Abstract
Chitin is an essential component of the fungal cell wall. Chitin synthases (Chss) catalyze chitin formation and translocation across the membrane and are targets of antifungal agents, including nikkomycin Z and polyoxin D. Lack of structural insights into the action of these inhibitors on Chs has hampered their further development to the clinic. We present the cryo-EM structures of Chs2 from Candida albicans (CaChs2) in the apo, substrate-bound, nikkomycin Z-bound, and polyoxin D-bound states. CaChs2 adopts a unique domain-swapped dimer configuration where a conserved motif in the domain-swapped region controls enzyme activity. CaChs2 has a dual regulation mechanism where the chitin translocation tunnel is closed by the extracellular gate and plugged by a lipid molecule in the apo state to prevent non-specific leak. Analyses of substrate and inhibitor binding provide insights into the chemical logic of Chs inhibition, which can guide Chs-targeted antifungal development.
Similar content being viewed by others
Main
Chitin is a linear polymer of β-(1,4)-linked d-glucosamine (GlcNAc) and the second most abundant naturally occurring polymer on Earth, after cellulose, with annual biosynthesis of more than 100,000,000,000 tons1. Chitin is an essential constituent of fungal cell walls and the exoskeletons of crustaceans, mollusks, and insects2. Chitin and chitosan have also been widely studied for biomedical applications such as polymeric tissue and nanocarrier engineering3,4. Chitin is synthesized by integral membrane Chss. Because chitin and Chs are not present in vertebrates and plants, Chs has been pursued as the target for fungicides, insecticides, and acaricides.
Fungal infections pose growing threats to human health and agricultural economics worldwide. In particular, fungal coinfections of patients with COVID-19 substantially increase mortality rates5,6,7,8. Unfortunately, there are limited classes of effective antifungal drugs while drug resistance is on the rise9,10. Development of new antifungals is challenging due to the similarities in cellular machineries between fungi and hosts. Chs is one of the few targets whose inhibition causes highly selective growth inhibition or death of fungi without harming vertebrates or plants. Chss are conserved throughout the fungal kingdom, including but not limited to yeast, molds, and all species of mushrooms and toadstools. Chs is a membrane-embedded processive glycosyltransferase11, and catalyzes homopolymerization of GlcNAc using UDP-GlcNAc as substrate12 to produce chitin, an essential polysaccharide component of the fungal cell wall13. Chs also mediates the extrusion of the polymer product across the membrane to the extracellular side. A naturally occurring Chs inhibitor, polyoxin D, has been widely used as a safe fungicide in agriculture14. Nikkomycin Z has been clinically investigated to treat endemic fungal infectious disease (coccidioidomycosis)15,16. However, the development of polyoxin D and nikkomycin Z beyond their current applications has not been realized because many fungal pathogens contain Chss that are less sensitive to nikkomycins and polyoxins17 and structural and mechanistic insights into their action on Chs are lacking. Here, we report atomic-resolution structures and structure-guided functional studies of Chs2 from Candida albicans, the leading cause of invasive fungal infections in humans. The results provide the structural basis of Chs regulation and inhibition by antifungal agents nikkomycin Z and polyoxin D and offer an unexplored avenue for the development of antifungal drugs9.
Results
Structure determination
There are seven different classes of Chs in fungi, of which Class 1 has been identified as the target of nikkomycins and polyoxins. CaChs2 (Class 1) was chosen as our target for structural analysis because C. albicans is one of the leading pathological fungi in humans. CaChs2 shares 70–90% sequence identity with other Class 1 fungal Chss, 50–70% sequence identity with other classes of fungal Chss, and ~35% sequence identity to insect Chss (Supplementary Fig. 1). We heterologously expressed CaChs2 in insect cells and purified it to homogeneity (Extended Data Fig. 1a). The purified recombinant CaChs2 catalyzed formation of water-insoluble chitin polymers from UDP-GlcNAc with Km = 6.0 ± 0.7 mM and kcat = 8.6 ± 2.7 min−1 (Fig. 1a, Extended Data Fig. 1b and Supplementary Table 1). We used FTIR spectroscopy to demonstrate that the produced chitin polymer adopts the α-chitin configuration (Fig. 1b), consistent with the form of chitin found in fungal cell wall. Mg2+ is essential for CaChs2 activity (Fig. 1c). The purified CaChs2 is inhibited by nikkomycin Z and polyoxin D with Ki = 1.5 ± 0.5 and 3.2 ± 1.4 μM, respectively (Fig. 1d and Supplementary Table 1). Using isothermal titration calorimetry (ITC), we measured the Kd of nikkomycin Z for CaChs2 to be ~190 nM (Extended Data Fig. 1c). These observations are consistent with the previous characterization of the crude preparation of CaChs218 and its homolog in Saccharomyces cerevisiae (ScChs1)19, suggesting successful preparation of physiologically relevant CaChs2.
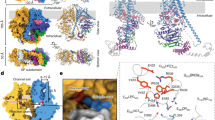
a, Activity assay of CaChs2 measuring the rate of GlcNAc incorporated into insoluble chitin at different UDP-GlcNAc concentrations. The solid line represents a fit to the Michaelis–Menten equation. Data points represent the mean of three independent experiments (n = 3) with enzyme prepared from two biological replicates and error bars represent 1 s.d. b, Characterization of CaChs2 product. Left: FTIR spectra of CaChs2 product (black trace) and commercial shrimp chitin (rose trace) as an α-chitin standard. Right: Region of spectra showing the divided amide I band characteristic of α-chitin. c, GlcNAc incorporated into insoluble chitin over time in the presence or absence of Mg2+. Data points are the mean of three independent experiments (n = 3) with enzyme prepared from two biological replicates with error bars representing 1 s.d. d, CaChs2 activity in response to varying concentrations of nikkomycin Z (circles) and polyoxin D (squares). Each data point is an average of three independent experiment repeats with enzyme prepared from two biological replicates with error bars representing 1 s.d. (n = 3). The solid lines represent a non-linear curve fit to the four-parameter logistic equation. e,f, Unsharpened cryo-EM map of CaChs2 dimer in two orientations (e) and (f). The cryo-EM densities for two Chs2 protomers are colored blue and yellow, respectively. The cryo-EM density for the detergent micelle is shown transparent and colored gray. g,h, Cartoon representation (g) and topology diagram (h) of a CaChs2 monomer. The transmembrane domain (TMD) is colored blue; the interfacial domain (IFD) green; the catalytic helical module (CHM) red; the glycosyltransferase domain (GTD) dark yellow. The β-sheets of the GTD core, the CHM, and the domain-swapped region are highlighted in h. Data for graphs in a, c and d are available as source data.
We determined four single-particle cryo-EM reconstructions of CaChs2 (Extended Data Figs. 2–4 and Table 1): (1) the ligand-free state resolved to ~2.95 Å, (2) in complex with UDP-GlcNAc and Mg2+ to ~3.02 Å, (3) in complex with nikkomycin Z to ~3.19 Å, and (4) in complex with polyoxin D to ~3.15 Å. CaChs2 forms a domain-swapped homodimer (Fig. 1e,f) with each protomer composed of an N-terminal cytoplasmic glycosyltransferase domain (GTD, colored dark yellow in Fig. 1g), an interfacial domain (IFD, green), and a C-terminal transmembrane domain (TMD, blue) constituting the chitin polymer channel (Fig. 1g,h). The separation of the N-terminal cytoplasmic GTD and the C-terminal TMD in the primary sequence contrasts with cellulose synthases, the other membrane-embedded processive GT with structural characterization20,21, indicating a unique architectural design of Chs (Extended Data Fig. 5). GTD is composed of the active site containing core GT-A fold decorated by additional substructures including part of the N-terminal region (134–172)12,22,23. The dimer interaction at TM2 and TM5 is facilitated by two phospholipids in the extracellular leaflet of the membrane. Two additional phospholipids are located outside the dimer interface near the intracellular leaflet of the membrane (Extended Data Fig. 6).
Chitin polymer translocation tunnel
Within the TMD, there is a central tunnel continuous with the cytoplasm (Fig. 2a). The tunnel is about 30 Å long with its radius ~3–4 Å (average 3.5 Å), which can accommodate a chitin chain of about five or six GlcNAc units (Fig. 2b,c). In comparison, the tunnel of the bacterial cellulose synthase BcsA is ~48 Å long but with a radius of 2.5–3.5 Å (Extended Data Fig. 7a,b)20. Similarly, the tunnel of the plant cellulose synthase CesA is ~45 Å long but with a radius of 2.5–4.0 Å. Both structures are consistent with the smaller diameter of cellulose (Extended Data Fig. 7a,b)21. Notably, the tunnel of CaChs2 has a narrow restriction site near the extracellular side and it is occluded by four conserved hydrophobic residues (L706, M775, L804, and L971) (Fig. 2d). This structural feature is analogous to the gate of ion channels. Alanine mutations of residues M775, L804, and L971 reduce CaChs2 activity significantly while L706A slightly increases CaChs2 activity (Fig. 2i) These results underscore the functional importance of this region and a distinct role of L706 in Chs function. Therefore, we tentatively assign the extracellular exit of the tunnel as the gate of CaChs2, although the definite gate assignment requires determination of the chitin polymer-bound CaChs2 structure.
a, Cross-section view of the CaChs2 dimer. One monomer is shown in surface representation, revealing the location and shape of the tunnel, and the other monomer is shown in cartoon representation. b, Ribbon representation of a CaChs2 monomer with the tunnel shown as surface representation. c, Length and radius of the tunnel calculated using the HOLE program. The starting point of the tunnel is defined at the location of W647 and the end point the gate. d, Extracellular gate of the tunnel shown in cartoon and surface representation. The gate residues and the lipid inside the tunnel are shown as sticks. e,f, The chitin translocation tunnel shown as surface (e) and the residues lining the tunnel in stick representation (f). g, Cross-section view of the chitin translocation tunnel. The phospholipid modeled as phosphatidylethanolamine (PE) inside the tunnel and residues interacting with the head group of the lipid are shown as sticks. The position of the lateral opening in this figure is above the plane of the paper. h, Surface representation of the lateral opening of the tunnel viewed from within the membrane leaflet. The lipid molecule and residues that enclose the lateral opening are shown as sticks. i, Comparison of the activities of CaChs2 WT and the tunnel, gate, and lateral opening mutants of CaChs2. Columns represent the mean of at least two independent experiments from two biological replicates each (n = 4 for all except WT where n = 9), and error bars represent 1 s.d. Significance was calculated using two-tailed t-test to compare the activity of the mutant enzymes to the WT enzyme (*P < 0.05, ***P < 0.001, ****P < 0.0001). Data for graph i and exact P values are available as source data.
At the base of the tunnel, there are many polar and charged amino acids. The middle and the outer sections of the tunnel are spatially separated with hydrophobic and hydrophilic regions (Fig. 2e, f). All tunnel-lining residues are conserved among Chss (Supplementary Fig. 1 and Extended Data Fig. 8a,b), indicating a shared mechanism of chitin translocation among Chss. The hydrophilic region may be involved in chitin polymer translocation by interacting with N-acetyl groups of GlcNAc of the growing chitin chain. Consistent with the idea, alanine substitutions of hydrophilic tunnel-lining residues E516, Y517, S695, Y815, Y835, and R985 largely abolish the CaChs2 activity (Fig. 2i). We did not observe cryo-EM density in the central tunnel interpretable as chitin. In contrast, cellulose is bound in the tunnels of all the reported structures of cellulose synthases20,21. This difference may represent a difference in the mechanism of product release and processivity between Chs and cellulose synthase.
Interestingly, we observed cryo-EM density within the tunnel that can be modeled as a phosphatidylethanolamine (PE) molecule (Fig. 2g and Extended Data Fig. 9). Consistent with the cryo-EM density, lipid analysis by LC–MS showed that PE and ceramide are the most abundant lipids that are copurified with detergent solubilized CaChs2 (Extended Data Fig. 9). The central tunnel has a lateral opening toward the lipid membrane where acyl chains of the PE are located (Fig. 2h). Because this lateral opening faces the hydrophobic lipid membrane, it is likely that the portal is for PE to move in and out of the tunnel rather than an exit for the chitin polymer. The closed extracellular gate and the absence of a chitin in the tunnel suggest that the observed CaChs2 structure represents an off state in which the enzyme is reset for another round of chitin synthesis after the release of the chitin polymer. An unregulated opening of the chitin channel domain in the absence of chitin polymer would lead to non-specific leakage of water, ions, and many small metabolites, disrupting cellular homeostasis. Therefore, the chitin channel domain must be tightly regulated before, during, and after the cell wall synthesis, which could be the reason for the dual regulation of the Chs channel by lipid and the gate. Once the enzyme is active, this lipid will be displaced out of the tunnel by a growing nascent chitin polymer. Consistent with this hypothesis, altering the lateral opening by introducing bulky aromatic side chains (T808W, I836W) reduced enzyme activity (Fig. 2i). The fact that T808W and I836W mutants decrease Chs2 activity indicates that the lipid is likely incorporated into the tunnel during folding and that reduction of the lateral opening size would impede chitin translocation within the tunnel by obstructing the displacement of the lipid. In addition, replacing Y812 with alanine also reduced enzyme activity (Fig. 2i). Y812 interacts with F832 in IF3 (Fig. 2h), both of which line the portal, and this interaction may be important to maintain the lateral opening structure. It is possible that the lipid may be partially displaced during the chitin synthesis, but understanding the detailed motion of the lipid would require capturing intermediate states of Chs2.
Substrate binding
The UDP-GlcNAc binding site is close to the cytoplasmic entrance of the tunnel. The extensive interactions between the IFD and the GTD make the central tunnel accessible from the active site of the GTD only, which serves as the cytoplasmic entry point for the donor UDP-GlcNAc (Fig. 3a,b). Substructures from IFD (IF2) and GTD (β4, β10, H5, H7, and H10) form a pocket for UDP-GlcNAc and Mg2+ (Fig. 3b,c). The EM density for UDP-GlcNAc and Mg2+ allows us to build the molecule without ambiguity, although the signal of the GlcNAc moiety is relatively weaker than the UDP moiety (Extended Data Fig. 8c,d), indicating that the GlcNAc moiety is more flexible. The uracil base interacts with Y319 via a π-π interaction and D364 via a H-bond with its N3. The ribose group is surrounded by E321, D465, and K441 with E321 and D465 interacting with 2′-OH and 3′-OH of ribose, respectively. Mg2+ interacts with the β phosphate of UDP and possibly D465. Other than the presence of UDP-GlcNAc and Mg2+, the substrate-bound structure is almost identical to the apo structure (Cα r.m.s.d.~ 0.2 Å). The lipid is still present in the tunnel and the tunnel gate remains closed.
a, Cartoon representation of a CaChs2 monomer bound to UDP-GlcNAc, shown as spheres. b,c, Detailed interactions between CaChs2 and UDP-GlcNAc (b), (c). Hydrophilic interactions are shown as dashed lines except for those of E603 and D604, which were shown to illustrate their long distance to the GlcNAc moiety of UDP-GlcNAc. d, Comparison of the activities of CaChs2 WT and mutants related to the UDP-GlcNAc binding site. Columns represent the mean of at least two independent experiments from two biological replicates each (n = 4 for all except WT where n = 9), and error bars represent 1 s.d. Significance was calculated using two-tailed t-test to compare the activity of the mutant enzymes to the WT enzyme (****P < 0.0001). Data for graph d and exact P values are available as source data.
Q643, R644, R646, and W647 form the conserved Q(R/Q)XRW motif24 in IF2 and appear to play a crucial role in UDP-GlcNAc binding, acceptor binding, and catalysis. First, R646 interacts with the UDP α-phosphate and GlcNAc. The 6′-OH of UDP-GlcNAc and the indole ring of W647 are located above the sugar ring. These interactions position the GlcNAc moiety of UDP-GlcNAc for catalysis (Fig. 3c). Second, Q643, R644, and W647 form a triad, and the indole ring of W647 lines a cavity that is continuous with the central tunnel (Fig. 3c). Trp in the Q(R/Q)XRW motif has been established as the binding site for the acceptor oligosaccharide in the cellulose synthase BcsA20,25. Therefore, we posit that the triad interactions help position the indole ring of W647 to serve as the acceptor binding site in CaChs2. Overlay of the GTDs of Chs2, BcsA and CesA shows that the residues that interact with the substrate and Mg2+are largely conserved (Extended Data Fig. 7c–f).
The putative catalytic base residue D604 is located at the N terminus of H10 and is not within reach of UDP-GlcNAc. The helix containing the corresponding residue in BcsA was termed a ‘finger helix’25. Compared to BcsA, H10 is displaced away from the putative acceptor site about a half-helical turn so that D604 is too far to activate 4′-OH of GlcNAc of the nascent chitin chain (Extended Data Fig. 7d,f). Therefore, UDP-GlcNAc bound CaChs2 represents an off state, consistent with the closed gate and the lipid in the central tunnel. H10 must therefore move closer to the putative acceptor site for catalysis. However, the movement of H10 is likely very different from that of the finger helix in BcsA. Unlike the finger helix that is flanked by loops (Extended Data Fig. 7d,f), H10 in CaChs2 forms a subdomain with H7–H9 (Figs. 1g,h and 3a,b, colored red). Therefore, in CaChs2, H10 motion must involve H7–H9 either by concerted movement or rearrangements. We term H7–H10 as the ‘catalytic helical module (CHM)’.
Consistent with the structure, alanine substitutions of residues for UDP-GlcNAc binding or catalysis (Y319, E321, K441, D465, E603, D604, Q643, R644, R646, W647) lead to complete or significant loss of activity (Fig. 3d). Our data are consistent with the previous identification of the three functionally important sequence motifs (D, (E/D)DX and Q(R/Q)XRW) of the conserved region 1 (dubbed as CON1) in ScChs2 (refs. 24,26). In the CaChs2 structure, these motifs are either involved in UDP-GlcNAc binding or catalysis (Fig. 3c).
Domain swap and the DISW motif
Near the cytoplasmic part of the dimer interface mediated by TM2 and TM5, a large lasso-shaped insertion between IF3 and TM5 (I854–E919) reaches out with the structured region (H11, β16, and β17) as a hook to anchor into the GTD of the neighboring protomer (Fig. 4a). This domain-swapped configuration of the insertion between IF3 and TM5 is confirmed by continuous cryo-EM density of loops connecting the insertion at a lower contour level and structure-guided cysteine disulfide crosslinking experiments (Extended Data Fig. 10). In the GTD, a substructure for the docking of the insertion hook is provided by elements distal to the core GT-A subdomain: a loop in the N-terminal region (Q134–F136), β1, β2, Η2, Η4, and β8 (Fig. 4b). Notably, the loop between IF3 and H10 in this domain-swapped region contains a conserved stretch of amino acids (856D-I-S-W-G-T-K-G863, DISW (domain swap) motif), a section of which was previously identified as the second conserved region in the fungal Chss27. The DISW motif appears to restrict the motion of the CHM and IF2 by forming an extensive interaction network with CHM, TM5, IF2, and H5 (Fig. 4a,c). Therefore, for the CHM and IF2 to move closer to the UDP-GlcNAc binding site for catalysis, either the DISW motif moves together with the CHM and IF2 or the interaction network involving the DISW motif needs to be rearranged, suggesting the importance of the DISW motif in catalysis. Within the DISW motif, individual mutations of D856, S858, and W859 and a simultaneous alanine substitution of the D856–W859 region significantly affected the catalytic activity of CaChs2 (Fig. 4d). In addition, mutation of residues involved in the DISW interaction network (K440, Y600, R645, and Y922) also reduced CaChs2 activity significantly (Fig. 4d). Consistent with these observations, ScChs2 mutants at sites corresponding to D856, W859, or T861 did not produce chitin27. Taken together, we suggest that the domain-swapped dimer configuration of CaChs2 serves a regulatory role, although the detailed mechanism requires additional studies.
a, Cartoon representation of a CaChs2 dimer. The domain-swapped regions are highlighted with surface representation. b, Close-up view of the inter-subunit interaction between the domain-swapped region of one monomer and the GTD of the other monomer. c, Close-up view of the intra-subunit interactions between the DISW motif and multiple regions surrounding the substrate binding site. H-bonds and hydrophobic interactions are labeled as dashed lines. d, Comparison of the activity of WT CaChs2 to DISW motif mutants. Simultaneous alanine substitution of the D856–W859 region is named DISW. Columns represent the mean of at least two independent experiments from two biological replicates each (n = 4 for all except WT where n = 9, H569A where n = 8, and K862A and W859A where n = 3), and error bars represent 1 s.d. Significance was calculated using two-tailed t-test to compare the activity of the mutant enzymes to the WT enzyme. (NS P > 0.05 **P < 0.01, ****P < 0.0001. Data for graph d and exact P values are available as source data.
Nikkomycin Z
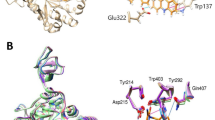
Nikkomycin Z and polyoxin D are competitive inhibitors of Chs and pseudo-dipeptides with an N-terminal amino acid linked to a six-carbon nucleoside, aminohexuronic acid (AHA) (Fig. 5b,g). These compounds are thought to structurally mimic UDP-GlcNAc, with the conserved AHA mimicking the uridine moiety of UDP-GlcNAc28,29. The electrostatic surface representation shows that the substrate entry pocket is spatially divided into electronegative and electropositive sides (Fig. 5e). Nikkomycin Z is bound to CaChs2 in an extended conformation in this pocket where its AHA 5′-carboxylate faces the electropositive side (Fig. 5a,d,e). The nucleobase and the 5′-carboxylate of AHA interact with Y319 and R646 via a π-π interaction and a salt bridge, respectively. 2′-OH of AHA also interacts with E321. The N-terminal amino acid of nikkomycin Z, 4-hydroxypyridylhomothreonine (HPHT), occupies the pocket between the UDP site and the central tunnel. The hydroxypyridinyl ring of HPHT forms the most critical contacts with CaChs2, which interacts with W647 via π-π interactions and with Y517 via an H-bond (Fig. 5d,e). Consistent with our structure, W647A reduces the affinity to nikkomycin Z by about 18-fold whereas Q643A only moderately affects the affinity (Extended Data Fig. 1d,e).
a, Cross-section surface representation of a CaChs2 monomer bound to nikkomycin Z (yellow spheres). b,c, Chemical structure (b) and cryo-EM density (c) of nikkomycin Z. d, Detailed interactions between CaChs2 and nikkomycin Z. Hydrophilic and π-π stacking interactions are labeled with dashed lines. e, Electrostatic surface representation of the nikkomycin Z binding site. Key side chains are shown as sticks. f, Cross-section surface representation of a CaChs2 monomer bound to polyoxin D (yellow spheres). g,h, Chemical structure (g) and cryo-EM density (h) of polyoxin D. i, Detailed interactions between CaChs2 and polyoxin D. Hydrophilic and π-π stacking interactions are labeled with dashed lines. j, Electrostatic surface representation of the polyoxin D binding site. Key side chains are shown as sticks.
The nikkomycin Z-bound CaChs2 structure offers explanations to the previous structure–activity–relationship (SAR) studies on Chs from Coprinus cinereus30. First, the SAR studies showed that the l-configuration of the N-terminal amino acid is important, likely due to the optimal interactions of the pyridinyl ring of HPHT with Chs. Second, the p-hydroxyl group of HPHT contributes to affinity, likely due to its increased π-π interaction with W647 and/or enhanced water-mediated H-bonding with Y517. Third, introducing a methyl group at C-3″ of HPHT increases affinity, likely due to the increased van der Waals interactions and the reduced conformational flexibility of the molecule. Fourth, a pseudo-tripeptide nikkomycin with glutamate conjugated to the 5′-carboxylate of AHA binds with several-fold lower affinity, as the additional glutamate disrupts the interaction of AHA with R646.
Comparison between nikkomycin Z and UDP-GlcNAc binding to CaChs2 reveals that although the AHA binding site overlaps with that of UDP of UDP-GlcNAc, there are noticeable differences in their positions with CaChs2, resulting in the differential interactions with CaChs2 (Fig. 6a). The 5′-carboxyl of AHA, the phosphate mimic of UDP28,29, gains additional interaction with K441. While the ribose 3′-OH of UDP-GlcNAc interacts with D465 (2.5 Å), this interaction was lost or severely weakened in AHA (3.7 Å) because D465 rotates away from sugar to form a salt bridge with K441. Notably, the active site of CaChs2 underwent rearrangements to accommodate nikkomycins Z, indicating the structural plasticity of CaChs2 (Fig. 6a).
a, Overlay of the substrate binding sites of UDP-GlcNAc-bound (brown) and nikkomycin Z-bound (green) CaChs2. b, Overlay of the substrate binding sites of nikkomycin Z-bound (green) and polyoxin D-bound (magenta) CaChs2. H-bond and π-π stacking interactions between substrate/ligand to CaChs2 are labeled with dashed lines in indicated colors.
Together, the unique interactions of HPHT and AHA with CaChs2 contribute to the specificity of nikkomycin Z for CaChs2. These observations suggest that nikkomycin Z is not merely a mimic of UDP-GlcNAc as previously thought and explain the substantially stronger affinity of nikkomycin Z to CaChs2 than UDP-GlcNAc (three orders of magnitude difference in Ki and Km).
Polyoxin D
Polyoxin D comprises a non-aromatic N-terminal amino acid, carbamoylpolyoxamic acid (CPOAA), and AHA with a 5-carboxyuracil base (5-carboxy-AHA) (Fig. 5g). CPOAA forms extensive interactions with the QRXRW motif. Particularly, Q643 plays a critical role by interacting with 3″-OH and 4″-OH of polyoxin D. This is in stark contrast to nikkomycin Z binding, where Q643 does not form an H-bond interaction. The 5-carboxyl group to the uracil base likely contributes to CaChs2 binding by its increased π-π interactions with Y319 and interaction with K440 (Fig. 5i,j). The polyoxin-bound CaChs2 structure is consistent with previous SAR studies28,31,32,33. First, 3″-OH, 4″-OH, and 2″-NH2 of CPOAA play crucial roles in the binding of polyoxin to Chs. The importance of 3″-OH and 4″-OH are well supported by their interactions with Q643. While the 2″-NH2 does not make direct interactions with the protein, it faces the electronegative part of the enzyme pocket, so it may interact with CaChs2 through either electrostatic interactions or via water-mediated H-bonding with nearby residues such as T631, D632, or the backbone carbonyl of C466 (Fig. 5i, j). Consistent with this idea, we found that alanine mutation of T631 decreases its affinity for polyoxin D by about threefold (Supplementary Table 1). Second, the 5-carboxyl group of the uracil base also contributes to the polyoxin binding to Chs31. Notably, because the interaction of K440 with the DISW motif is important for CaChs2 function (Fig. 5c,d), the rearranged interaction of K440 with the 5-carboxyl of the uracil base inhibits CaChs2 activity (Fig. 5i). Thus, polyoxin D has an additional inhibition mechanism of disrupting the DISW motif interaction network.
Interaction of polyoxin D is different from nikkomycin Z in that i) the pyridinyl ring of nikkomycin Z reaches further into the central tunnel while polyoxin D stops at the donor binding site, ii) a different set of residues in the QRXRW motif are utilized for binding nikkomycin Z and polyoxin D, and iii) polyoxin D has an additional interaction through 5-carboxyl of uracil base. The more potent inhibition of CaChs2 by nikkomycin Z than polyoxin D likely derives from the extended interaction through the pyridinyl ring (Fig. 6b). The 5-carboxyl of polyoxin D partially compensates for the lack of pyridinyl ring interactions.
Discussion
CaChs2 forms a domain-swapped dimer with its chitin translocation tunnel controlled by the extracellular gate and a lipid molecule, and this design is important for regulating Chs functions (Fig. 4). The presence of the gate raises the question of whether the gate opening is mediated by either direct pushing by a translocating chitin polymer or allosteric control by chitin polymer occupancy in the tunnel. The lipid-dependent regulation is specific to Chs. It is well known that Chs function is highly regulated because chitin synthesis is tightly coordinated with cytokinesis and hyphal extension34,35
Ambiguity remains about the mechanism of polymer release and reinitiation of polymerization in processive GTs. Our observation of the apo structure of a catalytically active CaChs2 suggests that CaChs2 releases the polymer product and returns to the apo state before initiating synthesis of another chitin chain (Fig. 7a). The presence of the chitin-free state in catalytically active Chss is consistent with the previous observations that Chss can be partially purified by product entrapment, where Chs is bound to its insoluble chitin product under turnover conditions and released to solution in the absence of substrate19. It is not known if the apo state is also present in cellulose synthases, as purified cellulose synthases contain cellulose20.
a, Lipid modulation and drug inhibition. In the off state, the extracellular gate (blue) of the chitin translocation tunnel is closed. The tunnel is further obstructed by a PE molecule that enters from the lateral opening to prevent non-specific leakage of ions, water, and small molecules. The nascent chitin chain extends into the tunnel and expels the lipid out of the tunnel. When the chitin chain reaches the extracellular side, the gate opens, and the chain continues to elongate toward the extracellular space. TM3 and EH1 (orange triangle) of CaChs2 protrude out of the membrane, and they may provide interactions with the nascent chitin chain and facilitate the elongation. Nikkomycin Z and polyoxin D inhibit chitin synthesis by two mechanisms. First, they occupy the UDP-GlcNAc binding site competitively. Second, once bound, their N-terminal amino acid extends into the tunnel, and either partially overlaps with the acceptor binding site or obstructs chitin translocation. b, Higher order chitin assembly. When the nascent chitin chain reaches a certain length, the reducing end could fold back and form antiparallel chitin strands with the GlcNAc units near the non-reducing end. The dimeric architecture of CaChs2 can facilitate assembly of two antiparallel chitin strands, one from each monomer, into α-chitin. At Chs2-enriched regions of the plasma membrane, the α-chitin strands can further assemble into the higher-order structures of protofibril and microfibril.
The length of the central tunnel in the CaChs2 structure is shorter than those of cellulose synthases (Fig. 2c and Extended Data Fig. 7a,b). Chs enzymes produce chitin with the degree of polymerization (DOP) of 50–200 (refs. 19,36,37), which is substantially shorter than cellulose produced by cellulose synthase (DOP of 7800 in cotton and 1600 and 2900 in bacteria)38. Since the timing of termination and product release could be partly determined by the affinity of the product to the enzyme, the difference in the tunnel length between Chs and cellulose synthase could contribute to the difference in the lengths of polymer products (Fig. 2c and Extended Data Fig. 7a,b).
Chitin exists in three different forms: α-chitin has antiparallel chains, β-chitin has parallel chains, and γ-chitin is a mix of parallel and antiparallel chains. Fungal cell walls are composed of α-chitin microfibrils39. Because our recombinant CaChs2 produces α-chitin fibrils (Fig. 1b), we posit that the structural design of CaChs2 facilitates α-chitin fibril formation. As it is believed that during chitin formation, the nascent chitin chain folds back on itself to form an antiparallel chitin strand34, two nascent chitin strands in the CaChs2 dimer should be spatially separated to avoid parallel chitin strand formation. The putative chitin polymer exits are separated within the CaChs2 dimer by ~30 Å, and the extracellular extension of TM3 and EH1 near the gate could guide the nascent chitin chain to extend away from the dimer interface (Figs. 1 and 7b). This architectural arrangement will facilitate an individual chitin chain to fold back on itself and two antiparallelly folded back chitin strands to interact within the dimer, facilitating protofibril formation (Fig. 7b). CaChs2 dimers may cluster and arrange in a specific fashion in the plasma membrane so that protofibrils can mature into microfibrils40. For plant cellulose synthase, its trimer arrangement was proposed to facilitate protofilament formation21. Therefore, the architectural design of processive GTs with respect to oligomerization and the tunnel exit may provide mechanisms to control the macrostructure of the polysaccharide products.
Finally, analysis of substrate- and inhibitor-bound CaChs2 structures together with previous SAR and our functional studies reveals the differential interactions of nikkomycin Z, polyoxin D, and UDP-GlcNAc with CaChs2, and the structural plasticity of CaChs2 (Fig. 6), suggesting that there is ample room to improve the potency and/or efficacy of nikkomycin Z and polyoxin D. The antifungal efficacy of nikkomycin Z against C. albicans is moderate to low (MIC80 < 0.5–32 µg ml−1)41, largely because of the weaker effects of nikkomycin Z on CaChs1 and the requirement for the inhibition of both CaChs1 and CaChs2 to be fungicidal42. Because CaChs2 shows high sequence similarity with other Chss, including CaChs1, our studies may provide a critical platform to develop nikkomycin and polyoxin analogs to treat C. albicans and other pathogenic fungal infections.
Methods
CaChs2 protein expression and purification
The full-length CHS2 gene from C. albicans was synthesized with codon-optimization for Spodoptera frugiperda 9 (Sf9) cells and cloned into a modified pFastBac vector (Invitrogen), in frame with C-terminal PreScission protease cite, FLAG-tag, and 10x His-tag. Baculovirus was generated according to the manufacturer’s protocol (Bac-to-Bac, Invitrogen). All mutants were made on wild-type Chs2 construct by site-directed mutagenesis using KOD hot start DNA polymerase (Novagen). For CaChs2 expression, Sf9 insect cells were infected with a baculovirus at a density of 2.5–3 M cells ml−1 and grown at 27 °C for ~48 h in an orbital shaker. Cells were then harvested by centrifugation at 2,500g for 10 min at 4 °C. Cell pellets were resuspended in buffer A (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 10% glycerol, 2 mM β-mercaptoethanol (BME), 12 μg ml−1 leupeptin, 12 μg ml−1 pepstatin, 12 μg ml−1 aprotinin, 1 mM PMSF, and DNaseI) and lysed by sonication (5 × 30 pulses). All CaChs2 purification steps were performed at 4 °C. To solubilize CaChs2, 30 mM n-dodecyl-β-d-maltopyranoside (DDM, Anatrace) and 3 mM cholesteryl hemisuccinate tris salt (CHS, Anatrace) was added to the lysate and stirred at 4 °C for 1.5 h. Insoluble material was removed by centrifugation (16,000g, 30 min), and anti-FLAG M2 resin (Sigma-Aldrich) was added to the supernatant and incubated for 1 h at 4 °C. The resin was then washed with 10 column volumes of buffer B (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 10% glycerol, 0.04% glyco-diosgenin (GDN, Anatrace), 2 mM BME, 5 mM ATP-Mg), followed by 10 column volumes of buffer C (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 10% glycerol, 0.04% GDN, 2 mM BME). CaChs2 was eluted with five column volumes of elution buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 10% glycerol, 0.04% GDN, 2 mM BME, 0.2 mg ml−1 FLAG peptide (GenScript)). The FLAG and 10x His affinity tags were removed by incubation with PreScission protease at room temperature for 15 min. CaChs2 was further purified by size-exclusion chromatography (Superose 6 Increase 10/300 GL column (Cytiva)) with SEC buffer containing 20 mM HEPES pH 7.5, 150 mM NaCl, 0.015% GDN, 2 mM DTT. CaChs2 mutants were purified similarly but without tag cleavage by PreScission protease. The size-exclusion chromatography buffer for CaChs2 mutants is 20 mM HEPES pH 7.5, 150 mM NaCl, 0.015% GDN, 0.2 mM tris(2-carboxyethyl) phosphine (TCEP, Thermo Scientific). Enzymes for activity assays were flash-frozen in liquid nitrogen with the addition of 30% glycerol.
Cryo-electron microscopy sample preparation and data collection
Peak fractions containing CaChs2 from the size-exclusion chromatography were concentrated to 3–5 mg ml−1. All cryo-EM samples in this study were prepared on freshly glow-discharged UltrAuFoil R1.2/1.3 300 mesh grids (Quantifoil), using a Leica EM GP2 plunge freezer with the chamber set at 4 °C and 80% humidity. For the apo-CaChs2 sample, 3 μl of CaChs2 was applied to the grid, incubated in the chamber for 90 s, and then blotted for 1.5–2 s, followed by plunge-freezing in liquid ethane cooled by liquid nitrogen. For the CaChs2-UDP-GlcNAc sample, concentrated CaChs2 was mixed with 20 mM MgCl2 and 20 mM UDP-GlcNAc. Immediately after mixing, 3 μl of the mixture was then applied to the grid, incubated for 60 s, and blotted for 1.5–2 s, followed by plunge-freezing in liquid ethane cooled by liquid nitrogen. For the CaChs2-nikkomycin sample, concentrated CaChs2 was mixed with 5 mM MgCl2 and 4 mM nikkomycin Z on ice for 30 min. Then 3 μl of the mixture was applied to the grid, incubated for 90 s, and blotted for 1.5–2 s, followed by plunge-freezing in liquid ethane cooled by liquid nitrogen. For the CaChs2-polyoxin sample, concentrated CaChs2 was mixed with 5 mM MgCl2 and 1.5 mM polyoxin D on ice for 1 min. Then 3 μl of the mixture was applied to the grid, incubated for 90 s, and blotted for 1.5–2 s, followed by plunge-freezing in liquid ethane cooled by liquid nitrogen.
The apo-Chs2, Chs2-UDP-GlcNAc, Chs2-nikkomycin, and Chs2-polyoxin datasets were collected with a Titan Krios microscope (Thermo Fisher) operating at 300 kV and equipped with a K3 detector (Gatan) in counting mode, using Serial EM or the Latitude-S automated data-acquisition program. Movie datasets were collected at a nominal magnification of ×81,000 with a pixel size of 1.08 Å per pixel at specimen level. Each movie contained 60 frames over a 4.6-s exposure time, using a dose rate of ~15 e− pixel−1 s−1, resulting in the total accumulated dose of ~60 e−/Å2. The nominal defocus range was set from −1 to −2 μm.
Cryo-electron microscopy data processing
All datasets were processed using similar procedures. Beam-induced motion correction and dose-weighting were performed using MotionCor2 (ref. 43). The motion-corrected micrographs were then subjected to contrast transfer function (CTF) estimation using Gctf44. Micrographs were subsequently selected based on CTF fit quality and CTF estimated resolution and then imported into cryoSPARC45. An initial set of ~5,000 particles was manually picked and subjected to a reference-free two-dimensional classification, from which the classes with clear protein features were selected as reference for template-based particle picking. Picked particles were extracted with the 216-pixel box size. Iterative two-dimensional classification was performed in cryoSPARC, and classes showing clear secondary structure features of CaChs2 were selected for ab initio reconstruction with three classes. Heterogeneous refinement was performed with the models from ab initio reconstruction. The only class that shows clear protein density was selected for non-uniform refinement46. After non-uniform refinement, the particles were transferred to RELION 3.147 and subject to several rounds of Bayesian polishing and CTF refinement, which improved resolution and map quality.
Model building, refinement, and alignment
The apo-CaChs2 structure was built de novo in Coot48. During model building, residue registration was guided by the presence of large aromatic side chains. The structure was manually refined in Coot with ideal geometry restraints. The ligand-bound CaChs2 models used the apo-CaChs2 structure as a reference. The restraints for lipids and ligands, including PE, UDP-GlcNAc, nikkomycin Z, and polyoxin D, were generated in eLBOW (as implemented in Phenix49) from isomeric SMILES strings and optimized using the REEL QM2 method. Ligand restraints were then inspected and adjusted manually to ensure correct stereochemistry before being fitted into the cryo-EM maps in Coot. The MolProbity50 server was utilized to identify problematic regions in the models, which were then manually adjusted in Coot. The final refinement was performed using the phenix-real_space_refine function with global minimization and secondary structure restraints as implemented in the Phenix suite49. The Fourier shell correlation of the half- and full-maps against the model, calculated in Phenix, were in good agreement, indicating that the models were not over-refined. Structural analyses and illustrations were performed using PYMOL (Schrödinger) and UCSF Chimera and ChimeraX51. Radius calculation was performed using the HOLE program52. Structure alignments and cryo-EM density map alignments were performed using Fit-In-Map in UCSF Chimera. Based on the aligned map, each structural model was aligned to its corresponding map by Fit-In-Map.
Isothermal titration calorimetry
Protein samples were purified as described above and concentrated to around 7–10 μM. CaChs2 was in the ITC buffer that contains 20 mM HEPES pH 7.5, 150 mM NaCl, 0.015% GDN, and 0.2 mM TCEP. The ITC measurements were performed with a Nano ITC microcalorimeter (TA Instruments) at 25 °C. For CaChs2 WT and Q643A, nikkomycin Z stock at 100 μM was prepared in the same ITC buffer and injected 19 times (0.4 μl for injection 1 and 2 μl for injections 2–19), with 150–250 s intervals between injections. For CaChs2 W647A, 300 μM nikkomycin Z was used with the same titration protocol. The background data obtained from injecting nikkomycin into the ITC buffer were subtracted before the data analysis. The data were fitted using NanoAnalyze v3.11.0 (TA Instruments).
Cysteine crosslinking
Cysteine mutations are engineered and expressed as described above. After the cells were harvested, cell membrane was prepared under non-reducing conditions. Cells were resuspended in low salt buffer (10 mM Tris-HCl, pH 8.0, 10 mM NaCl, 12 μg ml−1 leupeptin, 12 μg ml−1 pepstatin, 12 μg ml−1 aprotinin, 1 mM PMSF, and DNaseI) and lysed by osmosis pressure. Cell membrane was collected by centrifugation and homogenized in high salt buffer (50 mM Tris-HCl, pH 8.0, 1 M NaCl, 12 μg ml−1 leupeptin, 12 μg ml−1 pepstatin, 12 μg ml−1 aprotinin, 1 mM PMSF, and DNaseI). Then cell membrane was collected by centrifugation and resuspended in lysis buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 10% glycerol). The membrane was first incubated with 20 mM N-ethylmaleimide for 10 min at room temperature and then purified as described above in the absence of reducing reagent and subjected to SDS–PAGE analysis.
CaChs2 activity assay
Aliquots of the flash-frozen enzyme were thawed on ice immediately before the assay. Typical reactions were performed by adding one volume of enzyme to one volume of 2x assay mixture (4 mM UDP-[U-14C]GlcNAc (specific activity ~1,000 c.p.m. nmol−1), 100 mM Tris-HCl pH 7.5, 64 mM GlcNAc, 10 mM MgCl2). Reactions were incubated at 30 °C, and at each time point, an aliquot (5 μl) was quenched in 10% TCA (w/v). The insoluble product was filtered through a glass filter and washed five times with 70:30 ethanol:1 M acetic acid in water. The radioactivity retained on the filter was analyzed on a liquid scintillation counter with 3 ml of scintillation fluid. Steady-state kinetic analyses were performed in triplicate from two biological replicates.
CaChs2 inhibition assays
CaChs2 assays were performed as described above, except that the assay contained specified concentrations of nikkomycin Z or polyoxin D. The antifungal stock solutions were prepared in 10 mM ammonium acetate pH 6.0 to prevent hydrolysis of the amide bond and were diluted five- or tenfold in the final assay mixture. Fractional activities of each reaction were plotted and fit to the four-parameter logistic equation to determine the IC50 values.
Lipid mass spectrometry
Extraction of lipids copurified with Chs2 protein
Lipids copurified with the CaChs2 protein sample were extracted using the method of Bligh and Dyer as previously described53. PBS was added to the Chs2 protein solution in a glass tube to a final volume of 1.6 ml. Subsequently, 2 ml of chloroform and 4 ml of methanol were added to make the single-phase Bligh-Dyer mixture comprising chloroform/methanol/PBS (1:2:0.8, v/v/v). This mixture solution was subjected to sonic irradiation in a bath apparatus for 5 min. The subsequent single-phase extraction mixture was then centrifuged at 500g for 10 min using a clinical centrifuge and the protein precipitate was pelleted. The supernatant was transferred to a fresh glass tube where chloroform (2 ml) and PBS (2 ml) were added to generate the two-phase Bligh-Dyer mixture composed of chloroform/methanol/PBS (2:2:1.8, v/v/v)53. After mixing and centrifugation (500g) for 10 min, the upper phase was removed, and the lower phase was dried under a nitrogen stream. The dried lipid extract was stored at −20 °C until LC–MS analysis.
Lipid identification by liquid chromatography–mass spectrometry/mass spectrometry
Normal phase LC was performed on an Agilent 1200 Quaternary LC system equipped with an Ascentis Silica HPLC column, 5 μm, 25 cm × 2.1 mm (Sigma-Aldrich, St Louis, MO) as described previously54. Mobile phase A comprised chloroform/methanol/aqueous ammonium hydroxide (800:195:5, v/v/v); mobile phase B comprised chloroform/methanol/water/aqueous ammonium hydroxide (600:340:50:5, v/v/v); mobile phase C comprised chloroform/methanol/water/aqueous ammonium hydroxide (450:450:95:5, v/v/v). The elution program was as follows: 100% mobile phase A was held for 2 min, followed by a linear increase to 100% mobile phase B over 14 min, and then held at 100% B for 11 min. The LC gradient was changed to 100% mobile phase C over 3 min, held at 100% C for 3 min, returned to 100% A over 0.5 min, and held at 100% A for 5 min (ref. 54). The LC eluent (with a total flow rate of 300 μl/min) was introduced into the ESI source of a high resolution TripleTOF5600 mass spectrometer (Sciex, Framingham, MA). Instrumental settings for negative ion ESI and MS/MS analysis of lipid species were as follows: IS = −4500 V; CUR = 20 psi; GSI = 20 psi; DP = −55 V; and FP = −150 V as described54. The MS/MS analysis used nitrogen as the collision gas. Data analysis was performed using Analyst TF1.5 software (Sciex, Framingham, MA).
Fourier-transform infrared spectroscopy
Chitin from CaChs2 was prepared as described in the CaChs2 activity assay section in a 100 μl scale with 10 mM UDP-GlcNAc for 20 h. The cloudy reaction mixture was quenched by adding 100 μl of a 5% (w/v) SDS solution, and the insoluble chitin was collected by centrifuging for 15 minutes at 20,000g. The pellet was washed with 5 × 200 μl of 2% (w/v) SDS, water, and ethanol. The pellet was then dried by speedvac for 30 min. The IR spectra were collected (128 scans, 4 cm−1 resolution) on a Perkin-Elmer Frontier instrument with a Pike MIRacle attenuated total reflectance (ATR) optic and a Mercury-Cadmium-Telluride (MCT) detector. Before collecting sample spectra, baseline scans of air were taken and subtracted automatically from the sample spectra in the Spectrum software (Perkin-Elmer). Sample spectra were collected by placing the dried chitin samples directly on the ATR crystal.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The coordinates are deposited in the Protein Data Bank with the PDB IDs 7STL (apo), 7STM (UDP-GlcNAc-bound), 7STN (nikkomycin Z-bound state) and 7STO (polyoxin D-bound state), respectively. The cryo-EM density maps are deposited in EMDB with the IDs EMD-25432 (apo), EMD-25433 (UDP-GlcNAc-bound), EMD-25434 (nikkomycin Z-bound state) and EMD-25435 (polyoxin D-bound state), respectively. Source data are provided with this paper.
References
Tharanathan, R. N. & Kittur, F. S. Chitin–the undisputed biomolecule of great potential. Crit. Rev. Food Sci. Nutr. 43, 61–87 (2003).
Steinfeld, L., Vafaei, A., Rosner, J. & Merzendorfer, H. Chitin prevalence and function in bacteria, fungi and protists. Adv. Exp. Med. Biol. 1142, 19–59 (2019).
Mahmoud, Y. A., El-Naggar, M. E., Abdel-Megeed, A. & El-Newehy, M. Recent advancements in microbial polysaccharides: synthesis and applications. Polymers (Basel) 13, 4136 (2021).
Mikusova, V. & Mikus, P. Advances in chitosan-based nanoparticles for drug delivery. Int. J. Mol. Sci. 22, 5776 (2021).
Bhatt, K. et al. High mortality co-infections of COVID-19 patients: mucormycosis and other fungal infections. Discoveries (Craiova) 9, e126 (2021).
Gangneux, J. P., Bougnoux, M. E., Dannaoui, E., Cornet, M. & Zahar, J. R. Invasive fungal diseases during COVID-19: We should be prepared. J. Mycol. Med. 30, 100971 (2020).
Marr, K. A. et al. Aspergillosis complicating severe coronavirus disease. Emerg. Infect. Dis. 27, 18–25 (2021).
Fisher, M. C. et al. Emerging fungal threats to animal, plant and ecosystem health. Nature 484, 186–194 (2012).
Lima, S. L., Colombo, A. L. & de Almeida Junior, J. N. Fungal cell wall: emerging antifungals and drug resistance. Front. Microbiol. 10, 2573 (2019).
Roemer, T. & Krysan, D. J. Antifungal drug development: challenges, unmet clinical needs, and new approaches. Cold. Spring Harb. Perspect. Med. 4, a019703 (2014).
Coutinho, P. M., Deleury, E., Davies, G. J. & Henrissat, B. An evolving hierarchical family classification for glycosyltransferases. J. Mol. Biol. 328, 307–317 (2003).
Dorfmueller, H. C., Ferenbach, A. T., Borodkin, V. S. & van Aalten, D. M. F. A structural and biochemical model of processive chitin synthesis. J. Biol. Chem. 289, 23020–23028 (2014).
Orlean, P. & Funai, D. Priming and elongation of chitin chains: Implications for chitin synthase mechanism. Cell Surf. 5, 100017 (2019).
Osada, H. Discovery and applications of nucleoside antibiotics beyond polyoxin. J. Antibiot. (Tokyo) 72, 855–864 (2019).
Nix, D. E., Swezey, R. R., Hector, R. & Galgiani, J. N. Pharmacokinetics of nikkomycin Z after single rising oral doses. Antimicrob. Agents Chemother. 53, 2517–2521 (2009).
Shubitz, L. F., Roy, M. E., Nix, D. E. & Galgiani, J. N. Efficacy of Nikkomycin Z for respiratory coccidioidomycosis in naturally infected dogs. Med. Mycol. 51, 747–754 (2013).
Draelos, M. M. & Yokoyama, K. in Comprehensive Natural Products III (eds Liu, H.-W. & Begley, T. P.) 613–641 (Elsevier, 2020).
Kim, M. K., Park, H. S., Kim, C. H., Park, H. M. & Choi, W. Inhibitory effect of nikkomycin Z on chitin synthases in Candida albicans. Yeast 19, 341–349 (2002).
Kang, M. S. et al. Isolation of chitin synthetase from Saccharomyces cerevisiae. Purification of an enzyme by entrapment in the reaction product. J. Biol. Chem. 259, 14966–14972 (1984).
Morgan, J. L., Strumillo, J. & Zimmer, J. Crystallographic snapshot of cellulose synthesis and membrane translocation. Nature 493, 181–186 (2013).
Purushotham, P., Ho, R. & Zimmer, J. Architecture of a catalytically active homotrimeric plant cellulose synthase complex. Science 369, 1089–1094 (2020).
Martinez-Rucobo, F. W., Eckhardt-Strelau, L. & Terwisscha van Scheltinga, A. C. Yeast chitin synthase 2 activity is modulated by proteolysis and phosphorylation. Biochem. J. 417, 547–554 (2009).
Uchida, Y., Shimmi, O., Sudoh, M., Arisawa, M. & Yamada-Okabe, H. Characterization of chitin synthase 2 of Saccharomyces cerevisiae. II: Both full size and processed enzymes are active for chitin synthesis. J. Biochem. 119, 659–666 (1996).
Nagahashi, S. et al. Characterization of chitin synthase 2 of Saccharomyces cerevisiae. Implication of two highly conserved domains as possible catalytic sites. J. Biol. Chem. 270, 13961–13967 (1995).
Morgan, J. L. et al. Observing cellulose biosynthesis and membrane translocation in crystallo. Nature 531, 329–334 (2016).
Cos, T. et al. Molecular analysis of Chs3p participation in chitin synthase III activity. Eur. J. Biochem. 256, 419–426 (1998).
Yabe, T. et al. Mutational analysis of chitin synthase 2 of Saccharomyces cerevisiae. Identification of additional amino acid residues involved in its catalytic activity. Eur. J. Biochem. 258, 941–947 (1998).
Hori, M., Kakiki, K. & Misato, T. Further study on the relation of polyoxin structure to chitin synthetase inhibition. Agric. Biol. Chem. 38, 691–698 (1974).
Jackson, K. E., Pogula, P. K. & Patterson, S. E. Polyoxin and nikkomycin analogs: recent design and synthesis of novel peptidyl nucleosides. Heterocycl. Commun. 19, 375–386 (2013).
Decker, H., Zahner, H., Heitsch, H., Konig, W. A. & Fiedler, H. P. Structure-activity relationships of the nikkomycins. J. Gen. Microbiol. 137, 1805–1813 (1991).
Hori, M., Kakiki, K., Suzuki, S. & Misato, T. Studies on the mode of action of polyoxins: Part III. Relation of polyoxin structure to chitin synthetase inhibition. Agric. Biol. Chem. 35, 1280–1291 (1971).
Shenbagamurthi, P., Smith, H. A., Becker, J. M., Steinfeld, A. & Naider, F. Design of anticandidal agents: synthesis and biological properties of analogs of polyoxin L. J. Med. Chem. 26, 1518–1522 (1983).
Khare, R. K., Becker, J. M. & Naider, F. R. Synthesis and anticandidal properties of polyoxin L analogs containing.alpha.-amino fatty acids. J. Med. Chem. 31, 650–656 (1988).
Lenardon, M. D., Munro, C. A. & Gow, N. A. Chitin synthesis and fungal pathogenesis. Curr. Opin. Microbiol. 13, 416–423 (2010).
Gooday, G. W. & Schofield, D. A. Regulation of chitin synthesis during growth of fungal hyphae: the possible participation of membrane stress. Can. J. Bot. 73, 114–121 (1995).
Orlean, P. Two chitin synthases in Saccharomyces cerevisiae. J. Biol. Chem. 262, 5732–5739 (1987).
Grabinska, K. A., Magnelli, P. & Robbins, P. W. Prenylation of Saccharomyces cerevisiae Chs4p affects chitin synthase III activity and chitin chain length. Eukaryot. Cell 6, 328–336 (2007).
McManus, J. B., Yang, H., Wilson, L., Kubicki, J. D. & Tien, M. Initiation, elongation, and termination of bacterial cellulose synthesis. ACS Omega 3, 2690–2698 (2018).
Lenardon, M. D., Whitton, R. K., Munro, C. A., Marshall, D. & Gow, N. A. Individual chitin synthase enzymes synthesize microfibrils of differing structure at specific locations in the Candida albicans cell wall. Mol. Microbiol. 66, 1164–1173 (2007).
Vermeulen, C. A. & Wessels, J. G. Chitin biosynthesis by a fungal membrane preparation. Evidence for a transient non-crystalline state of chitin. Eur. J. Biochem. 158, 411–415 (1986).
Li, R. K. & Rinaldi, M. G. In vitro antifungal activity of nikkomycin Z in combination with fluconazole or itraconazole. Antimicrob. Agents Chemother. 43, 1401–1405 (1999).
Sudoh, M. et al. Identification of a novel inhibitor specific to the fungal chitin synthase. Inhibition of chitin synthase 1 arrests the cell growth, but inhibition of chitin synthase 1 and 2 is lethal in the pathogenic fungus Candida albicans. J. Biol. Chem. 275, 32901–32905 (2000).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K., Gctf & Real-time, C. T. F. determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7 (2018).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Goddard, T. D., Huang, C. C. & Ferrin, T. E. Visualizing density maps with UCSF Chimera. J. Struct. Biol. 157, 281–287 (2007).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–60, 376 (1996).
Haig, N. A. et al. Identification of self-lipids presented by CD1c and CD1d proteins. J. Biol. Chem. 286, 37692–37701 (2011).
Joyce, L. R. et al. Identification of a novel cationic glycolipid in Streptococcus agalactiae that contributes to brain entry and meningitis. PLoS Biol. 20, e3001555 (2022).
Acknowledgments
Cryo-EM data were screened and collected at the Duke University Shared Materials Instrumentation Facility (SMIF), the Pacific Northwest Center for Cryo-EM (PNCC) in OHSU, and the UNC CryoEM core facility. We thank N. Bhattacharya at SMIF, J. Myers at PNCC, and J. Peck and J. Strauss of the UNC CryoEM Core Facility for assistance with the microscope operation. We thank A. Stelling for assistance with the FTIR measurements, M. M. Draelos for preparation of nikkomycin Z and J. Fedor for his critical reading of the manuscript. This research was supported by a National Institutes of Health grant R01GM115729 (K.Y) and Duke startup funds (S.-Y.L). A portion of this research was supported by NIH grant U24GM129547 and performed at the PNCC at OHSU and accessed through EMSL (grid.436923.9), a DOE Office of Science User Facility sponsored by the Office of Biological and Environmental Research. Duke SMIF is affiliated with the North Carolina Research Triangle Nanotechnology Network, which is in part supported by the NSF (ECCS-2025064). The UNC CryoEM core facility is supported by NIH grant P30CA016086.
Author information
Authors and Affiliations
Contributions
Z.R. conducted biochemical preparation, sample freezing, single-particle 3D reconstruction of the enzyme, crosslinking experiment and ITC experiments under the guidance of S.-Y.L. A.C. carried out enzymatic assays, and the FTIR measurement under the guidance of K.Y. Y.S. and Z.R. collected cryo-EM data. Z.R. and S.-Y.L. performed model building and refinement. Z.G. performed the lipid MS analysis. S.-Y.L, Z.R., K.Y. and A.C. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Purification and enzymatic reaction of CaChs2 and isothermal titration calorimetry (ITC) analysis.
a, Representative gel-filtration profile and SDS-PAGE of purified CaChs2. The experiments have been repeated more than ten times independently with essentially the same results. b, Enzymatic reaction of CaChs2. c-e. Representative ITC data of CaChs2 WT (c), W647A (d) and Q643A (e) vs Nikkomycin. The Kd is the average of three independent repeats. The uncropped gel image for a is available as source data.
Extended Data Fig. 2 Cryo-EM analysis of CaChs2 in apo, UDP-GlcNAc bound, nikkomycin Z bound, and polyoxin D bound states.
a, General flowchart for data processing. b, Representative micrographs for each dataset. The number of total micrographs in each dataset is shown in Extended Data Fig.3. c, Representative 2D classes. d, Local resolution maps. e, Euler angle distribution of the final reconstructions. f, The gold-standard Fourier shell correlation (FSC) curves for the final reconstructions. g, FSC curves of the refined models versus the corresponding full map (red) and half maps (green and blue).
Extended Data Fig. 3 Data processing flowchart.
Data processing for CaChs2 in apo (a), UDP-GlcNAc bound (b), nikkomycin Z bound (c), and polyoxin D bound (d) states.
Extended Data Fig. 4 Representative EM densities.
Representative densities for CaChs2 in apo (a), UDP-GlcNAc bound (b), nikkomycin Z bound (c), and polyoxin D bound (d) states.
Extended Data Fig. 5 Comparison of architectures of chitin synthase CaChs2 and cellulose synthases RsBcsA and PttCesA8.
a, Domain arrangement of CaChs2, RsBcsA and PttCesA8. b-c, Topology diagrams of RsBcsA (b) and PttCesA8 (c).
Extended Data Fig. 6 The dimer interface of CaChs2.
a, Cartoon representation of a CaChs2 dimer. The dimer interface is highlighted in the rectangle. b-c, Close-up view of the dimer interface in two orientations.
Extended Data Fig. 7 Structural comparison of polymer translocation tunnels and substrate binding sites of CaChs2, RsBcsA and PttCesA8.
a, Left: Ribbon representation of RsBcsA in the product bound state (PDB code: 5EJZ) with the tunnel highlighted in surface representation and the cellulose polymer in sticks. Trp383, which defines the intracellular entry of the tunnel is highlighted in sticks. Right: Length and radius of the tunnel calculated by the HOLE program. b. Ribbon representation of PttCesA (PDB code: 6WLB) with the tunnel highlighted in surface representation and the cellulose polymer in sticks. Trp718, which defines the intracellular entry of the tunnel is highlighted in sticks. Right: Length and radius of the tunnel calculated by the HOLE program. c-d. Overlay of the substrate binding sites of CaChs2 (green) and substrate-bound RsBcsA (orange) (PDB code: 5EIY). Conserved residues that interact with the substrates and the catalytic residues are shown as sticks. e-f. Overlay of binding sites of CaChs2 (green) and product-bound PttCesA8 (purple) (PDB code: 6WLB). Conserved residues that interact with the substrates and the catalytic residues are shown as sticks.
Extended Data Fig. 8 Conservation mapping of CaChs2 and EM density of UDP-GlcNAc and Mg2+ at the substrate binding site.
a, Cross-section surface representation of a CaChs2 monomer bound to nikkomycin Z (yellow spheres). b, Detailed interactions between CaChs2 and nikkomycin Z. In both a and b, CaChs2 is colored based on the conservation score calculated by the Consurf Server (https://consurf.tau.ac.il/) from 150 Chs homologs whose sequence identity varies from 35% to 99%. Conservation scores: 1, ~15% conserved; 5, ~60% conserved; 9, 100% conserved. c-d, Density (Threshold = 0.007) of within 6 Å of UDP-GlcNAc, Mg2+, and residues interacting with the substrate, viewed from two orientations, is shown as grey surfaces. The protein model is shown as cartoon and the substrate and residues at the binding site are shown as sticks. The map and model of residues obstructing the view are hidden for clarity.
Extended Data Fig. 9 Identification of lipids copurified with CaChs2 by LC/MS.
a, Total ion chromatogram (TIC) of normal phase LC/MS of lipids extracted from the CaChs2 protein sample. b, Representative chemical structure of ceramide and mass spectrum of the 2.88 min TIC peak in a showing the chloride adduct [M + Cl]- ions of ceramide species. c, Representative chemical structure of and mass spectrum of the 16.20 min TIC peak in a showing the [M-H]- ions of PE molecular species. This identification of ceramide and PE was supported by exact mass measurement and MS/MS. The TIC peaks at 14.09, 21.16 and 27.18 min are consistent with glycodiosgenins (GDN) containing two, three and four sugars whose [M + Cl]- ions are observed at m/z 875.5, 1037.5 and 1199.5, respectively.
Extended Data Fig. 10 Map and model of the domain-swapped region and crosslinking experiments to validate the domain-swapped assembly.
a-b, Unsharpened map of the domain-swapped region of UDP-GlcNAc bound CaChs2 shown at threshold 0.00475, viewed from two orientations. c. Representative density of the domain-swapped region. The protein model is shown as cartoon and side chains are shown as sticks. Some of the side chains are not built due to lack of signal. d. CaChs2 viewed from the intracellular side. The Cα atom of P906 and N917, which define the N and C-terminus of the loop between H10 and TM5, are shown as spheres. The distances from the Cα of P906 to that of N917 within the same monomer or in different monomers are labeled with dashed lines. It is noteworthy that the distance between P906 and N917 is closer in the domain swapped arrangement (30 Å) than the non-domain swapped arrangement (46.5 Å). Because the length of an amino acid in a linear chain is ~3.5 Å, non-domain swapped configuration is physically impossible. e. Model of CaChs2 dimer with one protomer in blue and the other in salmon. Cα atoms of residues where cysteine was introduced are shown as spheres. The crosslinking cysteine pairs are labeled with dashed lines. f. SDS-PAGE analysis of single and double cysteine mutants of CaChs2. The experiment has been repeated four times from two biological repeats with essentially the same results. Uncropped gel images for f are available as source data.
Supplementary information
Supplementary Information
Supplementary Fig. 1 and Supplementary Table 1
Supplementary Data 1
Source data for Supplementary Table 1.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 1
Uncropped gel for 1a.
Source Data Extended Data Fig. 10
Uncropped gels for 10f.
Rights and permissions
About this article
Cite this article
Ren, Z., Chhetri, A., Guan, Z. et al. Structural basis for inhibition and regulation of a chitin synthase from Candida albicans. Nat Struct Mol Biol 29, 653–664 (2022). https://doi.org/10.1038/s41594-022-00791-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-022-00791-x
This article is cited by
-
UV-activated green-synthesized ZnO NPs from Camellia sinensis extract: a potent antimicrobial strategy
Chemical Papers (2024)
-
Curcumin interferes with chitin synthesis in Aedes aegypti: a computational and experimental investigation
Molecular Diversity (2024)
-
The N-terminal disordered region of ChsB regulates its efficient transport to the hyphal apical surface in Aspergillus nidulans
Current Genetics (2023)
-
Structural basis for directional chitin biosynthesis
Nature (2022)