Abstract
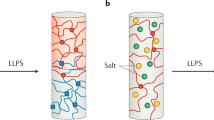
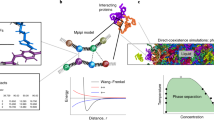
The low-complexity domain of the RNA-binding protein FUS (FUS LC) mediates liquid−liquid phase separation (LLPS), but the interactions between the repetitive SYGQ-rich sequence of FUS LC that stabilize the liquid phase are not known in detail. By combining NMR and Raman spectroscopy, mutagenesis, and molecular simulation, we demonstrate that heterogeneous interactions involving all residue types underlie LLPS of human FUS LC. We find no evidence that FUS LC adopts conformations with traditional secondary structure elements in the condensed phase; rather, it maintains conformational heterogeneity. We show that hydrogen bonding, π/sp2, and hydrophobic interactions all contribute to stabilizing LLPS of FUS LC. In addition to contributions from tyrosine residues, we find that glutamine residues also participate in contacts leading to LLPS of FUS LC. These results support a model in which FUS LC forms dynamic, multivalent interactions via multiple residue types and remains disordered in the densely packed liquid phase.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Chemical shift assignments and relaxation parameters for the condensed phase can be accessed using the BMRB accession 27887. All other data are available from the corresponding author upon reasonable request.
Code availability
Simulation software described in Methods section are publicly available and can be found at http://www.gromacs.org/ for the atomistic resolution simulations and (http://glotzerlab.engin.umich.edu/hoomd-blue/) for coarse-grained simulations HOOMD.
References
Brangwynne, C. P., Mitchison, T. J. & Hyman, A. A. Active liquid-like behavior of nucleoli determines their size and shape in Xenopus laevis oocytes. Proc. Natl Acad. Sci. USA 108, 4334–4339 (2011).
Brangwynne, C. P. et al. Germline P granules are liquid droplets that localize by controlled dissolution/condensation. Science 324, 1729–1732 (2009).
Mitrea, D. M. & Kriwacki, R. W. Phase separation in biology; functional organization of a higher order. Cell Commun. Signal 14, 1 (2016).
King, O. D., Gitler, A. D. & Shorter, J. The tip of the iceberg: RNA-binding proteins with prion-like domains in neurodegenerative disease. Brain Res. 1462, 61–80 (2012).
Burke, K. A., Janke, A. M., Rhine, C. L. & Fawzi, N. L. Residue-by-residue view of in vitro FUS granules that bind the C-terminal domain of RNA polymerase II. Mol. Cell 60, 231–241 (2015).
Patel, A. et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 162, 1066–1077 (2015).
Murakami, T. et al. ALS/FTD mutation-induced phase transition of FUS liquid droplets and reversible hydrogels into irreversible hydrogels impairs RNP granule function. Neuron 88, 678–690 (2015).
Kato, M. et al. Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell 149, 753–767 (2012).
Sun, Z. et al. Molecular determinants and genetic modifiers of aggregation and toxicity for the ALS disease protein FUS/TLS. PLoS Biol. 9, e1000614 (2011).
Couthouis, J. et al. A yeast functional screen predicts new candidate ALS disease genes. Proc. Natl Acad. Sci. USA 108, 20881–20890 (2011).
Fushimi, K. et al. Expression of human FUS/TLS in yeast leads to protein aggregation and cytotoxicity, recapitulating key features of FUS proteinopathy. Protein Cell 2, 141–149 (2011).
Kryndushkin, D., Wickner, R. B. & Shewmaker, F. FUS/TLS forms cytoplasmic aggregates, inhibits cell growth and interacts with TDP-43 in a yeast model of amyotrophic lateral sclerosis. Protein Cell 2, 223–236 (2011).
Bogaert, E. et al. Molecular dissection of FUS points at synergistic effect of low-complexity domains in toxicity. Cell Reports 24, 529–537.e4 (2018).
Hofweber, M. et al. Phase separation of FUS is suppressed by its nuclear import receptor and arginine methylation. Cell 173, 706–719.e13 (2018).
Riggi, N., Cironi, L., Suvà, M.-L. & Stamenkovic, I. Sarcomas: genetics, signalling, and cellular origins. Part 1: The fellowship of TET. J. Pathol. 213, 4–20 (2007).
Kwon, I. et al. Phosphorylation-regulated binding of RNA polymerase II to fibrous polymers of low-complexity domains. Cell 155, 1049–1060 (2013).
Janke, A. M. et al. Lysines in the RNA polymerase II C-terminal domain contribute to TAF15 fibril recruitment. Biochemistry 57, 2549–2563 (2018).
Murray, D. T. et al. Structure of FUS protein fibrils and its relevance to self-assembly and phase separation of low-complexity domains. Cell 171, 615–627.e16 (2017).
Luo, F. et al. Atomic structures of FUS LC domain segments reveal bases for reversible amyloid fibril formation. Nat. Struct. Mol. Biol. 25, 341–346 (2018).
Hughes, M. P. et al. Atomic structures of low-complexity protein segments reveal kinked β sheets that assemble networks. Science 359, 698–701 (2018).
Tsang, B. et al. Phosphoregulated FMRP phase separation models activity-dependent translation through bidirectional control of mRNA granule formation. Proc. Natl Acad. Sci. USA 116, 4218–4227 (2019).
Brady, J. P. et al. Structural and hydrodynamic properties of an intrinsically disordered region of a germ cell-specific protein on phase separation. Proc. Natl Acad. Sci. USA 114, E8194–E8203 (2017).
Reichheld, S. E., Muiznieks, L. D., Keeley, F. W. & Sharpe, S. Direct observation of structure and dynamics during phase separation of an elastomeric protein. Proc. Natl Acad. Sci. 114, E4408–E4415 (2017).
Tompa, P. Intrinsically unstructured proteins. Trends Biochem. Sci. 27, 527–533 (2002).
Nott, T. J. et al. Phase transition of a disordered nuage protein generates environmentally responsive membraneless organelles. Mol. Cell 57, 936–947 (2015).
Vernon, R. M. et al. Pi-Pi contacts are an overlooked protein feature relevant to phase separation. eLife 7, e31486 (2018).
Wilkins, D. K. et al. Hydrodynamic radii of native and denatured proteins measured by pulse field gradient NMR techniques. Biochemistry 38, 16424–16431 (1999).
Ryan, V. H. et al. Mechanistic View of hnRNPA2 Low-Complexity Domain Structure, Interactions, and Phase Separation Altered by Mutation and Arginine Methylation. Mol. Cell 69, 465–479.e7 (2018).
Wei, M. T. et al. Phase behaviour of disordered proteins underlying low density and high permeability of liquid organelles. Nat. Chem. 9, 1118–1125 (2017).
Ying, Q. & Chu, B. Overlap concentration of macromolecules in solution. Macromolecules 20, 362–366 (1987).
Fawzi, N. L., Ying, J., Torchia, D. A. & Clore, G. M. Probing exchange kinetics and atomic resolution dynamics in high-molecular-weight complexes using dark-state exchange saturation transfer NMR spectroscopy. Nat. Protoc. 7, 1523–1533 (2012).
Fawzi, N. L., Ying, J., Ghirlando, R., Torchia, D. A. & Clore, G. M. Atomic-resolution dynamics on the surface of amyloid-β protofibrils probed by solution NMR. Nature 480, 268–272 (2011).
Parekh, S. H., Lee, Y. J., Aamer, K. A. & Cicerone, M. T. Label-free cellular imaging by Broadband coherent anti-stokes raman scattering microscopy. Biophys. J. 99, 2695–2704 (2010).
Monahan, Z. et al. Phosphorylation of the FUS low-complexity domain disrupts phase separation, aggregation, and toxicity. EMBO J. 36, 2951–2967 (2017).
Hernández, B., Coïc, Y. M., Pflüger, F., Kruglik, S. G. & Ghomi, M. All characteristic Raman markers of tyrosine and tyrosinate originate from phenol ring fundamental vibrations. J. Raman Spectrosc. 47, 210–220 (2016).
Siamwiza, M. N. et al. Interpretation of the doublet at 850 and 830 cm−1 in the raman spectra of tyrosyl residues in proteins and certain model compounds. Biochemistry 14, 4870–4876 (1975).
Riback, J. A. et al. Stress-triggered phase separation is an adaptive, evolutionarily tuned response. Cell 168, 1028–1040.e19 (2017).
Ribbeck, K. & Görlich, D. The permeability barrier of nuclear pore complexes appears to operate via hydrophobic exclusion. EMBO J. 21, 2664–2671 (2002).
Kroschwald, S., Maharana, S. & Simon, A. Hexanediol: a chemical probe to investigate the material properties of membrane-less compartments. Matters 3, e201702000010 (2017).
Conicella, A. E., Zerze, G. H., Mittal, J. & Fawzi, N. L. ALS mutations disrupt phase separation mediated by α-helical structure in the TDP-43 low-complexity c-terminal domain. Structure 24, 1537–1549 (2016).
Lin, Y., Currie, S. L. & Rosen, M. K. Intrinsically disordered sequences enable modulation of protein phase separation through distributed tyrosine motifs. J. Biol. Chem. 292, 19110–19120 (2017).
Wang, J. et al. A molecular grammar governing the driving forces for phase separation of prion-like RNA binding proteins. Cell 174, 688–699.e16 (2018).
Hofmeister, F. Zur Lehre von der Wirkung der Salze. Archiv. f. Exp. Pathol. u. Pharmakol. 24, 247–260 (1888).
Cho, Y. et al. Effects of Hofmeister Anions on the Phase Transition Temperature of Elastin-like Polypeptides. J. Phys. Chem. B 112, 13765–13771 (2008).
Wegmann, S. et al. Tau protein liquid–liquid phase separation can initiate tau aggregation. EMBO J. 37, e98049 (2018).
Chaplin, M. Hofmeister series. Water Structure and Science (2001). http://www1.lsbu.ac.uk/water/hofmeister_series.html.
Tüű-Szabó, B., Hoffka, G., Duro, N. & Fuxreiter, M. Altered dynamics may drift pathological fibrillization in membraneless organelles. Biochim. Biophys. Acta Proteins Proteom. (in the press).
Lin, Y., Protter, D. S. W., Rosen, M. K. & Parker, R. Formation and maturation of phase-separated liquid droplets by RNA-binding proteins. Mol. Cell 60, 208–219 (2015).
Franzmann, T. M. et al. Phase separation of a yeast prion protein promotes cellular fitness. Science. 359, eaao5654 (2018).
Peskett, T. R. et al. A liquid to solid phase transition underlying pathological huntingtin exon1 aggregation. Mol. Cell 70, 588–601.e6 (2018).
Langdon, E. M. et al. mRNA structure determines specificity of a polyQ-driven phase separation. Science 360, 922–927 (2018).
Rhys, N. H., Soper, A. K. & Dougan, L. The hydrogen-bonding ability of the amino acid glutamine revealed by neutron diffraction experiments. J. Phys. Chem. B 116, 13308–13319 (2012).
Delaglio, F. et al. NMRPipe: A multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Lee, W., Tonelli, M. & Markley, J. L. NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31, 1325–1327 (2015).
Wang, C., Grey, M. J. & Palmer, A. G. CPMG sequences with enhanced sensitivity to chemical exchange. J. Biomol. NMR 21, 361–366 (2001).
Fawzi, N. L., Ying, J., Torchia, D. A. & Clore, G. M. Kinetics of amyloid β monomer-to-oligomer exchange by NMR relaxation. J. Am. Chem. Soc. 132, 9948–9951 (2010).
Liu, Y., Lee, Y. J. & Cicerone, M. T. Broadband CARS spectral phase retrieval using a time-domain Kramers–Kronig transform. Opt. Lett. 34, 1363–1365 (2009).
Korson, L., Drost-hansen, W. & Millero, F. J. Viscosity of water various temperatures. J. Phys. Chem. 73, 34–39 (1969).
Farrow, N. A., Zhang, O., Szabo, A., Torchia, D. A. & Kay, L. E. Spectral density function mapping using 15N relaxation data exclusively. J. Biomol. NMR 6, 153–162 (1995).
Hess, B., Kutzner, C., Van Der Spoel, D. & Lindahl, E. GRGMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chem. Theory Comput. 4, 435–447 (2008).
Best, R. B., Zheng, W. & Mittal, J. Balanced protein-water interactions improve properties of disordered proteins and non-specific protein association. J. Chem. Theory Comput. 10, 5113–5124 (2014).
Sugita, Y. & Okamoto, Y. Replica-exchange molecular dynamics method for protein folding. Chem. Phys. Lett. 314, 141–151 (1999).
Bonomi, M. & Parrinello, M. Enhanced sampling in the well-tempered ensemble. Phys. Rev. Lett. 104, 190601 (2010).
Barducci, A., Bussi, G. & Parrinello, M. Well-tempered metadynamics: a smoothly converging and tunable free-energy method. Phys. Rev. Lett. 100, 020603 (2008).
Anderson, J. A., Lorenz, C. D. & Travesset, A. General purpose molecular dynamics simulations fully implemented on graphics processing units. J. Comput. Phys. 227, 5342–5359 (2008).
Dignon, G. L., Zheng, W., Kim, Y. C., Best, R. B. & Mittal, J. Sequence determinants of protein phase behavior from a coarse-grained model. PLoS Comput. Biol. 14, e1005941 (2018).
Dignon, G. L., Zheng, W., Best, R. B., Kim, Y. C. & Mittal, J. Relation between single-molecule properties and phase behavior of intrinsically disordered proteins. Proc. Natl Acad. Sci. USA 115, 9929–9934 (2018).
Acknowledgements
We thank M. Naik for helpful advice and assistance with NMR spectroscopy and W. Shing Tang for assistance with NOE calculations from simulated ensembles. Research was supported in part by NIGMS R01GM118530 (to N.L.F.) and Human Frontier Science Program RGP0045/2018 (to S.H.P. and N.L.F.), and NSF 1845734 (to N.L.F.). A.C.M. was supported in part by NIGMS training grant to the MCB graduate program at Brown University (T32GM007601) and NSF graduate fellowship (1644760, to A.C.M.). G.L.D., G.H.Z., and J.M. are supported by the U.S. Department of Energy (DOE), Office of Science, Basic Energy Sciences (BES), Division of Material Sciences and Engineering under Award DESC0013979 (to J.M.). Y.K. and S.H.P. are supported by the Deutsche Forschungsgemeinschaft (DFG) PA-252611-1 and Marie Curie Foundation CIG322284 (to S.H.P.). Use of the high-performance computing capabilities of the Extreme Science and Engineering Discovery Environment (XSEDE), which is supported by the NSF grant TG-MCB-120014, is gratefully acknowledged in addition to resources of the National Energy Research Scientific Computing Center, a DOE Office of Science User Facility supported by the Office of Science of the U.S. Department of Energy under contract DE-AC02-05CH11231. The content is solely the responsibility of the authors and does not necessarily represent the official views of the funding agencies.
Author information
Authors and Affiliations
Contributions
A.C.M and N.L.F. designed and performed experiments and analyzed data for NMR spectroscopy, phase separation assays, and microscopy. Y.K. and S.H.P. designed, performed, and analyzed data for CARS. G.L.D., G.H.Z., and J.M. designed and performed simulation experiments and analyzed the resulting data. A.C.M. and N.L.F. wrote the manuscript with comments from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Anke Sparmann was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 NMR characterization of conformational exchange processes within the condensed phase.
A) Chemical shift differences between the dispersed and condensed phase of FUS LC are small. B) Additional dark-state exchange saturation transfer profiles of condensed FUS LC as a function of the offsets from the 15N carrier frequency and applied saturation field. The differences between profiles can be explained by the differences in value of the transverse relaxation rate constant, 15N R2, of an individual residue. Data are plotted as mean ± s.d. (approximately the size of the circles) propagated from best-fit parameter confidence interval equal to 1 s.d. in the representative data set of two independent experiments. C) 15N CPMG 1D signal intensity profiles of the amide backbone for the reference experiment (τ=0, no relaxation) and as a function of refocusing pulses field (τ=60 ms, νCPMG as listed) indicate that there is no evidence for exchange on the μs-ms timescale for all backbone positions. D) Reduced spectral density mapping for 15N relaxation parameters measured at 850 MHz (black) and 500 MHz (red) 1H Larmor frequencies. Data represent values of J(ω) at frequencies of 0, 86.2, 740 and 0, 50.7, 435 MHz. Data are plotted as mean ± propagated best-fit parameter confidence interval equal to 1 s.d. in the representative data set of two independent experiments.
Supplementary Figure 2 Intermolecular contacts in the condensed phase.
A) Comparison of the glutamine/asparagine side chain plane of the 3D (1H)15N-HSQC-NOESY-1H13C-HSQC between a 1:1 15N/12C and 14N/13C sample (blue) and a 1:1 15N/12C + 14N/12C sample (red). Decreased intensity of the natural abundance control (red) suggests a low contribution of intramolecular artifact to the NOEs. B) 13C/12C filtered-edited NOESY-HSQC on a 1:1 15N/12C and 15N/13C sample. This experiment shows NOEs to all the different residue types. Effective NOE amplitudes calculated from a two-chain simulation of FUS120-163 for serine/threonine backbone (C), glutamine/tyrosine backbone (D), glutamine/asparagine side chains (E), and glycine backbone (F). The overall trend of interaction is similar to experimental NOE profiles, but slight differences in the amplitudes of the NOEs exist, possibly due to the absence of dynamical information in the simulation. Predicted intermolecular NOEs from the FUS 37-97 fibril structure (PDB ID: 5W3N) for serine/threonine backbone (G), glutamine/tyrosine backbone (H), glutamine/asparagine side chains (I), and glycine backbone (J). The overall pattern of NOEs in the fibril structure differs from the condensed phase. The difference between the observed and predicted random coil Cα and Cβ chemical shift values for condensed FUS LC and fibrillar FUS 37-97. Condensed FUS LC values are between -1 and 1, indicating that disorder is preserved in the condensed phase, while chemical shifts in the fibrillar form are closer to values associated with β-sheet structure.
Supplementary Figure 3 Validation of all-atom simulations of FUS120-163.
A) Comparison of experimental dispersed FUS LC and simulated NMR parameters. The simulated dataset is qualitatively similar to the experimental NMR observables, indicating that the all-atom simulation is capturing the molecular details of the 120-163 region. Differences in the N-terminal region between the simulation and experimental dataset arise from the use of a short 44 residue fragment in the simulation. As a result, the N-terminus of the simulated fragment is free to move, while the same region in the experimental dataset is constrained due to being in the middle of the chain. Simulation data are plotted as mean ± s.e.m of n=5 equal divisions of the total data. B) Comparison of secondary shifts between experimental dispersed FUS LC and simulated datasets. Simulation data are plotted as mean ± s.e.m of n=5 equal divisions of the total data. Experimental data are plotted as mean ± estimated uncertainty in peak position. C) Starting configurations of the two-chain simulation of the 120-163 region. This diverse array of configurations was used to avoid bias in the resulting conformations.
Supplementary Figure 4 Paramagnetic relaxation enhancement within the condensed phase.
A) 1H transverse relaxation rates of condensed FUS LC containing ~1% Ac-MTSL labeled A16C, S86C, and S142C. Labeling at position S86C was repeated (S86C #2) and shows a similar relaxation rate as the original experiment (S86C #1). Data are plotted as mean ± best-fit parameter confidence interval equal to 1 s.d. in one data set, with two data sets plotted for S86C. B) 1H transverse relaxation rates of condensed FUS LC containing ~1% MTSL labeled A16C, S86C, and S142C. C-E) Signal intensity ratios of the condensed FUS LC diamagnetic MTSL and paramagnetic MTSL. The global decrease in signal intensity indicates labeling with paramagnetic MTSL. Data are plotted as peak height ratio mean ± propagated uncertainty equal to r.m.s.d. of baseline noise in the representative data set of two experiments.
Supplementary Figure 5 Hydrogen bonding, π/sp2 and hydrophobic interactions calculated from two-chain simulations of FUS120-163.
A) Comparison of the amino acid frequencies found within the low complexity domain of FUS and average disordered and globular proteins24. B) Per-residue hydrogen bonds between each residue of the sequence and different residue types in the other chain. Error bars represent the standard error of the mean for five equal blocks. Tyrosine and glutamine residues are highlighted with gray and red bars respectively. C) Average number of hydrogen bonds formed by each residue of the FUS120-163 sequence in simulation. For the two-chain simulation (dimer) case, only intermolecular hydrogen bonds are considered, and for the single chain simulation (monomer), neighbors of sequence separation of 5 residues or fewer are not counted. D) Average number of hydrophobic contacts formed by each residue of the FUS120-163 sequence in simulation. For the two-chain (dimer) case, only intermolecular hydrogen bonds are considered, and for single chain (monomer), neighbors of sequence separation of 5 residues or fewer are not counted. E) Average number of intermolecular hydrogen bonds, sp2 or hydrophobic contacts between all residues of chain 1 with each residue of chain 2. F). Average total number of hydrogen bonds, sp2 or hydrophobic contacts between residues of type X in chain 1 with type Y in chain 2. Simulation data are plotted as mean ± s.e.m of n=5 equal divisions of the total data.
Supplementary Figure 6 Hofmeister salts do not change the structure of dispersed FUS LC.
A) 1H-15N HSQC spectral overlay of dispersed FUS LC in the presence of the Hofmeister salts.
Supplementary Figure 7 Substitution of tyrosines with phenylalanine in FUS LC leads to aggregation.
A) DIC microscopy of a FUS LC variant in which all 24 tyrosines are mutated to phenylalanine (FUS LC YF). This variant forms aggregates instead of spherical droplets. Phase separation as monitored by turbidity of MBP-FUS FL WT (B) or YF (C). The increased turbidity and light scattering and long incubation time in the YF variant indicates the presence of stable aggregates. Data are plotted as mean ± s.d. of n=3 technical replicates.
Supplementary information
Supplementary Information
Supplementary Figures 1–7, Supplementary Table 1
Rights and permissions
About this article
Cite this article
Murthy, A.C., Dignon, G.L., Kan, Y. et al. Molecular interactions underlying liquid−liquid phase separation of the FUS low-complexity domain. Nat Struct Mol Biol 26, 637–648 (2019). https://doi.org/10.1038/s41594-019-0250-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0250-x
This article is cited by
-
Expanding the molecular language of protein liquid–liquid phase separation
Nature Chemistry (2024)
-
The molecular basis for cellular function of intrinsically disordered protein regions
Nature Reviews Molecular Cell Biology (2024)
-
RNA structure promotes liquid-to-solid phase transition of short RNAs in neuronal dysfunction
Communications Biology (2024)
-
Sequence-dependent material properties of biomolecular condensates and their relation to dilute phase conformations
Nature Communications (2024)
-
A solid beta-sheet structure is formed at the surface of FUS droplets during aging
Nature Chemical Biology (2024)