Abstract
To discriminate between closely related members of a protein family that differ at a limited number of spatially distant positions is a challenge for drug discovery. We describe a combined computational design and experimental selection approach for generating binders targeting functional sites with large, shape complementary interfaces to read out subtle sequence differences for subtype-specific antagonism. Repeat proteins are computationally docked against a functionally relevant region of the target protein surface that varies in the different subtypes, and the interface sequences are optimized for affinity and specificity first computationally and then experimentally. We used this approach to generate a series of human Frizzled (Fz) subtype-selective antagonists with extensive shape complementary interaction surfaces considerably larger than those of repeat proteins selected from random libraries. In vivo administration revealed that Wnt-dependent pericentral liver gene expression involves multiple Fz subtypes, while maintenance of the intestinal crypt stem cell compartment involves only a limited subset.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Davis, M. I. et al. Comprehensive analysis of kinase inhibitor selectivity. Nat. Biotechnol. 29, 1046 (2011).
Massague, J. TGF-beta signal transduction. Annu. Rev. Biochem. 67, 753–791 (1998).
Luca, V. C. et al. Structural basis for Notch1 engagement of Delta-like 4. Science 347, 847–853 (2015).
Artavanis-Tsakonas, S., Rand, M. D. & Lake, R. J. Notch signaling: cell fate control and signal integration in development. Science 284, 770–776 (1999).
Mendoza, J. L. et al. The IFN-λ-IFN-λR1-IL-10Rβ complex reveals structural features underlying type III IFN functional plasticity. Immunity 46, 379–392 (2017).
Spangler, J. B., Moraga, I., Mendoza, J. L. & Garcia, K. C. Insights into cytokine–receptor interactions from cytokine engineering. Annu. Rev. Immunol. 33, 139–167 (2015).
Kummer, L. et al. Structural and functional analysis of phosphorylation-specific binders of the kinase ERK from designed ankyrin repeat protein libraries. Proc. Natl Acad. Sci. 109, E2248–E2257 (2012).
Schilling, J., Schöppe, J. & Plückthun, A. From DARPins to LoopDARPins: novel LoopDARPin design allows the selection of low picomolar binders in a single round of ribosome display. J. Mol. Biol. 426, 691–721 (2014).
Binz, H. K. et al. High-affinity binders selected from designed ankyrin repeat protein libraries. Nat. Biotechnol. 22, 575 (2004).
Plückthun, A. Designed ankyrin repeat proteins (DARPins): binding proteins for research, diagnostics, and therapy. Annu. Rev. Pharmacol. Toxicol. 55, 489–511 (2015).
Nusse, R. & Clevers, H. Wnt/β-catenin signaling, disease, and emerging therapeutic modalities. Cell 169, 985–999 (2017).
Reya, T. & Clevers, H. Wnt signalling in stem cells and cancer. Nature 434, 843 (2005).
Clevers, H. Wnt/β-catenin signaling in development and disease. Cell 127, 469–480 (2006).
Gurney, A. et al. Wnt pathway inhibition via the targeting of Frizzled receptors results in decreased growth and tumorigenicity of human tumors. Proc. Natl Acad. Sci. 109, 11717–11722 (2012).
Janda, C. Y., Waghray, D., Levin, A. M., Thomas, C. & Garcia, K. C. Structural basis of Wnt recognition by Frizzled. Science 337, 59–64 (2012).
Rohl, C. A., Strauss, C. E., Misura, K. M. & Baker, D. in Methods in Enzymology Vol. 383 (New York: Academic Press, 2004).
Fallas, J. A. et al. Computational design of self-assembling cyclic protein homo-oligomers. Nat. Chem. 9, 353 (2017).
Boder, E. T. & Wittrup, K. D. Yeast surface display for screening combinatorial polypeptide libraries. Nat. Biotechnol. 15, 553 (1997).
Whitehead, T. A. et al. Optimization of affinity, specificity and function of designed influenza inhibitors using deep sequencing. Nat. Biotechnol. 30, 543 (2012).
Janda, C. Y. et al. Surrogate Wnt agonists that phenocopy canonical Wnt and β-catenin signalling. Nature 545, 234–237 (2017).
Yan, K. S. et al. Non-equivalence of Wnt and R-spondin ligands during Lgr5+intestinal stem-cell self-renewal. Nature 545, 238–242 (2017).
Kuhnert, F. et al. Essential requirement for Wnt signaling in proliferation of adult small intestine and colon revealed by adenoviral expression of Dickkopf-1. Proc. Natl. Acad. Sci. 101, 266–271 (2004).
Kabiri, Z. et al. Stroma provides an intestinal stem cell niche in the absence of epithelial Wnts. Development 141, 2206–2215 (2014).
Wei, K. et al. A liver Hif-2α–Irs2 pathway sensitizes hepatic insulin signaling and is modulated by Vegf inhibition. Nat. Med. 19, 1331 (2013).
van Es, J. H. et al. Wnt signalling induces maturation of Paneth cells in intestinal crypts. Nat. Cell Biol. 7, 381 (2005).
Benhamouche, S. et al. Apc tumor suppressor gene is the “zonation-keeper” of mouse liver. Dev. Cell 10, 759–770 (2006).
Liu, C. Q., Bakeri, H., Li, T. S. & Swaroop, A. Regulation of retinal progenitor expansion by Frizzled receptors: implications for microphthalmia and retinal coloboma. Hum. Mol. Genet. 21, 1848–1860 (2012).
Kabsch, W. Integration, scaling, space-group assignment and post-refinement. Acta Crystallogr. D 66, 133–144 (2010).
Otwinowski, Z. & Minor, W. in Methods in Enzymology Vol. 276 (New York: Academic Press, 1997).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix. refine. Acta Crystallogr. D 68, 352–367 (2012).
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Acknowledgements
We thank A. Velasco and D. Waghray for assistance. GM/CA@APS has been funded in whole or in part with Federal funds from the National Cancer Institute (ACB-12002) and the National Institute of General Medical Sciences (AGM-12006). This research used resources of the Advanced Photon Source, a US Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under contract no. DE-AC02-06CH11357. The Eiger 16 M detector was funded by an National Institutes of Health (NIH) Office of Research Infrastructure Programs, High-End Instrumentation Grant (1S10OD012289-01A1). The Berkeley Center for Structural Biology is supported in part by the Howard Hughes Medical Institute. The Advanced Light Source is a Department of Energy Office of Science User Facility under contract no. DE-AC02-05CH11231. Use of the SSRL, Stanford Linear Accelerator Center (SLAC) National Accelerator Laboratory, is supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences under contract no. DE-AC02-76SF00515. The SSRL Structural Molecular Biology Program is supported by the Department of Energy Office of Biological and Environmental Research, and by the NIH, National Institute of General Medical Sciences (including P41GM103393). The work here is supported by the Ludwig Foundation and Mathers Fund (K.C.G.), Howard Hughes Medical Institute (K.C.G. and D.B.) and NIH (grants U01DK085527, U19AI116484, R01NS100904, and U01CA217851 to C.J.K., and grant 1R01DK115728 to C.J.K. and K.C.G.).
Author information
Authors and Affiliations
Contributions
L.T.D., Y.M., K.C.G., and D.B. conceived the project. L.T.D. computationally designed, optimized, and characterized DRPB_Fz8, and obtained site saturation mutagenesis data for specificity tuning. Y.M. constructed the Fz4/7 targeting library, performed yeast surface display selection, purified all proteins, determined complex structures, and performed cell line DRPB inhibition assays. A.H., K.Y., and N.H. performed duodenum organoid experiments and in vivo histology experiments. J.G.V. contributed to tissue collection and pathology assessment. M.V. and J.Y. packaged adenovirus. K.P. contributed to computational DRPB design. C.Y.J. generated the luciferase reporter cell line. K.M.J. contributed to crystallography data collection. K.M. contributed to yeast surface display selections. C.J.K. supervised the organoid experiments and in vivo experiments. K.C.G. and D.B. supervised the project and interpreted the data. L.T.D., Y.M., K.C.G., and D.B. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
K.C.G., D.B., Y.M., and L.T.D. are inventors on patent application 62/698576 submitted by Stanford University that covers the use of Frizzled-specific Wnt antagonists.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
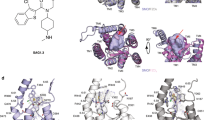
Supplementary Fig. 1 Structural analysis of DRPB_Fz8-Fz8CRD complex.
a, Titration original computation design (WT) and variants to Fz8CRD. The Phe42Ala and Thr66Ser point mutations to the original design (ptm9_5_1_5 in Supplementary Notes Table 1) are sufficient to confer binding to Fz8CRD. b, Titration of DRPB_Fz8 to biotinylated Fz 1/2/4/5/7/8 CRDs. DRPB_Fz8 was displayed on yeast surface. Biotinylated Fz 1/2/4/5/7/8 was added at different concentration. SA647 was subsequently added and the fluorescence data was analyzed and plotted in GraphPad Prism 7. DRPB_Fz8 showed strong binding to Fz5 and Fz8 whilst not interacting with Fz1, 2, 4 and 7 (Supplementary Notes Table 5). The yeast surface display titration has been repeated once with similar results. c, Structural superposition of DRPB_Fz8-Fz8CRD with XWnt8-Fz8CRD (PDB:4F0A). DRPB_Fz8 (light orange) and XWnt8 (magenta) are shown as cartoons. Fz8CRD (blue) is shown in surface representation. The lipid group of XWnt8 is shown as sticks. Zoomed view of the Fz8CRD hydrophobic groove showing steric clashes between DRPB_Fz8 “fingers” with XWnt8 lipid group. d, Phe42Ala mutations prevents steric clashes to neighboring residues.
Supplementary Fig. 2 Selection schematic of Fz-subtype specific DRPB.
Schematic of the selection protocol to obtain Fz-subtype specific DRPB molecules. The detailed protocol of each round of selection is described in the Supplementary Notes.
Supplementary Fig. 3 Binding affinity of individual DRPB to Fz 4/7/8 CRDs using surface plasmon resonance.
a-d, SRP sensogram data of individual DRPB binding to Fz 4/7/8 CRDs. Panels A to D represent DRPB_Fz8, DRPB_Fz7/8, DRPB_Fz7 and DRPB_Fz4. Biotinylated Fz4/7/8 CRD was captured on SA chip and DRPB was flown over the chip surface. The y-axis corresponds to RU change and x-axis corresponds to time (s). The data are analyzed by accompanying Biacore evaluation software and fitted to either a kinetic or steady-state binding model. Panel A to D represents DRPB_Fz8, DRPB_Fz7/8, DRPB_Fz7 and DRPB_Fz4. The sensor captured protein from left to right represents Fz4, 7 and 8. The SPR measurements have been repeated once with similar results. e, Measured Kd by SPR. In DRPB_Fz8 measurement, * represents the Kd measurement is approaching the detection limit of SPR, thus 0.2 nM Kd for BLI was used here for accuracy.
Supplementary Fig. 4 Structural superposition of DRPB in complex with Fz CRDs.
a, Superposition of the overall structures of the two complexes. The RMSD between the two complexes is 0.6 Å. Fz7 is colored in pink. Fz8 is colored in blue. DRPB_Fz7/8 is colored in cyan. DRPB_Fz8 is colored in orange. b, Zoomed view of Met9Arg mutation from DRPB_Fz8 to DRPB_Fz7/8. The Met9Arg mutation leads to hydrogen bond and salt bridge formation between Arg9 with Fz7 Asn75 and Asp78, as shown by grey dashed lines. c, Superposition of the overall structures of the DRPB_Fz7/8-Fz7 and DRPB_Fz7-Fz7. Fz7 is colored in light orange. DRPB_Fz7/8 is colored in light cyan. DRPB_Fz7 is colored in light blue. In the zoomed view, the carboxylate group of Asp112 forms a hydrogen bond and salt bridge with Lys46 of Fz7 CRD, enhancing the affinity.
Supplementary Fig. 5 Structural comparison of DRPB_Fz4-Fz4 and DRPB_Fz8-Fz8.
a, Superposition of the overall structures of the DRPB_Fz4-Fz4CRD and DRPB_Fz8-Fz8CRD showing backbone movement of DRPB_Fz4 (light green), but the original designed “fingers in groove” binding mode is retained. The two complexes were aligned by aligning the Fz4CRD (salmon) with Fz8CRD (light blue), with RMSD of 1 Å. DRPB_Fz8 is colored in light orange. There is no electron density for the C-terminal part of DRPB_Fz4 (residues 160–190), therefore this part of DRPB_Fz4 is not built and shown in the figure. b, Superposition of the overall structures of the DRPB_Fz4-Fz4CRD with XWnt8-Fz8CRD (PDB:4F0A). The two complexes were aligned by aligning the Fz4CRD with Fz8CRD. XWnt8 is colored in magenta. The lipid group of XWnt8 is sterically clashing with DRPB_Fz4 “fingers”. c, Representative zoomed view of Met9Arg mutation. This Met to Arg mutation in DRPB_Fz4 allows hydrogen bond and salt bridge formation with Asp74 of Fz4. d, Zoomed view of Ala101Phe mutation in DRPB_Fz4. Phe101 sidechain is deeply buried in the hydrophobic groove, conferring affinity. On the other hand, in DRPB_Fz8 the Ala101Phe mutation prevents the overall geometry adopted by DRPB_Fz8, eliminating Fz8 subtype binding. e, The loop regions of Fz4CRD interact with each other and adopt a usual dimer geometry within the crystal packing. The dimeric FzCRDs are colored in salmon and light yellow, respectively. Such dimer is not observed for the other DRPB-CRD complexes.
Supplementary Fig. 6 Fz-subtype specific DRPB antagonists inhibit β-catenin in Fz-specific manner.
a, In BeWo cell line, predominantly expressing Fz5, DRPB_Fz8and DRPB_Fz7/8 inhibits β-catenin signaling with IC50 of 0.1 nM and 3.7 nM. The y-axis is normalized Luciferase reporter signaling. In all panels of this figure, cells were stimulated with 20% Wnt3a conditioned media (ATCC) in combination with 25 nM R-spondin 2 and treated with gradient concentration of DRPB proteins. b, In A375BAR cell line predominantly expressing Fz2, DRPB_Fz7/8and DRPB_Fz7inhibits β-catenin signaling with IC50 of 0.3 nM and 1.8 nM. c, In HEK293STF cell line with transfected Fz4 receptor, saturating concentration of DRPB_Fz7/8 was added to inhibit Fz7 and Fz8 subtype Fz receptor-mediated Wnt signaling (shown in Supplementary Fig. 6a and 6b). The β-catenin response therefore is predominantly mediated with Fz4 receptor. Indeed, only DRPB_Fz4 showed inhibition with IC50 of 7.5 nM. Error bar represents S.E. of three replicates. The experiments in this figure were repeated once with similar results.
Supplementary Fig. 7 DRPB inhibition in mouse/human duodenum organoid and in vivo phenotype following DRPB injections.
a, b, Representative bright-field images of mouse (a) and human (b) duodenal organoids expanded for 7 days and 10 days, respectively, using standard submerged Matrigel in medium containing ENR (EGF/Noggin/R-spondin) with different Fz-subtype-specific DRPB antagonist treatments at serial concentrations from 0.1–100 nM. Each Fz antagonist was replenished every 3 days during medium change. In vitro experiments were repeated at least three times, and representative images are shown. Scale bars, 100 μm. c, Representative H & E stained sections of jejunum cross-sections from mice that received PBS or recombinant Fz-subtype-specific DRPB antagonists (conjugated with mouse serum albumin) at a concentration of 15 mg/kg once a day for 5 days. This experiment was performed once due to the protein yield limitation of mouse serum albumin conjugated DRBPs. The in vivo phenotype experiments are repeated using adenoviral delivery of DRBPs, with results shown in Fig. 4. Scale bar, 50 μm.
Supplementary Fig. 8 Time course of serum expression after a single intravenous injection of adenoviruses into C57BL/6 mice.
a, Time course (Day 2 and 7) of serum expression following single i.v. injection of adenoviruses into C57BL/6J mice. Anti-6x-His Western blot was performed on serum at the indicated times post-injection with adenovirus Fz-subtype-specific DRPB antagonist (His-tagged). b, Detection of control IgG2a Fc (Fc) or different Fz-subtype-specific DRPB antagonists in serum of mice 2 days after adenovirus injection using Western blot. 3 mice per condition. Time course serum expression experiment used 3 mice per group and was performed once.
Supplementary Information
Supplementary Information
Supplementary Figures 1–8 and Supplementary Notes
Rights and permissions
About this article
Cite this article
Dang, L.T., Miao, Y., Ha, A. et al. Receptor subtype discrimination using extensive shape complementary designed interfaces. Nat Struct Mol Biol 26, 407–414 (2019). https://doi.org/10.1038/s41594-019-0224-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0224-z
This article is cited by
-
Frizzled receptors (FZDs) in Wnt signaling: potential therapeutic targets for human cancers
Acta Pharmacologica Sinica (2024)
-
Exploring binding positions and backbone conformations of peptide ligands of proteins with a backbone-centred statistical energy function
Journal of Computer-Aided Molecular Design (2023)
-
Therapeutic blood-brain barrier modulation and stroke treatment by a bioengineered FZD4-selective WNT surrogate in mice
Nature Communications (2023)
-
De novo design of protein interactions with learned surface fingerprints
Nature (2023)