Abstract
Processivity clamps tether DNA polymerases to DNA, allowing their access to the primer–template junction. In addition to DNA replication, DNA polymerases also participate in various genome maintenance activities, including translesion synthesis (TLS). However, owing to the error-prone nature of TLS polymerases, their association with clamps must be tightly regulated. Here we show that fork-associated ssDNA-binding protein (SSB) selectively enriches the bacterial TLS polymerase Pol IV at stalled replication forks. This enrichment enables Pol IV to associate with the processivity clamp and is required for TLS on both the leading and lagging strands. In contrast, clamp-interacting proteins (CLIPs) lacking SSB binding are spatially segregated from the replication fork, minimally interfering with Pol IV-mediated TLS. We propose that stalling-dependent structural changes within clusters of fork-associated SSB establish hierarchical access to the processivity clamp. This mechanism prioritizes a subset of CLIPs with SSB-binding activity and facilitates their exchange at the replication fork.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data sets from the current study are available from the corresponding author on reasonable request. A structure of E. coli Pol IV was retrieved from the PDB with accession code 4Q43. Source data are provided with this paper.
Code availability
The custom-written computer codes from the current study are available from the corresponding author on reasonable request.
References
Fuchs, R. P. & Fujii, S. Translesion DNA synthesis and mutagenesis in prokaryotes. Cold Spring Harb. Perspect. Biol. 5, a012682 (2013).
Moldovan, G.-L., Pfander, B. & Jentsch, S. PCNA, the maestro of the replication fork. Cell 129, 665–679 (2007).
Dalrymple, B. P., Kongsuwan, K., Wijffels, G., Dixon, N. E. & Jennings, P. A. A universal protein-protein interaction motif in the eubacterial DNA replication and repair systems. Proc. Natl Acad. Sci. USA 98, 11627–11632 (2001).
Georgescu, R. E. et al. Structure of a sliding clamp on DNA. Cell 132, 43–54 (2008).
Krishna, T. S., Kong, X. P., Gary, S., Burgers, P. M. & Kuriyan, J. Crystal structure of the eukaryotic DNA polymerase processivity factor PCNA. Cell 79, 1233–1243 (1994).
Bunting, K. A., Roe, S. M. & Pearl, L. H. Structural basis for recruitment of translesion DNA polymerase Pol IV/DinB to the beta-clamp. EMBO J. 22, 5883–5892 (2003).
Patoli, A. A., Winter, J. A. & Bunting, K. A. The UmuC subunit of the E. coli DNA polymerase V shows a unique interaction with the β-clamp processivity factor. BMC Struct. Biol. 13, 12 (2013).
López de Saro, F. J. & O’Donnell, M. Interaction of the beta sliding clamp with MutS, ligase, and DNA polymerase I. Proc. Natl Acad. Sci. USA 98, 8376–8380 (2001).
Kurz, M., Dalrymple, B., Wijffels, G. & Kongsuwan, K. Interaction of the sliding clamp beta-subunit and Hda, a DnaA-related protein. J. Bacteriol. 186, 3508–3515 (2004).
Ozaki, S. et al. A replicase clamp-binding dynamin-like protein promotes colocalization of nascent DNA strands and equipartitioning of chromosomes in E. coli. Cell Rep. 4, 985–995 (2013).
Jeruzalmi, D. et al. Mechanism of processivity clamp opening by the delta subunit wrench of the clamp loader complex of E. coli DNA polymerase III. Cell 106, 417–428 (2001).
Tan, K. W., Pham, T. M., Furukohri, A., Maki, H. & Akiyama, M. T. Recombinase and translesion DNA polymerase decrease the speed of replication fork progression during the DNA damage response in Escherichia coli cells. Nucleic Acids Res. 43, 1714–1725 (2015).
Pluciennik, A., Burdett, V., Lukianova, O., O’Donnell, M. & Modrich, P. Involvement of the clamp in methyl-directed mismatch repair in vitro. J. Biol. Chem. 284, 32782–32791 (2009).
Furukohri, A., Nishikawa, Y., Akiyama, M. T. & Maki, H. Interaction between Escherichia coli DNA polymerase IV and single-stranded DNA-binding protein is required for DNA synthesis on SSB-coated DNA. Nucleic Acids Res. 40, 6039–6048 (2012).
Arad, G., Hendel, A., Urbanke, C., Curth, U. & Livneh, Z. Single-stranded DNA-binding protein recruits DNA polymerase V to primer termini on RecA-coated DNA. J. Biol. Chem. 283, 8274–8282 (2008).
Molineux, I. J. & Gefter, M. L. Properties of the Escherichia coli in DNA binding (unwinding) protein: interaction with DNA polymerase and DNA. Proc. Natl Acad. Sci. USA 71, 3858–3862 (1974).
Shereda, R. D., Kozlov, A. G., Lohman, T. M., Cox, M. M. & Keck, J. L. SSB as an organizer/mobilizer of genome maintenance complexes. Crit. Rev. Biochem. Mol. Biol. 43, 289–318 (2008).
Chang, S. et al. A gatekeeping function of the replicative polymerase controls pathway choice in the resolution of lesion-stalled replisomes. Proc. Natl Acad. Sci. USA 116, 25591–25601 (2019).
Sigal, N., Delius, H., Kornberg, T., Gefter, M. L. & Alberts, B. A DNA-unwinding protein isolated from Escherichia coli: its interaction with DNA and with DNA polymerases. Proc. Natl Acad. Sci. USA 69, 3537–3541 (1972).
Shereda, R. D., Reiter, N. J., Butcher, S. E. & Keck, J. L. Identification of the SSB binding site on E. coli RecQ reveals a conserved surface for binding SSB’s C terminus. J. Mol. Biol. 386, 612–625 (2009).
Ryzhikov, M., Koroleva, O., Postnov, D., Tran, A. & Korolev, S. Mechanism of RecO recruitment to DNA by single-stranded DNA binding protein. Nucleic Acids Res. 39, 6305–6314 (2011).
Uchida, K. et al. Overproduction of Escherichia coli DNA polymerase DinB (Pol IV) inhibits replication fork progression and is lethal. Mol. Microbiol. 70, 608–622 (2008).
Scotland, M. K. et al. A genetic selection for DinB mutants reveals an interaction between DNA polymerase IV and the replicative polymerase that is required for translesion synthesis. PLoS Genet. 11, e1005507 (2015).
Jarosz, D. F., Godoy, V. G., Delaney, J. C., Essigmann, J. M. & Walker, G. C. A single amino acid governs enhanced activity of DinB DNA polymerases on damaged templates. Nature 439, 225–228 (2006).
Bjedov, I. et al. Involvement of Escherichia coli DNA polymerase IV in tolerance of cytotoxic alkylating DNA lesions in vivo. Genetics 176, 1431–1440 (2007).
Heltzel, J. M. H., Maul, R. W., Scouten Ponticelli, S. K. & Sutton, M. D. A model for DNA polymerase switching involving a single cleft and the rim of the sliding clamp. Proc. Natl Acad. Sci. USA 106, 12664–12669 (2009).
Benson, R. W., Cafarelli, T. M., Rands, T. J., Lin, I. & Godoy, V. G. Selection of dinB alleles suppressing survival loss upon dinB overexpression in Escherichia coli. J. Bacteriol. 196, 3023–3035 (2014).
Indiani, C., Langston, L. D., Yurieva, O., Goodman, M. F. & O’Donnell, M. Translesion DNA polymerases remodel the replisome and alter the speed of the replicative helicase. Proc. Natl Acad. Sci. USA 106, 6031–6038 (2009).
Thrall, E. S., Kath, J. E., Chang, S. & Loparo, J. J. Single-molecule imaging reveals multiple pathways for the recruitment of translesion polymerases after DNA damage. Nat. Commun. 8, 2170 (2017).
Uphoff, S., Reyes-Lamothe, R., Garza de Leon, F., Sherratt, D. J. & Kapanidis, A. N. Single-molecule DNA repair in live bacteria. Proc. Natl Acad. Sci. USA 110, 8063–8068 (2013).
Godoy, V. G. et al. UmuD and RecA directly modulate the mutagenic potential of the Y family DNA polymerase DinB. Mol. Cell 28, 1058–1070 (2007).
Sladewski, T. E., Hetrick, K. M. & Foster, P. L. Escherichia coli Rep DNA helicase and error-prone DNA polymerase IV interact physically and functionally. Mol. Microbiol. 80, 524–541 (2011).
Cohen, S. E., Godoy, V. G. & Walker, G. C. Transcriptional modulator NusA interacts with translesion DNA polymerases in Escherichia coli. J. Bacteriol. 191, 665–672 (2009).
Cafarelli, T. M., Rands, T. J. & Godoy, V. G. The DinB•RecA complex of Escherichia coli mediates an efficient and high-fidelity response to ubiquitous alkylation lesions. Environ. Mol. Mutagen. 55, 92–102 (2014).
Kath, J. E. et al. Polymerase exchange on single DNA molecules reveals processivity clamp control of translesion synthesis. Proc. Natl Acad. Sci. USA 111, 7647–7652 (2014).
Jergic, S. et al. A direct proofreader–clamp interaction stabilizes the Pol III replicase in the polymerization mode. EMBO J. 32, 1322–1333 (2013).
Pagès, V., Mazón, G., Naiman, K., Philippin, G. & Fuchs, R. P. Monitoring bypass of single replication-blocking lesions by damage avoidance in the Escherichia coli chromosome. Nucleic Acids Res. 40, 9036–9043 (2012).
Li, G.-W., Burkhardt, D., Gross, C. & Weissman, J. S. Quantifying absolute protein synthesis rates reveals principles underlying allocation of cellular resources. Cell 157, 624–635 (2014).
Henrikus, S. S. et al. DNA polymerase IV primarily operates outside of DNA replication forks in Escherichia coli. PLoS Genet. 14, e1007161 (2018).
Heltzel, J. M. H., Maul, R. W., Wolff, D. W. & Sutton, M. D. Escherichia coli DNA Polymerase IV (Pol IV), but not Pol II, dynamically switches with a stalled Pol III* replicase. J. Bacteriol. 194, 3589–3600 (2012).
Reyes-Lamothe, R., Sherratt, D. J. & Leake, M. C. Stoichiometry and architecture of active DNA replication machinery in Escherichia coli. Science 328, 498–501 (2010).
Lau, I. F. et al. Spatial and temporal organization of replicating Escherichia coli chromosomes. Mol. Microbiol. 49, 731–743 (2003).
Davies, B. W. et al. Hydroxyurea induces hydroxyl radical-mediated cell death in Escherichia coli. Mol. Cell. 36, 845–860 (2009).
Butland, G. et al. Interaction network containing conserved and essential protein complexes in Escherichia coli. Nature 433, 531–537 (2005).
Arifuzzaman, M. et al. Large-scale identification of protein-protein interaction of Escherichia coli K-12. Genome Res. 16, 686–691 (2006).
Wu, C. A., Zechner, E. L., Reems, J. A., McHenry, C. S. & Marians, K. J. Coordinated leading- and lagging-strand synthesis at the Escherichia coli DNA replication fork. V. Primase action regulates the cycle of Okazaki fragment synthesis. J. Biol. Chem. 267, 4074–4083 (1992).
Pagès, V. & Fuchs, R. P. Uncoupling of leading- and lagging-strand DNA replication during lesion bypass in vivo. Science 300, 1300–1303 (2003).
Kim, S., Dallmann, H. G., McHenry, C. S. & Marians, K. J. Coupling of a replicative polymerase and helicase: a tau-DnaB interaction mediates rapid replication fork movement. Cell 84, 643–650 (1996).
Marceau, A. H. et al. Structure of the SSB-DNA polymerase III interface and its role in DNA replication. EMBO J. 30, 4236–4247 (2011).
Indiani, C., Patel, M., Goodman, M. F. & O’Donnell, M. E. RecA acts as a switch to regulate polymerase occupancy in a moving replication fork. Proc. Natl Acad. Sci. USA 110, 5410–5415 (2013).
Ogawa, T. & Okazaki, T. Discontinuous DNA replication. Annu. Rev. Biochem. 49, 421–457 (1980).
Wu, C. A., Zechner, E. L. & Marians, K. J. Coordinated leading- and lagging-strand synthesis at the Escherichia coli DNA replication fork. I. Multiple effectors act to modulate Okazaki fragment size. J. Biol. Chem. 267, 4030–4044 (1992).
Ikeda, M. et al. DNA polymerase IV mediates efficient and quick recovery of replication forks stalled at N2-dG adducts. Nucleic Acids Res. 42, 846-8472 (2014).
Yeeles, J. T. P. & Marians, K. J. Dynamics of leading-strand lesion skipping by the replisome. Mol. Cell 52, 855–865 (2013).
Morimatsu, K. & Kowalczykowski, S. C. RecFOR proteins load RecA protein onto gapped DNA to accelerate DNA strand exchange: a universal step of recombinational repair. Mol. Cell. 11, 1337–1347 (2003).
Bell, J. C., Plank, J. L., Dombrowski, C. C. & Kowalczykowski, S. C. Direct imaging of RecA nucleation and growth on single molecules of SSB-coated ssDNA. Nature 491, 274–278 (2012).
Isogawa, A., Ong, J. L., Potapov, V., Fuchs, R. P. & Fujii, S. Pol V-mediated translesion synthesis elicits localized untargeted mutagenesis during post-replicative gap repair. Cell Rep. 24, 1290–1300 (2018).
Fuchs, R. P. Tolerance of lesions in E. coli: chronological competition between translesion synthesis and damage avoidance. DNA Repair 44, 51–58 (2016).
Marians, K. J. Lesion bypass and the reactivation of stalled replication forks. Annu. Rev. Biochem. 87, 217–238 (2018).
Fujii, S. & Fuchs, R. P. Defining the position of the switches between replicative and bypass DNA polymerases. EMBO J. 23, 4342–4352 (2004).
Jarosz, D. F., Cohen, S. E., Delaney, J. C., Essigmann, J. M. & Walker, G. C. A DinB variant reveals diverse physiological consequences of incomplete TLS extension by a Y-family DNA polymerase. Proc. Natl Acad. Sci. USA 106, 21137–21142 (2009).
Tanner, N. A. et al. Single-molecule studies of fork dynamics in Escherichia coli DNA replication. Nat. Struct. Mol. Biol. 15, 170–176 (2008).
Pritchard, A. E., Dallmann, H. G., Glover, B. P. & McHenry, C. S. A novel assembly mechanism for the DNA polymerase III holoenzyme DnaX complex: association of δδ′ with DnaX4 forms DnaX3δδ′. EMBO J. 19, 6536–6545 (2000).
Sharan, S. K., Thomason, L. C., Kuznetsov, S. G. & Court, D. L. Recombineering: a homologous recombination-based method of genetic engineering. Nat. Protoc. 4, 206–223 (2009).
Lutz, R. & Bujard, H. Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraC/I1-I2 regulatory elements. Nucleic Acids Res. 25, 1203–1210 (1997).
Esnault, E., Valens, M., Espéli, O. & Boccard, F. Chromosome structuring limits genome plasticity in Escherichia coli. PLoS Genet. 3, e226 (2007).
Tokunaga, M., Imamoto, N. & Sakata-Sogawa, K. Highly inclined thin illumination enables clear single-molecule imaging in cells. Nat. Methods 5, 159–161 (2008).
Sliusarenko, O., Heinritz, J., Emonet, T. & Jacobs-Wagner, C. High-throughput, subpixel precision analysis of bacterial morphogenesis and intracellular spatio-temporal dynamics. Mol. Microbiol. 80, 612–627 (2011).
Jaqaman, K. et al. Robust single-particle tracking in live-cell time-lapse sequences. Nat. Methods 5, 695–702 (2008).
Aguet, F., Antonescu, C. N., Mettlen, M., Schmid, S. L. & Danuser, G. Advances in analysis of low signal-to-noise images link dynamin and AP2 to the functions of an endocytic checkpoint. Dev. Cell 26, 279–291 (2013).
Zawadzki, P. et al. The localization and action of topoisomerase IV in Escherichia coli chromosome segregation is coordinated by the SMC complex. Cell Rep. 13, 2587–2596 (2015).
Garza de Leon, F., Sellars, L., Stracy, M., Busby, S. J. W. & Kapanidis, A. N. Tracking low-copy transcription factors in living bacteria: the case of the lac repressor. Biophys. J. 112, 1316–1327 (2017).
Acknowledgements
We thank K. Arnett (Center for Macromolecular Interactions, Harvard Medical School) for training and assistance in using a CD spectropolarimeter, J. Keck (University of Wisconsin, Madison) for providing the fluorescein-conjugated SSB-Ct peptide, D. Li (University of Rhode Island) for providing the N2-FFdG-containing oligomer, J. Kath (Harvard Medical School) for construction of tetracycline-inducible strains, and S. Jergic (University of Wollongong) for providing the various purified Pol III polymerase complexes and helpful discussions. This work was supported by National Institutes of Health grants R01 GM114065 (to J.J.L.), The William F. Milton Fund (to S.C.), F32 GM113516 (to E.S.T.), and Agence Nationale de la Recherche (ANR) Grant (GenoBlock ANR-14-CE09-0010-01) (to V.P.).
Author information
Authors and Affiliations
Contributions
S.C., E.S.T., L.L., S.C.P., V.P., and J.J.L. designed and performed research. S.C., E.S.T., L L., S.C.P., V.P., and J.J.L. analyzed data. S.C., E.S.T., L.L., V.P., and J.J.L. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural and Molecular Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Beth Moorefield and Carolina Perdigoto, in collaboration with the Nature Structural & Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 An SSB-binding defective mutant Pol IV.
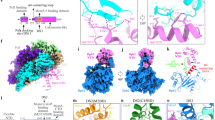
a. Formation of nucleoprotein complexes between SSB4 or SSBΔF4 and ssDNA. (Top) An FP-based binding assay scheme. (Bottom) Interaction of T71-FL with SSB4 or SSBΔF4 was measured by changes in FP. Means ± ranges of duplicate measurements. Similar results were reproduced twice. b. Over-expression-induced cell death by RecQ, Exonuclease I, Topoisomerase III, Pol II, or their variants. R503A of RecQ, and R418A, R316A, and Q311A of Exonuclease I, SSB-binding mutations; R448A of RecQ, a negative control mutation; flag, N-terminal FLAG epitope; exoI, topoIII, and polB, coding sequences for exonuclease I, topoisomerase III, and Pol II respectively. c. Location of T120 within Pol IV. (Left) A front view showing the location of T120 relative to the catalytic cleft (yellow arrow). (Right) A side view showing the location of T120 relative to the C-terminal end (P341). C-terminal unstructured region (a.a. 342-351) containing the clamp interacting motif (a.a. 346-351) is missing in the structure. A crystal structure of E. coli Pol IV (PDB 4q43) is modified.
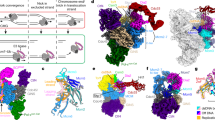
Extended Data Fig. 2 Fates of stalled replication and associated replication products in our rolling circle replication-based assays.
In these rolling circle DNA templates, the lesion-containing closed circular DNA is the template for leading strand synthesis. During rolling circle DNA replication, the growing leading strand serve as the template for lagging strand synthesis.
Extended Data Fig. 3 Pol IV must bind to the β2 clamp to mediate TLS at the fork.
The N2-FFdG-containing template was replicated in the presence of indicated concentrations of Pol IV or Pol IVΔC6 as shown in Fig. 2a. Pol IVΔC6, a mutant Pol IV lacking the C-terminal clamp binding motif.
Extended Data Fig. 4 Monitoring resolution pathway choice in cells.
a. Clamp binding motif (CBM) of the ε subunit of the Pol III core complex (αεθ). Shown are wild-type and mutant CBM sequences and their binding affinities to the β2 clamp. ε-cleft, the binding site on the β2 clamp for the CBM of the ε subunit; εQ, CBM with a weakening mutation; εL, CBM with a strengthening mutation. dnaQ(εQ), weakening mutation. θ subunit is omitted in the schematic diagram for simplicity. b. (Left) Site-specific insertion of a single N2-FFdG adduct into the E. coli genome in either the leading or lagging strand template. (Right) Resolution of lesion-stalled replication through TLS leads to the formation of blue-sectored colonies. White colonies represent resolution through DA. c. Deletion of lafU, which is replaced with an frt-kan-frt cassette as a dinB-linked marker for P1 transduction, does not influence pathway choice of N2-FFdG-stalled replisomes in cells. In all ΔlafU-containing strains, the kan cassette was flipped out. Means ± SDs (n = 6 for lafU+; n = 3 for ΔlafU; n, number of independent experiments). d. TLS over N2-FFdG in cells is primarily mediated by Pol IV. Deleting dinB reduces the resolution of N2-FFdG-stalled replisomes through TLS from ~60 to ~30%. Additional deletion of both polB and umuDC does not further reduce resolution through TLS. The residual TLS (~30%) in the ΔdinB ΔpolB ΔumuDC background is presumably mediated by Pol III. However, as Pol IV outcompetes Pol III in mediating TLS over N2-FFdG, the actual contribution of Pol III to TLS in the dinB+ background is likely less than estimated in the ΔdinB background. Ld, leading strand lesion; Lg, lagging strand lesion. Means ± SDs (n = 3; n, number of independent experiments). The statistical significance of the difference between the indicated pairs was determined by two-tailed Welch’s t-test; numbers, p values. e. Weakening the SSB-Pol IV interaction increases the MMS-induced SOS response. (Left) Resolution of stalled replication through repriming results in induction of the SOS damage response. TLS at the fork inhibits repriming. (Right) The MMS-induced SOS response in strains bearing indicated dinB alleles. The SOS reporter strains express green fluorescent protein (GFP) from a genomic expression cassette, in which the transcription of gfp is under the control of the sulA promoter (see ‘Methods’ for details). dinBplexA, a LexA repressor binding site mutant of the dinB promoter. Fluorescence intensities from single cells were measured for more than 90 × 103 cells using a flow cytometer. Means ± SDs (n > 90 × 103). The statistical significance of the difference between indicated pairs was determined by an unpaired t-test. ****, p < 0.0001. Similar results were reproduced twice.
Extended Data Fig. 5 SSB-driven ectopic localization of CLIPs.
a. Viable cell counts for +lacO250 dinB+ or ΔdinB strains (Fig. 5a) upon induction of LacIΔC11-SSB-Ct (Ct), LacIΔC11-ssb-CtDA,ΔF (CtDA,ΔF) or mock (-). Means ± SDs of triplicate measurements. Similar results were reproduced twice. b. Sensitivity of indicated assay strains to MMS upon induction of Ct or CtDA,ΔF. c. The C-terminal unstructured linker of SSB (SSBIDL in Fig. 1a) is not necessary for the formation of an ectopic SSB-Ct cluster that sensitizes cells to NFZ. Expression of LacIΔC11-SSB-Ct (Ct) or LacIΔC11-SSBIDL-Ct (IDL-Ct) comparably sensitized cells to NFZ. Addition of IPTG or binding mutations on SSB-Ct (CtDA,ΔF) eliminated this sensitization. SSBIDL-Ct, the entire C terminal unstructured linker of SSB including both SSBIDL and SSB-Ct. d. Sensitivity of the indicated strains to NFZ. Similar results were reproduced twice. e. Sensitivity of indicated assay strains to HU upon induction of Ct or CtDA,ΔF. Similar results were reproduced twice. f. ΔdinB is epistatic to polA-recQWH or hda-recQWH in sensitivity to NFZ. Sensitivity of the indicated strains to NFZ. polA- or hda-recQWH, the coding sequence for Pol I- or Hda-RecQWH. g. Sensitivity of indicated strains to NFZ. hda-recQWH or WH(R425A,R503A), the coding sequence for Hda-RecQWH or WH(R425A,R503A). h. Over-production of Crfc-RecQWH results in massive cell death. Expression of indicated genes was induced from a Tet-inducible genomic expression cassette. crfc-recQWH or WH(R425A,R503A), the coding sequence for Crfc-RecQWH or WH(R425A,R503A). Similar results were reproduced twice.
Supplementary information
Supplementary Information
Supplementary Figures 1–5.
Supplementary Table 1
E. coli strains and plasmids used in this study.
Source data
Source Data Fig. 1
Statistical Source Data.
Source Data Fig. 2
Statistical Source Data.
Source Data Fig. 3
Statistical Source Data.
Source Data Fig. 4
Statistical Source Data.
Source Data Fig. 5
Statistical Source Data.
Source Data Extended Data Fig. 1
Statistical Source Data.
Source Data Extended Data Fig. 3
Statistical Source Data.
Source Data Extended Data Fig. 4
Statistical Source Data.
Source Data Extended Data Fig. 5
Statistical Source Data.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chang, S., Thrall, E.S., Laureti, L. et al. Compartmentalization of the replication fork by single-stranded DNA-binding protein regulates translesion synthesis. Nat Struct Mol Biol 29, 932–941 (2022). https://doi.org/10.1038/s41594-022-00827-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-022-00827-2
This article is cited by
-
Focus and persistence: how Pol IV unblocks stalled DNA synthesis
Nature Structural & Molecular Biology (2022)