Abstract
Precise nucleosome organization at eukaryotic promoters is thought to be generated by multiple chromatin remodeler (CR) enzymes and to affect transcription initiation. Using an integrated analysis of chromatin remodeler binding and nucleosome occupancy following rapid remodeler depletion, we investigated the interplay between these enzymes and their impact on transcription in yeast. We show that many promoters are affected by multiple CRs that operate in concert or in opposition to position the key transcription start site (TSS)-associated +1 nucleosome. We also show that nucleosome movement after CR inactivation usually results from the activity of another CR and that in the absence of any remodeling activity, +1 nucleosomes largely maintain their positions. Finally, we present functional assays suggesting that +1 nucleosome positioning often reflects a trade-off between maximizing RNA polymerase recruitment and minimizing transcription initiation at incorrect sites. Our results provide a detailed picture of fundamental mechanisms linking promoter nucleosome architecture to transcription initiation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Code availability
Peak-calling software is available at https://gitlab.unige.ch/JLFalcone/peakmatic.
References
Lai, W. K. M. & Pugh, B. F. Understanding nucleosome dynamics and their links to gene expression and DNA replication. Nat. Rev. Mol. Cell Biol. 18, 548–562 (2017).
Brahma, S. & Henikoff, S. RSC-associated subnucleosomes define MNase-sensitive promoters in yeast. Mol. Cell 73, 238–249.e3 (2019).
Henikoff, J. G., Belsky, J. A., Krassovsky, K., MacAlpine, D. M. & Henikoff, S. Epigenome characterization at single base-pair resolution. Proc. Natl Acad. Sci. USA 108, 18318–18323 (2011).
Kent, N. A., Adams, S., Moorhouse, A. & Paszkiewicz, K. Chromatin particle spectrum analysis: a method for comparative chromatin structure analysis using paired-end mode next-generation DNA sequencing. Nucleic Acids Res. 39, e26 (2011).
Kubik, S. et al. Nucleosome stability distinguishes two different promoter types at all protein-coding genes in yeast. Mol. Cell 60, 422–434 (2015).
Weiner, A., Hughes, A., Yassour, M., Rando, O. J. & Friedman, N. High-resolution nucleosome mapping reveals transcription-dependent promoter packaging. Genome Res. 20, 90–100 (2010).
Xi, Y., Yao, J., Chen, R., Li, W. & He, X. Nucleosome fragility reveals novel functional states of chromatin and poises genes for activation. Genome Res. 21, 718–724 (2011).
Rhee, H. S. & Pugh, B. F. Genome-wide structure and organization of eukaryotic pre-initiation complexes. Nature 483, 295–301 (2012).
Kubik, S. et al. Sequence-directed action of RSC remodeler and general regulatory factors modulates +1 nucleosome position to facilitate transcription. Mol. Cell 71, 89–102.e5 (2018).
Clapier, C. R., Iwasa, J., Cairns, B. R. & Peterson, C. L. Mechanisms of action and regulation of ATP-dependent chromatin-remodelling complexes. Nat. Rev. Mol. Cell Biol. 18, 407–422 (2017).
Badis, G. et al. A library of yeast transcription factor motifs reveals a widespread function for Rsc3 in targeting nucleosome exclusion at promoters. Mol. Cell 32, 878–887 (2008).
Ganguli, D., Chereji, R. V., Iben, J. R., Cole, H. A. & Clark, D. J. RSC-dependent constructive and destructive interference between opposing arrays of phased nucleosomes in yeast. Genome Res. 24, 1637–1649 (2014).
Krietenstein, N. et al. Genomic nucleosome organization reconstituted with pure proteins. Cell 167, 709–721 e12 (2016).
Yen, K., Vinayachandran, V., Batta, K., Koerber, R. T. & Pugh, B. F. Genome-wide nucleosome specificity and directionality of chromatin remodelers. Cell 149, 1461–1473 (2012).
Boeger, H., Griesenbeck, J., Strattan, J. S. & Kornberg, R. D. Nucleosomes unfold completely at a transcriptionally active promoter. Mol. Cell 11, 1587–1598 (2003).
Floer, M. et al. ARSC/nucleosome complex determines chromatin architecture and facilitates activator binding. Cell 141, 407–418 (2010).
Klein-Brill, A., Joseph-Strauss, D., Appleboim, A. & Friedman, N. Dynamics of chromatin and transcription during transient depletion of the rsc chromatin remodeling complex. Cell Rep. 26, 279–292.e5 (2019).
Gkikopoulos, T. et al. A role for Snf2-related nucleosome-spacing enzymes in genome-wide nucleosome organization. Science 333, 1758–1760 (2011).
Whitehouse, I., Rando, O. J., Delrow, J. & Tsukiyama, T. Chromatin remodelling at promoters suppresses antisense transcription. Nature 450, 1031–1035 (2007).
Zentner, G. E., Tsukiyama, T. & Henikoff, S. ISWI and CHD chromatin remodelers bind promoters but act in gene bodies. PLoS Genet 9, e1003317 (2013).
Kobor, M. S. et al. A protein complex containing the conserved Swi2/Snf2-related ATPase Swr1p deposits histone variant H2A.Z into euchromatin. PLoS Biol. 2, E131 (2004).
Krogan, N. J. et al. A Snf2 family ATPase complex required for recruitment of the histone H2A variant Htz1. Mol. Cell 12, 1565–1576 (2003).
Mizuguchi, G. et al. ATP-driven exchange of histone H2AZ variant catalyzed by SWR1 chromatin remodeling complex. Science 303, 343–348 (2004).
Mohd-Sarip, A. et al. DOC1-dependent recruitment of NURD reveals antagonism with SWI/SNF during epithelial-mesenchymal transition in oral cancer cells. Cell Rep. 20, 61–75 (2017).
Morris, S. A. et al. Overlapping chromatin-remodeling systems collaborate genome wide at dynamic chromatin transitions. Nat. Struct. Mol. Biol. 21, 73–81 (2014).
Parnell, T. J., Schlichter, A., Wilson, B. G. & Cairns, B. R. The chromatin remodelers RSC and ISW1 display functional and chromatin-based promoter antagonism. eLife 4, e06073 (2015).
Tomar, R. S., Psathas, J. N., Zhang, H., Zhang, Z. & Reese, J. C. A novel mechanism of antagonism between ATP-dependent chromatin remodeling complexes regulates RNR3 expression. Mol. Cell Biol. 29, 3255–3265 (2009).
El-Brolosy, M. A. & Stainier, D. Y. R. Genetic compensation: A phenomenon in search of mechanisms. PLoS Genet. 13, e1006780 (2017).
Zentner, G. E., Kasinathan, S., Xin, B., Rohs, R. & Henikoff, S. ChEC-seq kinetics discriminates transcription factor binding sites by DNA sequence and shape in vivo. Nat Commun. 6, 8733 (2015).
Haruki, H., Nishikawa, J. & Laemmli, U. K. The anchor-away technique: rapid, conditional establishment of yeast mutant phenotypes. Mol. Cell 31, 925–932 (2008).
Bruzzone, M. J., Grunberg, S., Kubik, S., Zentner, G. E. & Shore, D. Distinct patterns of histone acetyltransferase and mediator deployment at yeast protein-coding genes. Genes Dev. 32, 1252–1265 (2018).
Churchman, L. S. & Weissman, J. S. Nascent transcript sequencing visualizes transcription at nucleotide resolution. Nature 469, 368–373 (2011).
Morawska, M. & Ulrich, H. D. An expanded tool kit for the auxin-inducible degron system in budding yeast. Yeast 30, 341–351 (2013).
van Bakel, H. et al. A compendium of nucleosome and transcript profiles reveals determinants of chromatin architecture and transcription. PLoS Genet. 9, e1003479 (2013).
Rawal, Y. et al. SWI/SNF and RSC cooperate to reposition and evict promoter nucleosomes at highly expressed genes in yeast. Genes Dev. 32, 695–710 (2018).
Hughes, A. L. & Rando, O. J. Mechanisms underlying nucleosome positioning in vivo. Annu Rev. Biophys. 43, 41–63 (2014).
Kornberg, R. D. & Stryer, L. Statistical distributions of nucleosomes: nonrandom locations by a stochastic mechanism. Nucleic Acids Res. 16, 6677–6690 (1988).
Shivaswamy, S. & Iyer, V. R. Stress-dependent dynamics of global chromatin remodeling in yeast: dual role for SWI/SNF in the heat shock stress response. Mol. Cell Biol. 28, 2221–2234 (2008).
Challal, D. et al. General regulatory factors control the fidelity of transcription by restricting non-coding and ectopic initiation. Mol. Cell 72, 955–969 e7 (2018).
Dreos, R., Ambrosini, G. & Bucher, P. Influence of rotational nucleosome positioning on transcription start site selection in animal promoters. PLoS Comput Biol. 12, e1005144 (2016).
Malabat, C., Feuerbach, F., Ma, L., Saveanu, C. & Jacquier, A. Quality control of transcription start site selection by nonsense-mediated-mRNA decay. eLife 4, e06722 (2015).
Fennessy, R. T. & Owen-Hughes, T. Establishment of a promoter-based chromatin architecture on recently replicated DNA can accommodate variable inter-nucleosome spacing. Nucleic Acids Res. 44, 7189–7203 (2016).
Vasseur, P. et al. Dynamics of nucleosome positioning maturation following genomic replication. Cell Rep. 16, 2651–2665 (2016).
Yadav, T. & Whitehouse, I. Replication-Coupled nucleosome assembly and positioning by ATP-dependent chromatin-remodeling enzymes. Cell Rep. 15, 715–723 (2016).
Ramachandran, S., Ahmad, K. & Henikoff, S. Capitalizing on disaster: Establishing chromatin specificity behind the replication fork. Bioessays https://doi.org/10.1002/bies.201600150 (2017).
Agalioti, T. et al. Ordered recruitment of chromatin modifying and general transcription factors to the IFN-beta promoter. Cell 103, 667–678 (2000).
Nocetti, N. & Whitehouse, I. Nucleosome repositioning underlies dynamic gene expression. Genes Dev. 30, 660–672 (2016).
Flaus, A. & Owen-Hughes, T. Dynamic properties of nucleosomes during thermal and ATP-driven mobilization. Mol. Cell Biol. 23, 7767–7779 (2003).
Kassabov, S. R., Zhang, B., Persinger, J. & Bartholomew, B. SWI/SNF unwraps, slides, and rewraps the nucleosome. Mol. Cell 11, 391–403 (2003).
Chaban, Y. et al. Structure of a RSC-nucleosome complex and insights into chromatin remodeling. Nat. Struct. Mol. Biol. 15, 1272–1277 (2008).
Dechassa, M. L. et al. Architecture of the SWI/SNF-nucleosome complex. Mol. Cell Biol. 28, 6010–6021 (2008).
Stockdale, C., Flaus, A., Ferreira, H. & Owen-Hughes, T. Analysis of nucleosome repositioning by yeast ISWI and Chd1 chromatin remodeling complexes. J. Biol. Chem. 281, 16279–16288 (2006).
Udugama, M., Sabri, A. & Bartholomew, B. The INO80 ATP-dependent chromatin remodeling complex is a nucleosome spacing factor. Mol. Cell Biol. 31, 662–673 (2011).
Bowman, G. D. & McKnight, J. N. Sequence-specific targeting of chromatin remodelers organizes precisely positioned nucleosomes throughout the genome. Bioessays 39, 1–8 (2017).
Goldmark, J. P., Fazzio, T. G., Estep, P. W., Church, G. M. & Tsukiyama, T. The Isw2 chromatin remodeling complex represses early meiotic genes upon recruitment by Ume6p. Cell 103, 423–433 (2000).
McKnight, J. N., Tsukiyama, T. & Bowman, G. D. Sequence-targeted nucleosome sliding in vivo by a hybrid Chd1 chromatin remodeler. Genome Res. 26, 693–704 (2016).
Wu, A. C. K. et al. Repression of divergent noncoding transcription by a sequence-specific transcription factor. Mol. Cell 72, 942–954.e7 (2018).
Yadon, A. N. et al. Chromatin remodeling around nucleosome-free regions leads to repression of noncoding RNA transcription. Mol. Cell Biol. 30, 5110–5122 (2010).
Hainer, S. J. et al. Suppression of pervasive noncoding transcription in embryonic stem cells by esBAF. Genes Dev. 29, 362–378 (2015).
David, F. P. et al. HTSstation: a web application and open-access libraries for high-throughput sequencing data analysis. PLoS One 9, e85879 (2014).
Khan, A. et al. JASPAR 2018: update of the open-access database of transcription factor binding profiles and its web framework. Nucleic Acids Res. 46, D260–D266 (2018).
Lerdrup, M., Johansen, J. V., Agrawal-Singh, S. & Hansen, K. An interactive environment for agile analysis and visualization of ChIP-sequencing data. Nat. Struct. Mol. Biol. 23, 349–357 (2016).
Acknowledgements
We thank M. Docquier and the iGE3 Genomics Platform (https://ige3.genomics.unige.ch/) at the University of Geneva for high-throughput sequencing services, N. Roggli for expert assistance with data presentation and artwork, B. Albert for help with fluorescence microscopy, and all members of the Shore lab for comments and discussions throughout the course of this work. M.J.B. was supported in part by an iGE3 PhD student fellowship. D.C. was supported by a fellowship from the Ligue Contre le Cancer. D.L. acknowledges support from the Centre National de la Recherche Scientifique (CNRS), the Fondation pour la Recherche Medicale (FRM, programme équipes 2013), l’Agence National pour la Recherche (ANR, grant ANR-16-CE12-0022-01 to D.L.), and the Labex Who Am I? (ANR-11-LABX-0071 et Idex ANR-11-IDEX-0005-02 to D.L.). D.S. acknowledges funding from the Swiss National Science Foundation (grant no. 31003A_170153) and the Republic and Canton of Geneva.
Author information
Authors and Affiliations
Contributions
Conceptualization: S.K., D.C., M.J.B., D.L. and D.S.; formal analysis: S.K., D.C., M.J.B., R.D., P.B., D.L. and D.S.; investigation: S.K., D.C., M.J.B. and S.M.; data curation: S.K., D.C., M.J.B., R.D., D.L. and P.B.; writing of original draft: S.K. and D.S.; funding acquisition: D.S., D.L. and P.B.; resources: D.S., D.L. and P.B.; supervision: D.S., D.L. and P.B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Beth Moorefield was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
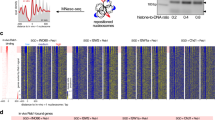
Supplementary Figure 1 Characterization of remodeler binding by ChEC-seq.
a, Growth assays (serial dilution “spot assays”) of the indicated MNase-tagged chromatin remodeler strains on YPAD medium, compared to the wild-type parent strain (WT). Plates were photographed following 24 or 48 hrs incubation at 30oC. b, Distribution of identified remodeler binding sites between different genomic features. c, Heatmaps displaying ChEC signal for every CR centered on the +1 nucleosome of 5,040 protein-coding genes, sorted (top to bottom) by decreasing signal calculated in a region -250 to -50 bp from +1 nucleosome dyad. d, Grid representing Pearson correlation coefficients between ChEC signal for different CRs calculated at all identified remodeler binding sites. e, Plots displaying average normalized ChEC signal for each CR calculated for all clusters. f, Average nucleosome occupancy in each cluster. g, Boxplots displaying expression of genes belonging to each cluster measured either as RNAPII ChIP-seq signal in the gene body or NET-seq signal. h, Fraction of genes in each cluster which contain a canonical TATA-box, dashed line indicates genomic average (~17%).
Supplementary Figure 2 Verification and characterization of remodeler depletion and effects on nucleosome occupancy and stability.
a, Fluorescence microscopy of cells bearing FRB-GFP fusions of Sth1, Snf2, Isw2 and Chd1; cells were treated with rapamycin for indicated times, fixed and stained with DAPI. b, Western blotting (anti-myc antibodies) of cell lysates from an untagged strain and strains bearing Ino80-AID*-myc and Isw1-AID*-myc fusions treated with auxin for indicated amount of time. c, Growth assays (serial dilution “spot assays”, as in Supplementary Fig. 1a) of the indicated anchor-away or AID depletion strains on YPAD medium. “WT” indicates the parental anchor-away and/or AID strains background. d, Pearson correlations for all pairwise comparisons of genome-wide nucleosome occupancy change over 100 bp windows for the indicated CR depletion strains in the absence of depletion (mock-treated). e, Screenshots of sample regions in which nucleosomes become stabilized (marked with a red rectangle) upon depletion of RSC (top) or SWI-SNF (bottom). f, Average plots of nucleosome occupancy for all nucleosomes becoming stabilized upon depletion of RSC (top) or SWI-SNF (bottom). g, Screenshots of sample regions in which nucleosomes become destabilized (marked with a red rectangle) upon depletion of ISW2 (top) or INO80 (bottom). h, Average plots of nucleosome occupancy for all nucleosomes destabilized upon depletion of ISW2 (top) or INO80 (bottom). i,j, Average plots of nucleosome occupancy with (red) or without (blue) depletion of ISW1 (i) or CHD1 (j), plotted separately for genes with the lowest (top) or highest (bottom) binding by the relevant CR (average binding profiles shown in green).
Supplementary Figure 3 Remodeler redundancy in nucleosome positioning.
a, Nucleosome occupancy change upon depletion of RSC (left) or SWI-SNF (right) at sites bound by each remodeler and displaying varying binding signal (+, +/-, -) of the other one. b, Nucleosome occupancy change upon depletion of ISW2 at sites bound by this remodeler and displaying varying binding signal of INO80 (as in (a)). c, Nucleosome occupancy change upon depletion of INO80 at sites bound by this remodeler and displaying varying binding signal of ISW2 (as in (a)). d, Boxplot of INO80 binding signal at INO80-bound sites displaying varying binding signal of ISW2. In (a-d), asterisks indicate significant differences (p<0.05, Mann-Whitney test). e, Snapshot of a sample genomic region displaying nucleosome occupancy change upon depletion of ISW1, CHD1 or both remodelers simultaneously.
Supplementary Figure 4 Multiple concordant and opposing remodeler activities control promoter nucleosome occupancy.
a, Nucleosome occupancy change upon RSC depletion at sites bound by RSC (left) and change upon SWI-SNF depletion at sites bound by SWI-SNF (right). In both cases comparisons are made between sites co-bound by ISW2 (+) or not (-). Asterisk indicates significant difference (p<0.05, Mann-Whitney test). b, Nucleosome occupancy change upon INO80 depletion at sites bound by INO80 and co-bound by RSC and/or SWI-SNF or not, as indicated below. c, Average plots of nucleosome occupancy for all nucleosomes destabilized upon depletion of ISW2, comparing wild-type cells and cells depleted of RSC, ISW2, or both remodelers simultaneously. d, Average plots of nucleosome occupancy for all nucleosomes destabilized upon depletion of INO80, comparing for wild-type cells and cells depleted of RSC, INO80, or both remodelers simultaneously. e, Average plots of nucleosome occupancy for all nucleosomes stabilized upon depletion of RSC, comparing wild-type cells and cells depleted of RSC, RSC and ISW2, RSC and INO80, or all three remodelers simultaneously.
Supplementary Figure 5 Links between nucleosome occupancy and transcriptional regulation.
a, Scatterplot showing relationship between TBP binding change at gene promoters and RNAPII binding change in corresponding gene bodies following SWI-SNF depletion, for all genes with a well-defined TSS; Pearson R value shown. b, Scatterplot showing relationship between TBP binding and nucleosome occupancy changes following SWI-SNF depletion at down-regulated genes. c, Average plots displaying nucleosome occupancy, RNAPII and TBP ChIP signals, in the presence and absence of SWI-SNF, for those genes down-regulated upon SWI-SNF depletion. d-f, Average plots displaying nucleosome occupancy together with RNAPII and TBP ChIP-seq signals for genes up-regulated by INO80 depletion (d), down-regulated by INO80 depletion (e), or up-regulated by ISW2 depletion (f), in each case with or without (+/-) the indicated CR. g, Scatter plot displaying the relationship between TBP binding and nucleosome occupancy changes following double “puller” depletion at genes where transcription was affected (see Fig. 5b, c). h, Scatterplot displaying the TBP - RNAPII ChIP-seq signal relationship at all genes following double “puller” depletion. i-k, Average plots displaying nucleosome occupancy together with RNAPII and TBP ChIP-seq signals for genes down-regulated upon ISW1 depletion (i), up-regulated upon ISW1 depletion (j), or up-regulated upon CHD1 depletion (k), in each case with or without (+/-) the indicated CR.
Supplementary Figure 6 Effects of remodeler depletion on TSS selection.
a, Snapshot of genomic region showing 5’-RACE signal for the Watson (w) and the Crick (c) strands as well as nucleosome occupancy in the presence and absence of RSC; upon RSC depletion the RPO49 gene transcription initiates more upstream comparing to wild-type conditions. b, As in (a) but for SWI-SNF depletion; upon SWI-SNF depletion the SNQ2 gene transcription initiates more frequently downstream comparing to wild-type conditions; additionally, there is more initiation events in the opposite strand just downstream from the upstream-most SNQ2 TSS. c, As in (a) but for ISW2 and INO80 simultaneous depletion; upon “pullers” depletion the FRP6 gene transcription initiates at a downstream position comparing to wild-type conditions. d, Average 5’-RACE signal at genes upregulated upon simultaneous depletion of ISW2 and INO80. e, As in (d) but for downregulated genes.
Supplementary Figure 7 Effect of “puller” remodelers on nucleosome occupancy.
Heatmaps showing nucleosome occupancy, centered at TSS positions determined in cells depleted of ISW2 and INO80, shown in wild type cells and cells depleted of both remodelers.
Supplementary information
Supplementary Information
Supplementary Figures 1–7
Supplementary Table 1
K-means clustering of ChEC-seq data
Supplementary Table 2
GO of clusters
Supplementary Table 3
Strain list
Source data
Rights and permissions
About this article
Cite this article
Kubik, S., Bruzzone, M.J., Challal, D. et al. Opposing chromatin remodelers control transcription initiation frequency and start site selection. Nat Struct Mol Biol 26, 744–754 (2019). https://doi.org/10.1038/s41594-019-0273-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0273-3
This article is cited by
-
Context-specific functions of chromatin remodellers in development and disease
Nature Reviews Genetics (2024)
-
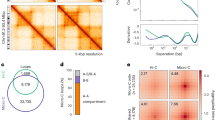
In vitro reconstitution of chromatin domains shows a role for nucleosome positioning in 3D genome organization
Nature Genetics (2024)
-
The BAF chromatin remodeler synergizes with RNA polymerase II and transcription factors to evict nucleosomes
Nature Genetics (2024)
-
Energy-driven genome regulation by ATP-dependent chromatin remodellers
Nature Reviews Molecular Cell Biology (2024)
-
BCL7A and BCL7B potentiate SWI/SNF-complex-mediated chromatin accessibility to regulate gene expression and vegetative phase transition in plants
Nature Communications (2024)