Abstract
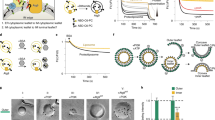
A key event in autophagy is autophagosome formation, whereby the newly synthesized isolation membrane (IM) expands to form a complete autophagosome using endomembrane-derived lipids. Atg2 physically links the edge of the expanding IM with the endoplasmic reticulum (ER), a role that is essential for autophagosome formation. However, the molecular function of Atg2 during ER–IM contact remains unclear, as does the mechanism of lipid delivery to the IM. Here we show that the conserved amino-terminal region of Schizosaccharomyces pombe Atg2 includes a lipid-transfer-protein-like hydrophobic cavity that accommodates phospholipid acyl chains. Atg2 bridges highly curved liposomes, thereby facilitating efficient phospholipid transfer in vitro, a function that is inhibited by mutations that impair autophagosome formation in vivo. These results suggest that Atg2 acts as a lipid-transfer protein that supplies phospholipids for autophagosome formation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Coordinates and structure factors of apo- and PE-bound SpAtg2NR have been deposited in the Protein Data Bank under accession codes 6A9E and 6A9J, respectively. Source data for Figs. 3e,h and 4c are available with the paper online. All other data are available from the corresponding authors upon reasonable request.
References
Mizushima, N., Yoshimori, T. & Ohsumi, Y. The role of Atg proteins in autophagosome formation. Annu. Rev. Cell. Dev. Biol. 27, 107–132 (2011).
Hailey, D. W. et al. Mitochondria supply membranes for autophagosome biogenesis during starvation. Cell 141, 656–667 (2010).
Axe, E. L. et al. Autophagosome formation from membrane compartments enriched in phosphatidylinositol 3-phosphate and dynamically connected to the endoplasmic reticulum. J. Cell Biol. 182, 685–701 (2008).
Hamasaki, M. et al. Autophagosomes form at ER-mitochondria contact sites. Nature 495, 389–393 (2013).
Puri, C. et al. The RAB11A-positive compartment is a primary platform for autophagosome assembly mediated by WIPI2 recognition of PI3P-RAB11A. Dev. Cell 45, 114–131 e118 (2018).
Ge, L. et al. Remodeling of ER-exit sites initiates a membrane supply pathway for autophagosome biogenesis. EMBO Rep. 18, 1586–1603 (2017).
Suzuki, K., Akioka, M., Kondo-Kakuta, C., Yamamoto, H. & Ohsumi, Y. Fine mapping of autophagy-related proteins during autophagosome formation in Saccharomyces cerevisiae. J. Cell Sci. 126, 2534–2544 (2013).
Gomez-Sanchez, R. et al. Atg9 establishes Atg2-dependent contact sites between the endoplasmic reticulum and phagophores. J. Cell Biol. 217, 2743–2763 (2018).
Obara, K., Sekito, T., Niimi, K. & Ohsumi, Y. The Atg18-Atg2 complex is recruited to autophagic membranes via phosphatidylinositol 3-phosphate and exerts an essential function. J. Biol. Chem. 283, 23972–23980 (2008).
Graef, M., Friedman, J. R., Graham, C., Babu, M. & Nunnari, J. ER exit sites are physical and functional core autophagosome biogenesis components. Mol. Biol. Cell 24, 2918–2931 (2013).
Hirata, E., Ohya, Y. & Suzuki, K. Atg4 plays an important role in efficient expansion of autophagic isolation membranes by cleaving lipidated Atg8 in Saccharomyces cerevisiae. PLoS One 12, e0181047 (2017).
Chowdhury, S. et al. Insights into autophagosome biogenesis from structural and biochemical analyses of the ATG2A-WIPI4 complex. Proc. Natl Acad. Sci. USA 115, E9792–E9801 (2018).
Zheng, J. X. et al. Architecture of the ATG2B-WDR45 complex and an aromatic Y/HF motif crucial for complex formation. Autophagy 13, 1870–1883 (2017).
Lang, A. B., John Peter, A. T., Walter, P. & Kornmann, B. ER-mitochondrial junctions can be bypassed by dominant mutations in the endosomal protein Vps13. J. Cell Biol. 210, 883–890 (2015).
Kotani, T., Kirisako, H., Koizumi, M., Ohsumi, Y. & Nakatogawa, H. The Atg2-Atg18 complex tethers pre-autophagosomal membranes to the endoplasmic reticulum for autophagosome formation. Proc. Natl Acad. Sci. USA 115, 10363–10368 (2018).
Wong, L. H., Copic, A. & Levine, T. P. Advances on the transfer of lipids by lipid transfer proteins. Trends Biochem. Sci. 42, 516–530 (2017).
Reinisch, K. M. & De Camilli, P. SMP-domain proteins at membrane contact sites: structure and function. Biochim. Biophys. Acta 1861, 924–927 (2016).
Jeong, H., Park, J. & Lee, C. Crystal structure of Mdm12 reveals the architecture and dynamic organization of the ERMES complex. EMBO Rep. 17, 1857–1871 (2016).
Kawano, S. et al. Structure–function insights into direct lipid transfer between membranes by Mmm1–Mdm12 of ERMES. J. Cell Biol. 217, 959–974 (2018).
Toulmay, A. & Prinz, W. A. A conserved membrane-binding domain targets proteins to organelle contact sites. J. Cell Sci. 125, 49–58 (2012).
Kumar, N. et al. VPS13A and VPS13C are lipid transport proteins differentially localized at ER contact sites. J. Cell Biol. 217, 3625–3639 (2018).
Lemus, L., Ribas, J. L., Sikorska, N. & Goder, V. An ER-localized SNARE protein is exported in specific COPII vesicles for autophagosome biogenesis. Cell Rep. 14, 1710–1722 (2016).
Davis, S., Wang, J. & Ferro-Novick, S. Crosstalk between the secretory and autophagy pathways regulates autophagosome formation. Dev. Cell 41, 23–32 (2017).
Yla-Anttila, P., Vihinen, H., Jokitalo, E. & Eskelinen, E. L. 3D tomography reveals connections between the phagophore and endoplasmic reticulum. Autophagy 5, 1180–1185 (2009).
Hayashi-Nishino, M. et al. A subdomain of the endoplasmic reticulum forms a cradle for autophagosome formation. Nat. Cell Biol. 11, 1433–1437 (2009).
Baba, M., Osumi, M. & Ohsumi, Y. Analysis of the membrane structures involved in autophagy in yeast by freeze-replica method. Cell Struct. Funct. 20, 465–471 (1995).
Toulmay, A. & Prinz, W. A. Lipid transfer and signaling at organelle contact sites: the tip of the iceberg. Curr. Opin. Cell Biol. 23, 458–463 (2011).
Sperandeo, P., Martorana, A. M. & Polissi, A. The lipopolysaccharide transport (Lpt) machinery: a nonconventional transporter for lipopolysaccharide assembly at the outer membrane of Gram-negative bacteria. J. Biol. Chem. 292, 17981–17990 (2017).
Okuda, S., Freinkman, E. & Kahne, D. Cytoplasmic ATP hydrolysis powers transport of lipopolysaccharide across the periplasm in E. coli. Science 338, 1214–1217 (2012).
Sherman, D. J. et al. Lipopolysaccharide is transported to the cell surface by a membrane-to-membrane protein bridge. Science 359, 798–801 (2018).
Matsuyama, A. et al. ORFeome cloning and global analysis of protein localization in the fission yeast Schizosaccharomyces pombe. Nat. Biotechnol. 24, 841–847 (2006).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Delano, W. L. The PyMOL Molecular Graphics System (DeLano Scientific LLC, Palo Alto, CA, 2002).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Kawaoka, T., Ohnuki, S., Ohya, Y. & Suzuki, K. Morphometric analysis of autophagy-related structures in Saccharomyces cerevisiae. Autophagy 13, 2104–2110 (2017).
Noda, T., Matsuura, A., Wada, Y. & Ohsumi, Y. Novel system for monitoring autophagy in the yeast Saccharomyces cerevisiae. Biochem. Biophys. Res. Commun. 210, 126–132 (1995).
Horvath, A. & Riezman, H. Rapid protein extraction from Saccharomyces cerevisiae. Yeast 10, 1305–1310 (1994).
Suzuki, K., Kamada, Y. & Ohsumi, Y. Studies of cargo delivery to the vacuole mediated by autophagosomes in Saccharomyces cerevisiae. Dev. Cell 3, 815–824 (2002).
Acknowledgements
We thank Y. Ishii for her help with protein expression, C. Kondo-Kakuta and Y. Kamada (National Institute for Basic Biology, Japan) for generation of the pCK6 plasmid, S. Kawano and T. Endo (Kyoto Sangyo University) for providing the plasmid for the Mmm1–Mdm12 complex, H. Tachikawa (University of Tokyo) for providing the cDNA of Vps13, and A. I. May for proofreading the manuscript. S. pombe Atg2 cDNA was provided by the RIKEN BioResource Research Center (BRC) through the National BioResource Project of the Ministry of Education, Culture, Sports, Science and Technology (MEXT) and Agency for Medical Research and Development (AMED), Japan. The synchrotron radiation experiments were performed using beamlines BL-1A and BL-17A at Photon Factory (KEK, Japan). This work was supported by the Japan Science and Technology Agency (JST) Core Research for Evolutionary Science and Technology (CREST, Grant Number JPMJCR13M7 to N.N.N. and H.N.), the Japan Society for the Promotion of Science (JSPS) KAKENHI Grant-in-Aid for Scientific Research 25111004 and 18H03989 (to N.N.N.), 25111003 (to H.N.), 16H06375 (to Y.O.), 18J13429 (to E.H.), 16H06280 and 18H04853 (to K.S.), and the Platform Project for Supporting in Drug Discovery and Life Science Research (Platform for Drug Discovery, Informatics, and Structural Life Science) from MEXT and AMED.
Author information
Authors and Affiliations
Contributions
T.O. and N.N.N. conceived the project. T.O. carried out protein expression, purification, crystallization, structure determination, and in vitro analyses of Atg2 and Atg18. N.N.N. performed microscopy of liposomes. T. Kawaoka, E.H., K.S., T. Kotani, H.N., and Y.O. performed the yeast experiments. T.O. and N.N.N. wrote the manuscript with input from other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Structural study of SpAtg2NR.
a, Secondary structure prediction of Chorein_N from several species. Chorein_N seems to conserve a fold quite similar to a part of SpAtg2NR. b, An artificial dimer found in SpAtg2NR crystals. SpAtg2NR dimerizes through formation of intermolecular antiparallel β-sheet.
Supplementary Figure 2 Structural study of PE-bound SpAtg2NR.
a, Structural superposition of SpAtg2NR and its PE-bound form. Both structures are represented by Cα-trace. b, Interaction between sn-2 acyl chains within the dimer found in crystal packing.
Supplementary Figure 3 Mutational analyses of ScAtg2.
a, SDS-PAGE of purified recombinant proteins used for in vitro analyses. Protein bands were stained with Coomassie Brilliant Blue. Uncropped gel images are shown in Supplementary Data Set 1. b, The effect of ATP on phospholipid transfer mediated by Atg2. c, Comparison of the MT activity between Atg2 and the Vps13 fragment analyzed by dynamic light scattering. d, MT activities of Atg2 as assessed by fluorescence microscopy of NBD-PE and Rhod-PE liposomes. Scale bar indicates 5 μm. e, Bulk autophagic activity measured by the ALP assay. In e and f, data are presented as mean±s.d. from n = 3 independent experiments. N.S. means not statistically significant (two-tailed Student’s test). f, Maturation of a selective autophagy cargo aminopeptidase I (Ape1) via autophagy monitored by western blotting. Uncropped blot image is shown in Supplementary Data Set 1. g, LT and MT activities of the K108E R216E mutant. h, Confocal microscopic analysis of liposomes mixed with the 3D mutant of ScAtg2. i, Circular dichroism spectra of wild-type and 3D mutant of MBP-fused SpAtg2NR.
Supplementary Figure 4 Structural feature of Atg2 supporting its lipid transfer activity.
a, Left, Displacement of the H4 helix creates a hydrophobic groove connecting the hydrophobic cavity and the membrane bound to the basic residues of Atg2NR. Arrows indicate a possible path for a phospholipid from the membrane to the cavity. Right, Superimposition of the structure of Vps13NR (PDB 6CBC) onto that of Atg2NR indicates the displacement of H4 in Vps13NR. b, Secondary structure prediction of ScAtg2 indicates that Atg2 contains repeated structures downstream of the N-terminal region that are abundant with β-strands. Boundaries of the repeated structures are roughly defined. The fifth repeated structure contains ATG2_CAD.
Supplementary information
Supplementary Information
Supplementary Figures 1–4, Supplementary Tables 1 and 2 and Supplementary Dataset 1
Supplementary Video 1
Effect of protease treatment on tethering of liposomes. Moving puncta of liposomes induced by Atg2 treatment as in Fig. 3h were subjected to proteinase K treatment, and NBD signals were monitored by fluorescence microscopy.
Source data
Rights and permissions
About this article
Cite this article
Osawa, T., Kotani, T., Kawaoka, T. et al. Atg2 mediates direct lipid transfer between membranes for autophagosome formation. Nat Struct Mol Biol 26, 281–288 (2019). https://doi.org/10.1038/s41594-019-0203-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0203-4
This article is cited by
-
Loss of WIPI4 in neurodegeneration causes autophagy-independent ferroptosis
Nature Cell Biology (2024)
-
ANKFY1 bridges ATG2A-mediated lipid transfer from endosomes to phagophores
Cell Discovery (2024)
-
ALLO-1- and IKKE-1-dependent positive feedback mechanism promotes the initiation of paternal mitochondrial autophagy
Nature Communications (2024)
-
Biogenesis of Rab14-positive endosome buds at Golgi–endosome contacts by the RhoBTB3–SHIP164–Vps26B complex
Cell Discovery (2024)
-
Complete set of the Atg8–E1–E2–E3 conjugation machinery forms an interaction web that mediates membrane shaping
Nature Structural & Molecular Biology (2024)