n = 3); two-sided paired t-test; ***P < 0.001. l, Representative immunofluorescence images of NPM1 (red), LoNA RNA FISH (magenta), and DAPI (blue) in control and LoNA KD 2C-like cells. Scale bar: 2 μm. m, Representative immunofluorescence images of RPL27 (red), LoNA RNA FISH (magenta), and DAPI (blue) in control and LoNA KD 2C-like cells. Scale bar: 2 μm. n, Representative transmission electron microscopy (TEM) images of nucleolar structure in control and LoNA KD 2C-like cells and quantification of GC density in TEM images (n = 15 nucleoli); two-sided paired t-test; ***P < 0.001. Asterisks indicate the fibrillar centers. Scale bar: 500 nM. Data are presented as mean values ± s.e.m.
Extended Data Fig. 2 FBL-NPM1 protein interaction depends on LoNA-mediated tethering.
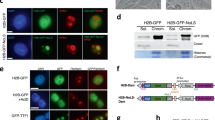
a-c, Control and LoNA KD 2C-like cells were electroporated with an equal amount of LoNA-FBL/NPM1-BD (a), LoNA-FBL-BD (b), or LoNA-NPM1-BD (c), then subjected to immunoprecipitation with an anti-FBL antibody and immunoblotting with an anti-NPM1 antibody. d,e, Control and LoNA KD 2C-like cells were introduced with an equal amount of LoNA-NPM1-BD (d) or LoNA-FBL/NPM1-BD (e), then subjected to immunoprecipitation with an anti-NPM1 antibody and immunoblotting with an anti-FBL antibody. f and h, 2C-like cell lysates treated with indicated amounts of RNAse A/T1 (f) or DNase I (h), followed by immunoprecipitation with an anti-FBL antibody and immunoblotting with an anti-NPM1 antibody. g and i, Densiometric analyses for (f) and (h) (n = 3); two-sided paired t-test; n.s. P > 0.05, ***P < 0.001. j, Representative images of LoNA and FBL PLA signals (magenta), NPM1 (green), and DAPI (blue) in control and LoNA KD 2C-like cells. Scale bar: 10 μm. k, Representative images of LoNA and NPM1 PLA signals (magenta), FBL (green), and DAPI (blue) in control and LoNA KD 2C-like cells. Scale bar: 10 μm. l, Representative immunofluorescence images of FBL and NPM1 PLA signals (magenta), LoNA RNA FISH (red), and DAPI (blue) in control and LoNA KD 2C-like cells, wherein KD cells were treated with an equal amount of LoNA-WT or LoNA-FBL/NPM1-BD. Scale bar: 10 μm. m, Upper: Schematic illustration of a nucleolus, and distance from the center of the nucleolus to the edge of the fluorescence signals for the indicated biomolecules. R1 represents the distance from the center to the outer edge of NPM1 signal; R2 represents the distance from the center to the outer edge of LoNA signal; R3 represents the distance from the center to the outer edge of FBL signal. Lower: the R1, R2, and R3 were calculated in nucleoli of 2C-like cells (n = 30 nucleoli). n,o, Quantification for number (n) (n = 14) and size (o) (n = 19) of droplets with the bilayer structure in the presence of 1 μM FBL, 10 μM NPM1, and indicated concentration of LoNA in in vitro phase separation assays; two-sided paired t-test; ***P < 0.001. Data are presented as mean values ± s.e.m.
Extended Data Fig. 3 LoNA promotes the fluidity of phase-separated condensates of NPM1 and FBL in vivo.
a, Representative in vitro images of mixture of recombinant NPM1 and FBL proteins and visualization of droplets in the presence of indicated amounts of recombinant proteins and rRNA. NPM1 was labeled with DyLight 594 (red) and FBL was labeled with Alexa Fluor 488 (green). Scale bar: 25 μm. b, c, Quantification for number (b) (n = 14) and size (c) (n = 19) of droplets with the bilayer structure in the presence of 1 μM FBL, 10 μM NPM1, and indicated concentration of rRNA in in vitro phase separation assays; two-sided paired t-test; ***P < 0.001. d, Indicated amounts of LoNA plasmid were electroporated into 2C-like cells for LoNA expression, followed by electroporation with 1 μg NPM1-mCherry plasmid and 1 μg FBL-zsGreen plasmid for expression of NPM1-mCherry and FBL-zsGreen. Quantification for the nucleolus area per nucleus to indicate the size of nucleolus (n = 25 nuclei); two-sided paired t-test; ***P < 0.001. e, Levels of NPM1 and FBL in control and LoNA KD 2C-like cells, assessed by immunoblotting with anti-NPM1 and anti-FBL antibodies and densiometric analyses (n = 3); two-way ANOVA with Šidák’s test; n.s. P > 0.05. f,h, 2C-like cells were electroporated with indicated amounts of LoNA plasmid, followed by electroporation with NPM1-mCherry and FBL-ZsGreen plasmids. f, Representative images of fluorescence recovery after photobleaching (FRAP) measurement of mCherry and ZsGreen fluorescence in vivo. White arrowheads indicate the same area before (pre-bleach), at 1, 180, 360, and 480 seconds (s) after photobleaching. Scale bar: 1 μm. g,h, FRAP curves of mCherry (g) and ZsGreen (h) fluorescence over a 480 s period (n = 3). i-k, Visualization of the movement of fluorescently labeled recombinant proteins (mCherry-NPM1 and ZsGreen-FBL), in the presence of indicated amounts of RNA LoNA. i, Representative images of FRAP measurements in vitro. White arrowheads indicate the same area before (pre-bleach), at 1, 180, 360, and 480 seconds (s) after photobleaching. Scale bar: 5 μm. j,k, FRAP curves of mCherry (j) and ZsGreen (k) fluorescence over a 480 s period (n = 3). Data are presented as mean values ± s.e.m.
Extended Data Fig. 4 Loss of LoNA leads to two-cell embryonic arrest.
a, Generation of LoNA knockout mice. Schematic of the genomic target sites in Lona locus and genotyping result of the founder line mice. b, Body weight of LoNA heterozygous knockout (Lona+/−) and wild-type (WT) mice (n = 10); two-sided paired t-test, n.s. P > 0.05. c, Appearance of Lona+/− and WT mice. d, Genotype distribution of embryos produced from Lona+/− intercrossed mice, at embryonic days (E) 7.5, 9.5, and 13.5. e, Genotype distribution of zygote, two-cell, and four-cell embryos produced from Lona+/− mice gametes from three independent experiments. f, Quantification of zygotes and two-cell embryos developed from WT or Lona+/− mice gametes at 24 h after IVF. The numbers of embryos are indicated above the bars. g, Quantification of two- and four-cell embryos developed from WT or Lona+/− mice gametes at 48 h after IVF. h, LoNA genotype in normal four-cell and arrested two-cell embryos produced from WT or Lona+/− mice gametes, assessed by RT-PCR. i, Genotype distribution of arrested two-cell embryos produced from Lona+/− gametes from three independent experiments. j,k, WT zygotes were microinjected with a control ASO or an ASO targeting Lona. j, Quantification of two- and four-cell embryos from three independent experiments. k, Relative LoNA levels in normal developing and arrested embryos (n = 3); two-sided paired t-test, ***P < 0.001. Data are presented as mean values ± s.e.m.
Extended Data Fig. 5 LoNA knockout at advanced developmental stages results in developmental arrest and failed nucleolar formation.
a-h, Cultured four-cell, eight-cell, and sixteen-cell embryos produced from CAG-CreER/Lonaflox/flox mice gametes (from a cross of CAG-CreER and Lonaflox/flox mice) were treated with tamoxifen (LoNA KO) or DMSO (control). a-c, Quantification of four-cell/eight-cell (a), eight-cell/sixteen-cell (b), sixteen-cell/thirty-two-cell (c) embryos. The numbers of embryos quantified in three independent experiments are indicated above the bars. d-h, Representative immunofluorescence images of FBL (green), NPM1 (red), LoNA RNA FISH (magenta), and DAPI (blue) in four-cell embryos (d), eight-cell embryos (e), and sixteen-cell embryos (g). (f and h), Intensity profiles of FBL (gray) and NPM1 (red) in control or LoNA KO embryos obtained using ImageJ software, along an ideal straight line (white). Scale bar: 5 μm.
Extended Data Fig. 6 LoNA-WT, not its binding-dead mutants, rescues two-cell embryonic arrest in Lona−/− embryos.
a-c, Embryos developed from WT or Lona+/− mice gametes were microinjected with control RNA, LoNA-WT, LoNA-FBL-BD, LoNA-NPM1-BD, or LoNA-FBL/NPM1-BD at 6 h after IVF. a, Quantification of two- and four-cell embryos at 48 h after IVF from three independent experiments. b,c, Genotype distribution of embryos at 48 h after IVF from three independent experiments. The numbers of embryos quantified from three independent experiments are indicated above the bars (in c). d, Relative levels of LoNA-WT and BD LoNA mutants, assessed by qPCR (n = 3); two-sided paired t-test; ***P < 0.001. e,f, WT zygotes were microinjected with PBS (Mock), control RNA or control ASO at 6 h after IVF. Quantification of two-cell and four-cell embryos at 48 h after IVF. The numbers of embryos quantified from three independent experiments are indicated above the bars (in e). Data are presented as mean values ± s.e.m.
Extended Data Fig. 7 LoNA KD leads to repression of 2-cell transcriptional program in 2C-like cells.
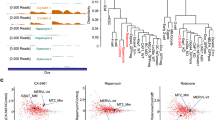
a, Pie chart showing number of up- and down-regulated differentially expressed genes (DEGs) in Lona−/− vs. WT embryos at 24 h after IVF. b-d, Volcano plots showing the DEGs in Lona−/− vs. WT embryos at 18 h (b), 20 h (c), and 22 h (d) after IVF (n = 4); two-sided paired t-test. e, Volcano plots showing the DEGs in LoNA KD vs control 2C-like cells (n = 3); two-sided paired t-test. f, The intersection of down-regulated genes in LoNA KD vs. control 2C-like cells (at least 2-fold) and major reported 2C genes, visualized in a Venn diagram. g, Gene Ontology (GO) analysis of genes that were down-regulated at least 2-fold in LoNA KD vs. control 2C-like cells. h, PCA scores plot for RNA-seq data generated from our 2C-like cells, published mESCs, and published 2C-like cells. i, Relative levels of selective minor ZGA genes (MuERV-L and Klf5) and LoNA in control and α–amanitin (DRB) treated zygotes (6 h after IVF) (n = 3); two-way ANOVA with Šidák’s test; ***P < 0.001. j, Relative levels of selective minor ZGA genes (MuERV-L and Klf5) and LoNA in control and Actinomycin D (ActD) treated zygotes (6 h after IVF) (n = 3); two-way ANOVA with Šidák’s test; ***P < 0.001. Data are presented as mean values ± s.e.m.
Extended Data Fig. 8 LoNA deficiency leads to repression of 2C transcriptional program via increasing H3K27me3 enrichment on 2C gene loci.
a, Representative immunofluorescence images of H3K27me3 (red) and DAPI (blue) in WT and Lona−/− two-cell embryos. Scale bar: 20 μm. b, Enrichment of H3K27me3 on the representative 2C genes (Zscan4d, Bc147527, Tcstv3, Gm4981, and Mbd3l2) was analyzed by H3K27me3 ChIP-seq in WT and Lona−/− two-cell embryos (n = 150). c, Heatmap visualization and band plot of H3K27me3 around the peak center (transcriptional start site, TSS) for WT and Lona−/− two-cell embryos (n = 150). d, Representative 2C marker gene levels in WT and Lona−/− two-cell embryos, assessed by qPCR (n = 3); two-way ANOVA with Šidák’s test; ***P < 0.001. e, Overlap gene number between genes enriched with H3K27me3 and reported 2C genes, visualized in a Venn diagram. f, Representative immunofluorescence images of H3K4me3 (red) and DAPI (blue) in WT and Lona−/− two-cell embryos. Scale bar: 10 μm. g, Enrichment of H3K4me3 on the representative 2C genes (Gm4981 and Mbd3l2) was analyzed by H3K4me3 ChIP-seq in WT and Lona−/− two-cell embryos (n = 150). h, Heatmap visualization and band plot of H3K4me3 around the peak center (transcriptional start site, TSS) for WT and Lona−/− two-cell embryos (n = 150). Data are presented as mean values ± s.e.m.
Extended Data Fig. 9 Deposition of H3K27me3 on 2C gene loci is dependent on p300-mediated NPM1 acetylation.
a, Representative immunofluorescence images of acNPM1 (red), LoNA RNA FISH (magenta), and DAPI (blue) in control and LoNA KD 2C-like cells. Scale bar: 2 μm. b, acNPM1 levels in LoNA KD 2C-like cells treated with or without C646, the p300 inhibitor, assessed by immunoblotting with an anti-acNPM1 antibody and subsequent densiometric analyses (n = 3); two-sided paired t-test; ***P < 0.001. c, Enrichment of acNPM1 on the representative 2C genes (Zscan4d, Bc147527, Tcstv3, Gm4981 and Mbd3l2) was analyzed by acNPM1 ChIP-seq for WT 2-cell embryos, Lona−/− two-cell embryos, and Lona−/− two-cell embryos treated with the p300 inhibitor (n = 200). d, Heatmap visualization and band plot of acNPM1 around the peak center (transcriptional start site, TSS) for WT two-cell embryos, Lona−/− two-cell embryos, and Lona−/− two-cell embryos treated with the p300 inhibitor (n = 200). e, Overlap of genes enriched with acNPM1 and reported 2C genes, visualized in a Venn diagram. f, Overlap of genes enriched with H3K27me3 and acNPM1, and reported 2C genes was visualized in a Venn diagram. g, Relative levels of 2C genes and non-2C genes enriched with both NPM1 and H3K27me3 in WT and Lona−/− embryos, visualized in a heatmap. h, The association of PRC2 complex and acNPM1 in control and LoNA KD 2C-like cells, assessed by immunoprecipitation with an anti-EZH2 antibody, followed by immunoblotting with anti-acNPM1, anti-EED, or anti-SUZ12 antibodies. i, Representative immunofluorescence images of acNPM1 and EZH2 PLA signals (red), LoNA RNA FISH (magenta), and DAPI (blue) in Lona−/− two-cell embryos supplemented with control RNA, LoNA-WT, or LoNA-FBL/NPM1-BD. Scale bar: 5 μm. Data are presented as mean values ± s.e.m.
Extended Data Fig. 10 PRC2 targeting 2C gene loci is dependent on acNPM1’s protein and DNA binding domains.
a, Levels of EZH2, EED, and SUZ12 in control and LoNA KD 2C-like cells (n = 3); two-way ANOVA with Šidák’s test; n.s. P > 0.05. b, Schematic drawing of NPM1 and its DNA or protein BD mutants. c-h, WT-NPM1-HA, NPM1-DBD-HA, or NPM1-PBD-HA plasmid was electroporated into LoNA KD 2C-like cells. c, Association of NPM1 or its mutants with PRC2 complex. d-f, acNPM1 in control and LoNA KD 2C-like cells overexpressing WT-NPM1-HA (d), NPM1-DBD-HA (e), or NPM1-PBD-HA (f). g, h, LoNA KD 2C-like cells electroporated with WT-NPM1-HA, NPM1-DBD-HA, or NPM1-PBD-HA. g, Enrichment of acNPM1 on 2C genes (n = 3); two-way ANOVA with Šidák’s test; n.s. P > 0.05; ***P < 0.001. h, Co-occupancy of acNPM1 and H3K27me3 on 2C genes (n = 3); two-way ANOVA with Šidák’s test; n.s. P > 0.05; ***P < 0.001. i, Zygotes generated from Lona+/− gametes were microinjected with a control or EZH2 ASO at 6 h after IVF. Quantification of two-cell and four-cell embryo numbers. j, MEF cells derived from CAG-CreER/Lonaflox/flox mice were treated with tamoxifen (LoNA KO) or DMSO (control). Representative images of NPM1 (green), FBL (red), LoNA RNA FISH (magenta), and DAPI (blue). Scale bar: 2 μm. k, Representative images of NPM1 (green), FBL (red), LoNA RNA FISH (magenta), and DAPI (blue) in control and LoNA KD mESC cells. Scale bar: 2 μm. l, Levels of NPM1 and NCL in 2C-like and N2a cells. m, LoNA associated NPM1 or NCL in 2C-like and N2a cells. n, Representative images of NPM1 (red), FBL (green), LoNA RNA FISH (magenta), and DAPI (blue) in control and LoNA KD N2a cells. Scale bar: 2 μm. o, Neurons were prepared from CAG-CreER/Lonaflox/flox mouse brains and then treated with tamoxifen (LoNA KO) or DMSO (control). Representative images of NPM1 (red), FBL (green), LoNA RNA FISH (magenta), and DAPI (blue). Scale bar: 2 μm. p, Levels of 45S rRNA at the indicated time points post insemination (n = 3). The thick line indicates the mean and the area shaded in lighter color indicates s.e.m. q, rRNA levels in control and LoNA KD 2C-like cells (n = 3); two-way ANOVA with Šidák’s test; ***P < 0.001. Data are presented as mean values ± s.e.m.
Supplementary information
Supplementary Tables 1 and 2
Table 1: ChIP–seq quality metrics Table 2: Sequence of oligos
Source data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 1
Unprocessed western blots
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 4
Statistical Source Data
Source Data Fig. 5
Statistical Source Data
Source Data Fig. 6
Statistical Source Data
Source Data Fig. 6
Unprocessed western blots
Source Data Extended Data Fig. 1
Statistical Source Data
Source Data Extended Data Fig. 1
Unprocessed western blots
Source Data Extended Data Fig. 2
Statistical Source Data
Source Data Extended Data Fig. 2
Unprocessed western blots
Source Data Extended Data Fig. 3
Statistical Source Data
Source Data Extended Data Fig. 3
Unprocessed western blots
Source Data Extended Data Fig. 4
Statistical Source Data
Source Data Extended Data Fig. 4
Unprocessed western blots/gels
Source Data Extended Data Fig. 5
Statistical Source Data
Source Data Extended Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 7
Statistical Source Data
Source Data Extended Data Fig. 8
Statistical Source Data
Source Data Extended Data Fig. 9
Statistical Source Data
Source Data Extended Data Fig. 9
Unprocessed western blots
Source Data Extended Data Fig. 10
Statistical Source Data
Source Data Extended Data Fig. 10
Unprocessed western Blots
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, D., Cao, R., Li, Q. et al. Nucleolus assembly impairment leads to two-cell transcriptional repression via NPM1-mediated PRC2 recruitment. Nat Struct Mol Biol 30, 914–925 (2023). https://doi.org/10.1038/s41594-023-01003-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-023-01003-w