Abstract
Trim-Away is a recently developed technology that exploits off-the-shelf antibodies and the RING E3 ligase and cytosolic antibody receptor TRIM21 to carry out rapid protein depletion. How TRIM21 is catalytically activated upon target engagement, either during its normal immune function or when repurposed for targeted protein degradation, is unknown. Here we show that a mechanism of target-induced clustering triggers intermolecular dimerization of the RING domain to switch on the ubiquitination activity of TRIM21 and induce virus neutralization or drive Trim-Away. We harness this mechanism for selective degradation of disease-causing huntingtin protein containing long polyglutamine tracts and expand the Trim-Away toolbox with highly active TRIM21–nanobody chimeras that can also be controlled optogenetically. This work provides a mechanism for cellular activation of TRIM RING ligases and has implications for targeted protein degradation technologies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
SAXS data are available upon request. All other data generated or analyzed during this study are included in this published article and associated files.
References
Zheng, N. & Shabek, N. Ubiquitin ligases: structure, function, and regulation. Annu. Rev. Biochem. 86, 129–157 (2017).
Schapira, M., Calabrese, M. F., Bullock, A. N. & Crews, C. M. Targeted protein degradation: expanding the toolbox. Nat. Rev. Drug Discov. 18, 949–963 (2019).
Yamada, T., Yang, Y. & Bonni, A. Spatial organization of ubiquitin ligase pathways orchestrates neuronal connectivity. Trends Neurosci. 36, 218–226 (2013).
Verma, R., Mohl, D. & Deshaies, R. J. Harnessing the power of proteolysis for targeted protein inactivation. Mol. Cell 77, 446–460 (2020).
Wu, T. et al. Targeted protein degradation as a powerful research tool in basic biology and drug target discovery. Nat. Struct. Mol. Biol. 27, 605–614 (2020).
James, L. C., Keeble, A. H., Khan, Z., Rhodes, D. A. & Trowsdale, J. Structural basis for PRYSPRY-mediated tripartite motif (TRIM) protein function. Proc. Natl Acad. Sci. USA 104, 6200–6205 (2007).
Mallery, D. L. et al. Antibodies mediate intracellular immunity through tripartite motif-containing 21 (TRIM21). Proc. Natl Acad. Sci. USA 107, 19985–19990 (2010).
Vaysburd, M. et al. Intracellular antibody receptor TRIM21 prevents fatal viral infection. Proc. Natl Acad. Sci. USA 110, 12397–12401 (2013).
Fan, W. et al. Swine TRIM21 restricts FMDV infection via an intracellular neutralization mechanism. Antiviral Res. 127, 32–40 (2016).
Caddy, S. L. et al. Intracellular neutralisation of rotavirus by VP6-specific IgG. PLoS Pathog. 16, e1008732 (2020).
Caddy, S. L. et al. Viral nucleoprotein antibodies activate TRIM21 and induce T cell immunity. EMBO J. https://doi.org/10.15252/embj.2020106228 (2020).
Rakebrandt, N., Lentes, S., Neumann, H., James, L. C. & Neumann-Staubitz, P. Antibody- and TRIM21-dependent intracellular restriction of Salmonella enterica. Pathog. Dis. 72, 131–137 (2014).
McEwan, W. A. et al. Intracellular antibody-bound pathogens stimulate immune signaling via the Fc receptor TRIM21. Nat. Immunol. 14, 327–336 (2013).
McEwan, W. A. et al. Cytosolic Fc receptor TRIM21 inhibits seeded tau aggregation. Proc. Natl Acad. Sci. USA 114, 574–579 (2017).
Foss, S. et al. TRIM21—from intracellular immunity to therapy. Front. Immunol. 10, 2049 (2019).
McEwan, W. A., Mallery, D. L., Rhodes, D. A., Trowsdale, J. & James, L. C. Intracellular antibody-mediated immunity and the role of TRIM21. Bioessays 33, 803–809 (2011).
Clift, D. et al. A method for the acute and rapid degradation of endogenous proteins. Cell 171, 1692–1706 (2017).
Clift, D., So, C., McEwan, W. A., James, L. C. & Schuh, M. Acute and rapid degradation of endogenous proteins by Trim-Away. Nat. Protoc. 13, 2149–2175 (2018).
Fletcher, A. J., Mallery, D. L., Watkinson, R. E., Dickson, C. F. & James, L. C. Sequential ubiquitination and deubiquitination enzymes synchronize the dual sensor and effector functions of TRIM21. Proc. Natl Acad. Sci. USA 112, 10014–10019 (2015).
Kiss, L. et al. A tri-ionic anchor mechanism drives Ube2N-specific recruitment and K63-chain ubiquitination in TRIM ligases. Nat. Commun. 10, 4502 (2019).
de Bie, P. & Ciechanover, A. Ubiquitination of E3 ligases: self-regulation of the ubiquitin system via proteolytic and non-proteolytic mechanisms. Cell Death Differ. 18, 1393–1402 (2011).
Peth, A., Uchiki, T. & Goldberg, A. L. ATP-dependent steps in the binding of ubiquitin conjugates to the 26S proteasome that commit to degradation. Mol. Cell 40, 671–681 (2010).
Hofmann, R. M. & Pickart, C. M. In vitro assembly and recognition of Lys-63 polyubiquitin chains. J. Biol. Chem. 276, 27936–27943 (2001).
Saeki, Y. et al. Lysine 63-linked polyubiquitin chain may serve as a targeting signal for the 26S proteasome. EMBO J. 28, 359–371 (2009).
Kim, H. T. et al. Certain pairs of ubiquitin-conjugating enzymes (E2s) and ubiquitin-protein ligases (E3s) synthesize nondegradable forked ubiquitin chains containing all possible isopeptide linkages. J. Biol. Chem. 282, 17375–17386 (2007).
Ohtake, F., Tsuchiya, H., Saeki, Y. & Tanaka, K. K63 ubiquitylation triggers proteasomal degradation by seeding branched ubiquitin chains. Proc. Natl Acad. Sci. USA 115, E1401–E1408 (2018).
Fletcher, A. J. et al. TRIM5α requires Ube2W to anchor Lys63-linked ubiquitin chains and restrict reverse transcription. EMBO J. 34, 2078–2095 (2015).
Fletcher, A. J. et al. Trivalent RING assembly on retroviral capsids activates TRIM5 ubiquitination and innate immune signaling. Cell Host Microbe 24, 761–775 (2018).
Fiorentini, F., Esposito, D. & Rittinger, K. Does it take two to tango? RING domain self-association and activity in TRIM E3 ubiquitin ligases. Biochem. Soc. Trans. 48, 2615–2624 (2020).
Koliopoulos, M. G., Esposito, D., Christodoulou, E., Taylor, I. A. & Rittinger, K. Functional role of TRIM E3 ligase oligomerization and regulation of catalytic activity. EMBO J. 35, 1204–1218 (2016).
Haubrich, K. et al. RNA binding regulates TRIM25-mediated RIG-I ubiquitylation. Preprint at bioRxiv https://doi.org/10.1101/2020.05.04.070177 (2020).
Sanchez, J. G. et al. TRIM25 binds RNA to modulate cellular anti-viral defense. J. Mol. Biol. 430, 5280–5293 (2018).
Diaz-Griffero, F. et al. Rapid turnover and polyubiquitylation of the retroviral restriction factor TRIM5. Virology 349, 300–315 (2006).
Li, X. & Sodroski, J. The TRIM5α B-box 2 domain promotes cooperative binding to the retroviral capsid by mediating higher-order self-association. J. Virol. 82, 11495–11502 (2008).
Yudina, Z. et al. RING dimerization links higher-order assembly of TRIM5α to synthesis of K63-linked polyubiquitin. Cell Rep. 12, 788–797 (2015).
Ganser-Pornillos, B. K. et al. Hexagonal assembly of a restricting TRIM5α protein. Proc. Natl Acad. Sci. USA 108, 534–539 (2011).
Li, Y.-L. et al. Primate TRIM5 proteins form hexagonal nets on HIV-1 capsids. Elife 5, e16269 (2016).
Dickson, C. et al. Intracellular antibody signalling is regulated by phosphorylation of the Fc receptor TRIM21. Elife 7, e32660 (2018).
McEwan, W. A. et al. Regulation of virus neutralization and the persistent fraction by TRIM21. J. Virol. 86, 8482–8491 (2012).
Hilpert, K. et al. Anti-c-myc antibody 9E10: epitope key positions and variability characterized using peptide spot synthesis on cellulose. Protein Eng. Des. Sel. 14, 803–806 (2001).
Krauss, N. et al. The structure of the anti-c-myc antibody 9E10 Fab fragment/epitope peptide complex reveals a novel binding mode dominated by the heavy chain hypervariable loops. Proteins 73, 552–565 (2008).
Young, G. et al. Quantitative mass imaging of single biological macromolecules. Science 360, 423–427 (2018).
Cole, D., Young, G., Weigel, A., Sebesta, A. & Kukura, P. Label-free single-molecule imaging with numerical-aperture-shaped interferometric scattering microscopy. ACS Photonics 4, 211–216 (2017).
Kubala, M. H., Kovtun, O., Alexandrov, K. & Collins, B. M. Structural and thermodynamic analysis of the GFP:GFP-nanobody complex. Protein Sci. 19, 2389–2401 (2010).
Gillingham, A. K. & Munro, S. The PACT domain, a conserved centrosomal targeting motif in the coiled‐coil proteins AKAP450 and pericentrin. EMBO Rep. 1, 524–529 (2000).
Shvets, E., Bitsikas, V., Howard, G., Hansen, C. G. & Nichols, B. J. Dynamic caveolae exclude bulk membrane proteins and are required for sorting of excess glycosphingolipids. Nat. Commun. 6, 6867 (2015).
Hayer, A., Stoeber, M., Bissig, C. & Helenius, A. Biogenesis of caveolae: stepwise assembly of large caveolin and cavin complexes. Traffic 11, 361–382 (2010).
Bates, G. P. et al. Huntington disease. Nat. Rev. Dis. Primers 1, 15005 (2015).
Owens, G. E., New, D. M., West, A. P. & Bjorkman, P. J. Anti-polyQ antibodies recognize a short polyQ stretch in both normal and mutant huntingtin exon 1. J. Mol. Biol. 427, 2507–2519 (2015).
Bugaj, L. J., Choksi, A. T., Mesuda, C. K., Kane, R. S. & Schaffer, D. V. Optogenetic protein clustering and signaling activation in mammalian cells. Nat. Methods 10, 249–252 (2013).
Taslimi, A. et al. An optimized optogenetic clustering tool for probing protein interaction and function. Nat. Commun. 5, 4925 (2014).
Park, H. et al. Optogenetic protein clustering through fluorescent protein tagging and extension of CRY2. Nat. Commun. 8, 30 (2017).
Kono, K. et al. Reconstruction of Par-dependent polarity in apolar cells reveals a dynamic process of cortical polarization. Elife 8, e45559 (2019).
La Spada, A. R. & Taylor, J. P. Repeat expansion disease: progress and puzzles in disease pathogenesis. Nat. Rev. Genet. 11, 247–258 (2010).
Soto, C. & Pritzkow, S. Protein misfolding, aggregation, and conformational strains in neurodegenerative diseases. Nat. Neurosci. 21, 1332–1340 (2018).
Ganser-Pornillos, B. K. & Pornillos, O. Restriction of HIV-1 and other retroviruses by TRIM5. Nat. Rev. Microbiol. 17, 546–556 (2019).
Mund, T. & Pelham, H. R. Substrate clustering potently regulates the activity of WW-HECT domain-containing ubiquitin ligases. J. Biol. Chem. 293, 5200–5209 (2018).
Słabicki, M. et al. Small-molecule-induced polymerization triggers degradation of BCL6. Nature 588, 164–168 (2020).
Zeng, J., Slodkowicz, G. & James, L. C. Rare missense variants in the human cytosolic antibody receptor preserve antiviral function. Elife 8, e48339 (2019).
Liman, E. R., Tytgat, J. & Hess, P. Subunit stoichiometry of a mammalian K+ channel determined by construction of multimeric cDNAs. Neuron 9, 861–871 (1992).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Schuck, P. On the analysis of protein self-association by sedimentation velocity analytical ultracentrifugation. Anal. Biochem. 320, 104–124 (2003).
Brautigam, C. A. in Methods in Enzymology Vol. 562 (ed. Cole, J. L.) 109–133 (Academic Press, 2015).
Acknowledgements
We thank J. T. Andersson (University of Oslo) for the 9C12 antibody variants and members of the James laboratory for helpful discussions. J.Z. was supported by a PhD Studentship from the Rosetrees Trust and the Frank Edward Elmore Fund (University of Cambridge). M.O. is part of the GABBA PhD program from the University of Porto. D.A.J. was supported by an NHMRC Early Career Fellowship (CJ Martin) (GNT1036521). C.F.D. was supported by an NHMRC Early Career Fellowship (GNT1110116). L.K. is supported by a PhD fellowship from the Boehringer Ingelheim Fonds. W.A.M. is a Lister Institute Prize Fellow and is supported by a Sir Henry Dale Fellowship jointly funded by the Wellcome Trust and the Royal Society (grant number 206248/Z/17/Z) and is supported by the UK Dementia Research Institute, which receives its funding from UK DRI Ltd, funded by the UK Medical Research Council, Alzheimer’s Society and Alzheimer’s Research UK. E.M.-d.-S. acknowledges FCT (Fundação para a Ciência e a Tecnologia) (contract CEECIND/00622/2017 and grant PTDC/BEX-BCM/0432/2014) and project Norte-01-0145-FEDER-000029, supported by NORTE 2020. This work was supported by the MRC (UK, U105181010) and a Wellcome Trust Investigator Award (200594/Z/16/Z).
Author information
Authors and Affiliations
Contributions
D.C. and L.C.J. conceptualized the study; J.Z., M.O., W.A.M., E.M.-d.-S., D.C. and L.C.J. developed the methodology; J.Z., A.F.S., A.S.M., M.O., D.A.J., C.F.D., S.H.M., C.M.J., L.K., J.L., N.R., M.V., W.A.M. and D.C. performed the experiments; D.C. and L.C.J. prepared the manuscript; J.Z., A.F.S., M.O., W.A.M., E.M.-d.-S., D.C. and L.C.J. revised and edited the manuscript; W.A.M., E.M.-d.-S., D.C. and L.C.J. supervised the work; W.A.M., E.M.-d.-S. and L.C.J. acquired funding.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Structural & Molecular Biology thanks Alessio Ciulli and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Inês Chen was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 TRIM21 autoubiquitination upon substrate engagement.
a, His-Ubiquitin pulldown from cells expressing the indicated TRIM21 mutants infected with AdV5 ± 9C12 or 9C12(H433A). b-d, TRIM21-His is expressed at endogenous TRIM21 levels (b) and is functional for AdV5 neutralisation (c) and NFκB signalling (d). Graphs in c and d show mean and s.e.m. of n = 3 technical replicates. e, TRIM21-His pulldown from cells infected with AdV5 ± 9C12 or 9C12(H433A). The AdV5 + 9C12 condition was split and one half treated with the deubiquitinase USP2. f, co-crystal structure of TRIM21 RING:Ube2N~ubiquitin complex (PDB: 6S53) showing the tri-anionic anchor motif (E12, E13 and D21) and second site residues (R67 and N71). g, His-Ubiquitin pulldown from cells expressing the indicated TRIM21 mutants infected with AdV5 ± 9C12 or 9C12(H433A). h, His-Ubiquitin pulldown from cells expressing the indicated His-Ubiquitin mutants infected with AdV5 ± 9C12 or 9C12(H433A). *T21 in immunoblots shown on top of panels a, e, g and h shows non-specific binding of unmodified TRIM21 to the NiNTA beads. To ensure equal His-Ubiquitin pulldown between conditions within a single experiment, equal amounts of His-Ubiquitin plasmids were transfected side-by-side for each condition, and His-pulldowns were performed side-by-side under identical conditions (see also Methods). Uncropped blots/membranes are in Supplementary Data 2. Source data for graphs are in Supplementary Data 1.
Extended Data Fig. 2 Further characterisation of TRIM21 RING domain mutants.
a,b, RING dimerisation mutants M10E and M72E are expressed at WT levels (a) and do not affect AdV5 infection efficiency (b). c, His-Ubiquitin pulldown from cells expressing the indicated TRIM21 mutants infected with AdV5 ± 9C12 or 9C12(H433A). *T21 shows non-specific binding of unmodified TRIM21 to the NiNTA beads. e,f, Second site mutants R67A and N71D are expressed at WT levels (e) and do not affect AdV5 infection efficiency (f). g, Neutralisation of AdV5 infection by increasing 9C12 concentrations in cells expressing the indicated TRIM21 mutants. h, AdV5-9C12-induced NFkB activation in cells expressing the indicated TRIM21 mutants. i, Trim-away of target protein IKKα following electroporation (+) of anti-IKKα IgG (anti-IKKα). Immunoblots show IKKα degradation is deficient in cells expressing the N71D mutant TRIM21. Graphs in b,f,g,h show mean and s.e.m for n = 3 independent experiments. Statistical significance is based on one-way ANOVA (b,f,h) and two-way ANOVA (g) and represented with symbols: ns(P>0.05), *(P≤0.05), **(P≤0.01), ***(P≤0.001), ****(P≤0.0001). Uncropped blots/membranes are in Supplementary Data 2. Source data for graphs and statistics are in Supplementary Data 1.
Extended Data Fig. 3 Structural and biophysical characterisation of TRIM21.
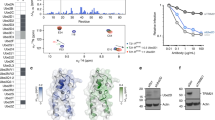
a, Schematic of TRIM21 protein sequence indicating RING, B Box, coiled-coil and PRYSPRY domain boundaries. b, SEC-MALS chromatograms of T21CC235 loaded at concentrations of 15 (black), 5 (green), 0.55 (blue) and 0.25 mg/ml (red) in which the refractive index is indicated by the solid lines while the molar mass evaluated from the light scattering analysis is indicated with the corresponding coloured dotted lines. c, Isothermal titration calorimetry (ITC) trace of T21CC235 fitted to a dimer dissociation model reveal an enthalpy of 65 kcal/mol and Kd of 7 µM. d, Analytical ultracentrifugation (AUC) sedimentation velocity analysis of T21RBCC in solution. The upper panel shows absorbance profiles with best fits of a c(s) model (coloured lines) and their residuals to the fits underneath. The different colours represent scans at different times: blue is the earliest time points where very little material has sedimented; through to red where all the material has sedimented and the signal is near baseline across the radius. The lower panel shows the c(s) distribution of specie. T21RBCC sedimented with coefficient of 2.5–2.6S (Sw,20 = 3.3–3.6S) with a frictional ratio of 1.49–1.52 corresponding to a mass of between 52–55 kDa, close to that expected for a dimeric species. e, Concentration-normalised scattering plots and DAM fits from SAXS data for T21CC235 (cyan, χ = 1.00), MBP-T21CC235 (green, χ = 1.31) and T21RBCC (purple, χ = 1.05). f, The linear Guinier regions from (e). f, Derived P(r) curves. The CC235 P(r) curve is consistent with an elongated rod, while the two peaks observed for MBP-T21CC235 and T21RBCC are consistent with dumbbell-shaped molecules. h, Scattering plot with DAM fit from TRIM21:Fc SAXS data (red line, 1.00), TRIM21:Fc atomic model fit (blue, χ = 1.04), and apo-TRIM21 atomic model fit (brown dashed,, χ = 2.00). i, The linear Guinier regions from (h). j, Derived P(r) curves.
Extended Data Fig. 4 Anti-GFP antibodies bind to GFP inside living cells.
RPE-1 TRIM21 KO cells expressing membrane-localised GFP (mem-mEGFP) were electroporated with PBS, control IgG or the indicated anti-GFP antibodies. Cells were fixed 3 hours post-electroporation and stained with alexa 647-conjugated anti-IgG secondary antibodies and imaged by confocal microscopy. Scale bar 10 µm. Representative examples from n = 3 independent experiments.
Extended Data Fig. 5 Proteasomal degradation of GFP-tagged proteins by TRIM21 RING-nanobody fusion.
a-d, NIH3T3-Caveolin-1-GFP (a,b) and RPE-1-H2B-mEGFP-FKBP (c,d) cells were electroporated with mRNA encoding mCherry-T21R-vhhGFP4, incubated with either DMSO (control) or MG132 and imaged (a,c) and GFP fluorescence quantified (b,d) with the IncuCyte system. Time, hours (h) post-electroporation. Scale bars, 20 µm. Error bars show s.e.m. from n = 4 technical replicates. Source data for graphs are in Supplementary Data 1.
Supplementary information
Supplementary Information
Supplementary Notes 1 and 2.
Supplementary Data 1
Data for graphs and statistics.
Supplementary Data 2
Uncropped blots, gels and membranes.
Supplementary Video 1
Time-lapse video of Drosophila S2 cells expressing the indicated constructs showing that light-induced clustering of mRFP–CRY2clust–TRIM21 triggers protein degradation, as fluorescence loss is inhibited by treatment with a proteasomal inhibitor (MG132). Clustering was triggered by blue light at time 0. Scale bar, 5 μm.
Rights and permissions
About this article
Cite this article
Zeng, J., Santos, A.F., Mukadam, A.S. et al. Target-induced clustering activates Trim-Away of pathogens and proteins. Nat Struct Mol Biol 28, 278–289 (2021). https://doi.org/10.1038/s41594-021-00560-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-021-00560-2
This article is cited by
-
TRIM21 reduces H1N1-induced inflammation and apoptosis by regulating the TBK1–IRF3 signaling pathway in A549 cells
Archives of Virology (2024)
-
Proteomic studies of VEGFR2 in human placentas reveal protein associations with preeclampsia, diabetes, gravidity, and labor
Cell Communication and Signaling (2024)
-
Protein degradation: expanding the toolbox to restrain cancer drug resistance
Journal of Hematology & Oncology (2023)
-
A TRIM21-based bioPROTAC highlights the therapeutic benefit of HuR degradation
Nature Communications (2023)
-
Trim-Away ubiquitinates and degrades lysine-less and N-terminally acetylated substrates
Nature Communications (2023)