Abstract
Gene regulation in the germline ensures the production of high-quality gametes, long-term maintenance of the species and speciation. Male germline transcriptomes undergo dynamic changes after the mitosis-to-meiosis transition and have been subject to evolutionary divergence among mammals. However, the mechanisms underlying germline regulatory divergence remain undetermined. Here, we show that endogenous retroviruses (ERVs) influence species-specific germline transcriptomes. After the mitosis-to-meiosis transition in male mice, specific ERVs function as active enhancers to drive germline genes, including a mouse-specific gene set, and bear binding motifs for critical regulators of spermatogenesis, such as A-MYB. This raises the possibility that a genome-wide transposition of ERVs rewired germline gene expression in a species-specific manner. Of note, independently evolved ERVs are associated with the expression of human-specific germline genes, demonstrating the prevalence of ERV-driven mechanisms in mammals. Together, we propose that ERVs fine-tune species-specific transcriptomes in the mammalian germline.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The H3K27ac ChIP-seq data reported in this study are described in the accompanying study by Maezawa et al.30 and are deposited to the Gene Expression Omnibus (GEO) under accession code GSE130652. H3K27ac native ChIP-seq data in WT and A-myb mutant PSs and RNA-seq data in CRISPRa ESCs are deposited under accession code GSE142173. All other next-generation sequencing datasets used in this study are publicly available and referenced in Supplementary Data 77,9,24,28,37,43,75,76,77,78,79,80. Source data are provided with this paper.
Code availability
Source code for all software and tools used in this study, with documentation, examples and additional information, is available at following URLs: https://github.com/GenomeImmunobiology/Sakashita_et_al_2020 (best match TE annotation set), https://github.com/alexdobin/STAR (STAR RNA-seq aligner), http://crispor.tefor.net (CRISPOR), http://daehwankimlab.github.io/hisat2 (HISAT2), https://ccb.jhu.edu/software/stringtie (StringTie), https://htseq.readthedocs.io/en/master (HTSeq), https://bedtools.readthedocs.io/en/latest/content/installation.html (BEDTools), https://bioconductor.org/packages/release/bioc/html/DESeq2.html (DESeq2), https://david.ncifcrf.gov/summary.jsp (DAVID), https://software.broadinstitute.org/morpheus (Morpheus), https://software.broadinstitute.org/software/igv/igvtools (IGVTools), http://bowtie-bio.sourceforge.net/bowtie2 (bowtie2), https://www.bioinformatics.babraham.ac.uk/projects/seqmonk (SeqMonk), https://github.com/taoliu/MACS (MACS), https://github.com/shenlab-sinai/ngsplot (ngsplot), https://cran.r-project.org/web/packages/gplots/index.html (gplots), https://github.com/tidyverse/ggplot2 (ggplot2), https://cran.r-project.org/web/packages/chromoMap/vignettes/chromoMap.html#install-chromomap (chromoMap), http://homer.ucsd.edu/homer (HOMER), ·http://great.stanford.edu/public/html (GREAT), https://useast.ensembl.org/info/data/biomart/index.html (BioMart), https://imagej.net/Fiji/Downloads (Fiji, ImageJ), https://systems.crump.ucla.edu/hypergeometric/ (Hypergeometric P value calculator).
References
Ramskold, D., Wang, E. T., Burge, C. B. & Sandberg, R. An abundance of ubiquitously expressed genes revealed by tissue transcriptome sequence data. PLoS Comput. Biol. 5, e1000598 (2009).
Brawand, D. et al. The evolution of gene expression levels in mammalian organs. Nature 478, 343–348 (2011).
Soumillon, M. et al. Cellular source and mechanisms of high transcriptome complexity in the mammalian testis. Cell Rep. 3, 2179–2190 (2013).
Lambert, S. A. et al. The human transcription factors. Cell 172, 650–665 (2018).
Shima, J. E., McLean, D. J., McCarrey, J. R. & Griswold, M. D. The murine testicular transcriptome: characterizing gene expression in the testis during the progression of spermatogenesis. Biol. Reprod. 71, 319–330 (2004).
Namekawa, S. H. et al. Postmeiotic sex chromatin in the male germline of mice. Curr. Biol. 16, 660–667 (2006).
Hasegawa, K. et al. SCML2 establishes the male germline epigenome through regulation of histone H2A ubiquitination. Dev. Cell 32, 574–588 (2015).
Sin, H. S., Kartashov, A. V., Hasegawa, K., Barski, A. & Namekawa, S. H. Poised chromatin and bivalent domains facilitate the mitosis-to-meiosis transition in the male germline. BMC Biol. 13, 53 (2015).
Lesch, B. J., Silber, S. J., McCarrey, J. R. & Page, D. C. Parallel evolution of male germline epigenetic poising and somatic development in animals. Nat. Genet. 48, 888–894 (2016).
Waterston, R. H. et al. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
McClintock, B. The origin and behavior of mutable loci in maize. Proc. Natl Acad. Sci. USA 36, 344–355 (1950).
Meyer, T. J., Rosenkrantz, J. L., Carbone, L. & Chavez, S. L. Endogenous retroviruses: with us and against us. Front. Chem. 5, 23 (2017).
Rebollo, R., Romanish, M. T. & Mager, D. L. Transposable elements: an abundant and natural source of regulatory sequences for host genes. Annu. Rev. Genet. 46, 21–42 (2012).
Friedli, M. & Trono, D. The developmental control of transposable elements and the evolution of higher species. Annu. Rev. Cell Dev. Biol. 31, 429–451 (2015).
Chuong, E. B., Elde, N. C. & Feschotte, C. Regulatory activities of transposable elements: from conflicts to benefits. Nat. Rev. Genet. 18, 71–86 (2017).
Garcia-Perez, J. L., Widmann, T. J. & Adams, I. R. The impact of transposable elements on mammalian development. Development 143, 4101–4114 (2016).
Thompson, P. J., Macfarlan, T. S. & Lorincz, M. C. Long terminal repeats: from parasitic elements to building blocks of the transcriptional regulatory repertoire. Mol. Cell 62, 766–776 (2016).
Zamudio, N. & Bourc’his, D. Transposable elements in the mammalian germline: a comfortable niche or a deadly trap? Heredity (Edinb.) 105, 92–104 (2010).
Crichton, J. H., Dunican, D. S., MacLennan, M., Meehan, R. R. & Adams, I. R. Defending the genome from the enemy within: mechanisms of retrotransposon suppression in the mouse germline. Cell. Mol. Life Sci. 71, 1581–1605 (2014).
Ku, H. Y. & Lin, H. PIWI proteins and their interactors in piRNA biogenesis, germline development and gene expression. Natl Sci. Rev. 1, 205–218 (2014).
Watanabe, T., Cheng, E. C., Zhong, M. & Lin, H. Retrotransposons and pseudogenes regulate mRNAs and lncRNAs via the piRNA pathway in the germline. Genome Res. 25, 368–380 (2015).
Davis, M. P. et al. Transposon-driven transcription is a conserved feature of vertebrate spermatogenesis and transcript evolution. EMBO Rep. 18, 1231–1247 (2017).
Maezawa, S., Yukawa, M., Alavattam, K. G., Barski, A. & Namekawa, S. H. Dynamic reorganization of open chromatin underlies diverse transcriptomes during spermatogenesis. Nucleic Acids Res. 46, 593–608 (2018).
Alavattam, K. G. et al. Attenuated chromatin compartmentalization in meiosis and its maturation in sperm development. Nat. Struct. Mol. Biol. 26, 175–184 (2019).
Patel, L. et al. Dynamic reorganization of the genome shapes the recombination landscape in meiotic prophase. Nat. Struct. Mol. Biol. 26, 164–174 (2019).
Wang, Y. et al. Reprogramming of meiotic chromatin architecture during spermatogenesis. Mol. Cell 73, 547–561 (2019).
Maezawa, S. et al. Polycomb protein SCML2 facilitates H3K27me3 to establish bivalent domains in the male germline. Proc. Natl Acad. Sci. USA 115, 4957–4962 (2018).
Adams, S. R. et al. RNF8 and SCML2 cooperate to regulate ubiquitination and H3K27 acetylation for escape gene activation on the sex chromosomes. PLoS Genet. 14, e1007233 (2018).
Maezawa, S. et al. Super-enhancer switching drives a burst in gene expression at the mitosis-to-meiosis transition. Nat. Struct. Mol. Biol. https://doi.org/10.1038/s41594-020-0488-3 (2020).
Reichmann, J. et al. Microarray analysis of LTR retrotransposon silencing identifies Hdac1 as a regulator of retrotransposon expression in mouse embryonic stem cells. PLoS Comput. Biol. 8, e1002486 (2012).
Ollinger, R. et al. Deletion of the pluripotency-associated Tex19.1 gene causes activation of endogenous retroviruses and defective spermatogenesis in mice. PLoS Genet. 4, e1000199 (2008).
Russ, B. E. et al. Regulation of H3K4me3 at transcriptional enhancers characterizes acquisition of virus-specific CD8+ T cell-lineage-specific function. Cell Rep. 21, 3624–3636 (2017).
Chuong, E. B., Rumi, M. A., Soares, M. J. & Baker, J. C. Endogenous retroviruses function as species-specific enhancer elements in the placenta. Nat. Genet. 45, 325–329 (2013).
Sin, H. S. et al. RNF8 regulates active epigenetic modifications and escape gene activation from inactive sex chromosomes in post-meiotic spermatids. Genes Dev. 26, 2737–2748 (2012).
Bolcun-Filas, E. et al. A-MYB (MYBL1) transcription factor is a master regulator of male meiosis. Development 138, 3319–3330 (2011).
Li, X. Z. et al. An ancient transcription factor initiates the burst of piRNA production during early meiosis in mouse testes. Mol. Cell 50, 67–81 (2013).
Hubley, R. et al. The Dfam database of repetitive DNA families. Nucleic Acids Res. 44, D81–D89 (2016).
Isbel, L. et al. Trim33 binds and silences a class of young endogenous retroviruses in the mouse testis; a novel component of the arms race between retrotransposons and the host genome. PLoS Genet. 11, e1005693 (2015).
Brind’Amour, J. et al. An ultra-low-input native ChIP-seq protocol for genome-wide profiling of rare cell populations. Nat. Commun. 6, 6033 (2015).
Nakasuji, T. et al. Complementary critical functions of Zfy1 and Zfy2 in mouse spermatogenesis and reproduction. PLoS Genet. 13, e1006578 (2017).
McCarrey, J. R. Toward a more precise and informative nomenclature describing fetal and neonatal male germ cells in rodents. Biol. Reprod. 89, 47 (2013).
The ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Ecco, G. et al. Transposable elements and their KRAB-ZFP controllers regulate gene expression in adult tissues. Dev. Cell 36, 611–623 (2016).
Imbeault, M., Helleboid, P. Y. & Trono, D. KRAB zinc-finger proteins contribute to the evolution of gene regulatory networks. Nature 543, 550–554 (2017).
Peaston, A. E. et al. Retrotransposons regulate host genes in mouse oocytes and preimplantation embryos. Dev. Cell 7, 597–606 (2004).
Veselovska, L. et al. Deep sequencing and de novo assembly of the mouse oocyte transcriptome define the contribution of transcription to the DNA methylation landscape. Genome Biol. 16, 209 (2015).
Franke, V. et al. Long terminal repeats power evolution of genes and gene expression programs in mammalian oocytes and zygotes. Genome Res. 27, 1384–1394 (2017).
Bogutz, A. B. et al. Evolution of imprinting via lineage-specific insertion of retroviral promoters. Nat. Commun. 10, 5674 (2019).
De Iaco, A. et al. DUX-family transcription factors regulate zygotic genome activation in placental mammals. Nat. Genet. 49, 941–945 (2017).
Hendrickson, P. G. et al. Conserved roles of mouse DUX and human DUX4 in activating cleavage-stage genes and MERVL/HERVL retrotransposons. Nat. Genet. 49, 925–934 (2017).
Kim, T. K. et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182–187 (2010).
Wang, M. et al. Single-cell RNA sequencing analysis reveals sequential cell fate transition during human spermatogenesis. Cell Stem Cell 23, 599–614 (2018).
Dunn-Fletcher, C. E. et al. Anthropoid primate-specific retroviral element THE1B controls expression of CRH in placenta and alters gestation length. PLoS Biol. 16, e2006337 (2018).
Chuong, E. B. The placenta goes viral: retroviruses control gene expression in pregnancy. PLoS Biol. 16, e3000028 (2018).
Yuan, C. L. & Hu, Y. C. A transgenic core facility’s experience in genome editing revolution. Adv. Exp. Med. Biol. 1016, 75–90 (2017).
Haeussler, M. et al. Evaluation of off-target and on-target scoring algorithms and integration into the guide RNA selection tool CRISPOR. Genome Biol. 17, 148 (2016).
Li, E., Bestor, T. H. & Jaenisch, R. Targeted mutation of the DNA methyltransferase gene results in embryonic lethality. Cell 69, 915–926 (1992).
Chavez, A. et al. Highly efficient Cas9-mediated transcriptional programming. Nat. Methods 12, 326–328 (2015).
Wang, W. et al. Chromosomal transposition of PiggyBac in mouse embryonic stem cells. Proc. Natl Acad. Sci. USA 105, 9290–9295 (2008).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Feng, J., Liu, T. & Zhang, Y. Using MACS to identify peaks from ChIP-Seq data. Curr. Protoc. Bioinformatics 34, 2.14.1–2.14.14 (2011).
Shen, L., Shao, N., Liu, X. & Nestler, E. ngs.plot: quick mining and visualization of next-generation sequencing data by integrating genomic databases. BMC Genomics 15, 284 (2014).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Kinsella, R. J. et al. Ensembl BioMarts: a hub for data retrieval across taxonomic space. Database 2011, bar030 (2011).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Jung, Y. H. et al. Chromatin states in mouse sperm correlate with embryonic and adult regulatory landscapes. Cell Rep. 18, 1366–1382 (2017).
Lavin, Y. et al. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell 159, 1312–1326 (2014).
Guo, J. et al. Chromatin and single-cell RNA-Seq profiling reveal dynamic signaling and metabolic transitions during human spermatogonial stem cell development. Cell Stem Cell 21, 533–546 (2017).
Li, D. et al. Chromatin accessibility dynamics during iPSC reprogramming. Cell Stem Cell 21, 819–833 (2017).
He, S. et al. Hemi-methylated CpG sites connect Dnmt1-knockdown-induced and Tet1-induced DNA demethylation during somatic cell reprogramming. Cell Discov. 5, 11 (2019).
Cao, S. et al. Chromatin accessibility dynamics during chemical induction of pluripotency. Cell Stem Cell 22, 529–542 (2018).
Acknowledgements
We thank M. Weirauch and members of the Namekawa laboratory for discussion and helpful comments regarding the manuscript, the CCHMC Research Flow Cytometry Core for sharing FACS equipment (supported by NIH S10OD023410), X. Li at the University of Rochester Medical Center for sharing A-myb mutant mice, the laboratory of B. Bernstein at Massachusetts General Hospital for providing human testis H3K27ac ChIP-seq data (ENCSR136ZQZ, ENCODE) and the Transgenic Animal and Genome Editing Core at CCHMC for generating the Zfy2 enhancer-deletion mice. We acknowledge the following funding sources: Lalor Foundation Postdoctoral Fellowship and JSPS Overseas Research Fellowship to A.S.; the Research Project Grant by the Azabu University Research Services Division, Ministry of Education, Culture, Sports, Science and Technology (MEXT) Supported Program for the Private University Research Branding Project (2016–2019), a Grant-in-Aid for Research Activity Start-up (19K21196) and the Uehara Memorial Foundation Research Incentive Grant (2018) to S.M.; an Albert J. Ryan Fellowship to K.G.A.; National Institute of Health (NIH) grant DP2 GM119134 to A.B.; a March of Dimes Prematurity Research Centre Collaborative Grant (#22-FY14-470) to M.P.; NIH R01 GM122776 grant to S.H.N.
Author information
Authors and Affiliations
Contributions
The manuscript was written by A.S., K.G.A. and S.H.N., with critical feedback from all other authors. A.S. and S.H.N. designed the study. S.M. performed crosslinking ChIP-seq experiments and A.S. performed native ChIP-seq experiments. A.S. analyzed A-myb mutant mice with the help of K.T. A.S. and K.T. performed CRISPRa experiments. A.S. performed immunostaining and dual-luciferase reporter assays. Y.-C.H. supervised the generation of the Zfy2 enhancer-deletion mice. A.S., K.G.A., M.Y., S.K., N.F.P., A.B., M.P. and S.H.N. designed and interpreted the computational analyses. A.S. performed the majority of computational analyses. S.H.N. supervised the project.
Corresponding author
Ethics declarations
Competing interests
A.B. is a cofounder of Datirium, LLC.
Additional information
Peer review information Beth Moorefield was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Analysis of repetitive element expression during mouse spermatogenesis.
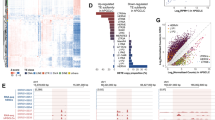
a, The RNA-seq pipeline for comprehensive quantification of TE copies. The flowchart indicates the various RNA-seq and data analysis processes that comprise the pipeline. Round-corner rectangles, input files; rectangles, output files; diamond, branch condition. The specific tools used are highlighted in red. b, The proportion of expressed and unexpressed copies of repetitive elements in each class during spermatogenesis. Of note, nearly half of rRNA genes are expressed in spermatogenic differentiation following the KIT+ spermatogonia stage.
Extended Data Fig. 2 ATAC-seq read enrichment at representative enhancer-like ERV loci and 5,000 randomly selected repetitive element loci.
a, Heatmap depicts RPKM-normalized ATAC-seq reads at enhancer-like RLTR10 and RMER17 loci (n = 694), and 5,000 randomly selected repetitive element loci in representative stages of spermatogenesis. b, Top: Venn diagram shows the intersection between total copy numbers of MMERVK10C loci (green) and total copy numbers of all RLTR10C loci (pink). Bottom: Venn diagram shows the intersection between total copy numbers of MMERVK10C loci (green) and total copy numbers of enhancer-like RLTR10C loci (red).
Extended Data Fig. 3 H3K4me3 enrichment at enhancer-like ERVs loci.
a, Average tag density plots and heatmaps show H3K27ac and H3K4me3 enrichments around enhancer-like ERVs (±1 kb around ±5 kb of ERVs) in PS. b, Scatter plot depicts H3K4me3 enrichments at enhancer-like ERV loci in PS. X-axis indicates relative distance of enhancer-like ERV loci from TSS of nearest genes. Y-axis indicates relative H3K4me3 enrichments at individual enhancer-like ERV loci. Red line shows a regression line.
Extended Data Fig. 4 The genomic features of enhancer-like ERVs in meiosis.
a, Representative track views show H3K27ac ChIP-seq, ATAC-seq, RNA-seq, and A-MYB ChIP-seq signals on chromosome X. The red highlight indicates an enhancer-like ERV locus. b, Pie charts indicate the distributions of enhancer-like ERVs on autosomes and sex chromosome. c, Top: Bar chart depicts the numbers of enhancer-like ERVs on each chromosome. Bottom: Chromosome map shows the distribution of enhancer-like ERVs throughout the mouse genome. Values for H3K27ac enrichment represent log2 fold enrichment of H3K27ac signal relative to input. d, Box-and-whisker plots show relative H3K27ac enrichment at enhancer-like ERV loci on autosomes and sex chromosomes. Values: log2 fold enrichment of H3K27ac signal relative to input. Central bars represent medians, the boxes encompass 50% of the data points, and the error bars indicate 90% of the data points. We detected no statistical difference in H3K27ac enrichment at autosome enhancer-like ERVs vs. sex chromosome enhancer-like ERVs: P = 0.307, Mann-Whitney U test. e, Bar chart shows enhancer-like ERVs distribution across genomic entities (intergenic, intronic, etc.) in autosomes versus the sex chromosomes: P = 3.6 × 10-5, Chi-square test with Yates’s correction. f, The consensus sequence of RLTR10B, listed in the Dfam database, contains two A-MYB binding motifs (GGCAGTT).
Extended Data Fig. 5 The generation of CRISPRa embryonic stem cell lines, and the evaluation of CRISPR-deletion mice.
a, qRT-PCR analyses of CRISPRa embryonic stem (ES) cells show expression level changes of the dCas9-VPR transgene 24 h after doxycycline (Dox) induction. Expression levels were normalized to the endogenous housekeeping gene Hprt. Upon addition of Dox, all ES cell clones evinced overt dCas9-VPR mRNA expression. Because clone #6 exhibited the highest upregulation of dCas9-VPR transcript, we restricted further experiments to clone #6. b, Representative image of CRISPRa ES cell colonies at day 4 after transduction with the sgRNA lentiviral construct. We validated the degree of sgRNA expression through observations of the red fluorescent reporter protein DsRed. Scale bar, 200 µm. c, Testis sections from wild-type (WT; left) and Zfy2 enhancer-deletion mice (right) at postnatal day 28 (P28). The sections were stained with hematoxylin and eosin. Scale bars, 100 μm. In our observations of Zfy2 enhancer-deletion samples, we noted no gross changes to testis morphology; however, we observed multinucleated cells (arrowheads).
Extended Data Fig. 6 The synteny of mouse meiosis-specific enhancer-like ERVs in rats and other placental mammals.
a, Pie charts indicate the genomic distribution of enhancer-like ERVs in the following genomes: mouse (mm10) and rat (rn6). Between the two species, genomic feature enrichment statistically differs: *** P < 0.001, Chi-square test with Yates’s correction. b, Representative track views show evolutionary conservation in regions adjacent to enhancer-like ERVs across several placental mammals. Red highlights indicate enhancer-like ERV loci; such loci exhibit low levels of conservation across placental mammals, including rats, a species closely related to mice.
Extended Data Fig. 7 MER57E3 is enriched in KRAB-ZF-encoding genes that have rapidly evolved in primates or humans.
a, Representative track views show H3K27ac ChIP-seq enrichment for whole, adult human testis tissue and RNA-seq signal in human KIT+ and PS. Red highlights indicate enhancer-like MER57E3s that overlap high levels of H3K27ac deposition. b, Pie charts indicate the genomic distribution of enhancer-like MER57E3 loci in the human genome (hg38). Most enhancer-like MER57E3s are located within the first intronic regions of KRAB-ZF-encoding genes.
Extended Data Fig. 8 A-MYB is highly expressed in both mouse and human spermatocytes.
a, Testis sections from mice at 12 weeks of age immunostained with antibodies raised against A-MYB (red) and γH2AX (green), and counterstained with DAPI (gray). The Roman numerals indicate stages of the seminiferous epithelium cycle. Scale bars, 20 µm. b, Representative testis sections from humans at 29-to-65 years of age immunohistochemically stained with an antibody raised against A-MYB (brown), counterstained with hematoxylin. Images of human testis sections were sourced and adapted from the Human Protein Atlas (www.proteinatlas.org/ENSG00000185697-MYBL1/tissue/testis). Scale bars, 20 µm.
Supplementary information
Supplementary Data 1
List of differentially expressed TE copies between two spermatogenic stages.
Supplementary Data 2
List of locations of meiosis-specific enhancer-like ERVs and their H3K27ac enrichments.
Supplementary Data 3
List of genes adjacent to meiosis-specific enhancer-like ERVs.
Supplementary Data 4
List of upregulated genes in Dox+, A-MYB+ and Dox+;A-MYB+ conditions.
Supplementary Data 5
List of locations of human enhancer-like ERVs in testis.
Supplementary Data 6
List of sequence information used in this study.
Supplementary Data 7
List of next-generation sequencing data from other sources.
Source data
Source Data Fig. 1
Statistical source data
Source Data Fig. 2
Statistical source data
Source Data Fig. 3
Statistical source data
Source Data Fig. 4
Statistical source data
Source Data Fig. 5
Statistical source data
Source Data Fig. 6
Statistical source data
Source Data Fig. 7
Statistical source data
Rights and permissions
About this article
Cite this article
Sakashita, A., Maezawa, S., Takahashi, K. et al. Endogenous retroviruses drive species-specific germline transcriptomes in mammals. Nat Struct Mol Biol 27, 967–977 (2020). https://doi.org/10.1038/s41594-020-0487-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-020-0487-4