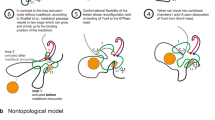
Abstract
The DNA loop extrusion model is a provocative new concept explaining the formation of chromatin loops that revolutionizes understanding of genome organization. Central to this model is the structural maintenance of chromosomes (SMC) protein family, which is now thought to function as a DNA motor. In this Perspective, we review and reinterpret the current knowledge of SMC structure and function and propose a novel mechanism for SMC motor activity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Riggs, A. D. DNA methylation and late replication probably aid cell memory, and type I DNA reeling could aid chromosome folding and enhancer function. Philos. Trans. R. Soc. Lond. B Biol. Sci. 326, 285–297 (1990).
Nasmyth, K. Disseminating the genome: joining, resolving, and separating sister chromatids during mitosis and meiosis. Annu. Rev. Genet. 35, 673–745 (2001).
Alipour, E. & Marko, J. F. Self-organization of domain structures by DNA-loop-extruding enzymes. Nucleic Acids Res. 40, 11202–11212 (2012).
Nichols, M. H. & Corces, V. G. A CTCF code for 3D genome architecture. Cell 162, 703–705 (2015).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl. Acad. Sci. USA 112, E6456–E6465 (2015).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320 (2017).
Haarhuis, J. H. I. et al. The cohesin release factor WAPL restricts chromatin loop extension. Cell 169, 693–707 (2017).
Wutz, G. et al. Topologically associating domains and chromatin loops depend on cohesin and are regulated by CTCF, WAPL, and PDS5 proteins. EMBO J. 36, 3573–3599 (2017).
Wang, X., Brandão, H. B., Le, T. B. K., Laub, M. T. & Rudner, D. Z. Bacillus subtilis SMC complexes juxtapose chromosome arms as they travel from origin to terminus. Science 355, 524–527 (2017).
Barrington, C., Finn, R. & Hadjur, S. Cohesin biology meets the loop extrusion model. Chromosome Res. 25, 51–60 (2017).
Racko, D., Benedetti, F., Dorier, J. & Stasiak, A. Transcription-induced supercoiling as the driving force of chromatin loop extrusion during formation of TADs in interphase chromosomes. Nucleic Acids Res. 46, 1648–1660 (2018).
Yamamoto, T. & Schiessel, H. Osmotic mechanism of the loop extrusion process. Phys. Rev. E 96, 030402 (2017).
Brackley, C. A. et al. Extrusion without a motor: a new take on the loop extrusion model of genome organization. Nucleus 9, 95–103 (2018).
Vian, L. et al. The energetics and physiological impact of cohesin extrusion. Cell 173, 1165–1178 (2018).
Wang, X. et al. In vivo evidence for ATPase-dependent DNA translocation by the Bacillus subtilis SMC condensin complex. Mol. Cell https://doi.org/10.1016/j.molcel.2018.07.006 (2018).
Terakawa, T. et al. The condensin complex is a mechanochemical motor that translocates along DNA. Science 358, 672–676 (2017).
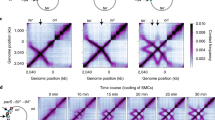
Ganji, M. et al. Real-time imaging of DNA loop extrusion by condensin. Science 360, 102–105 (2018). This study directly imaged real-time unidirectional loop extrusion by single condensin complexes. This represents the strongest evidence to date that SMC complexes are DNA motors and provides important insights into the mechanism by which extrusion occurs.
Stigler, J., Çamdere, G. Ö., Koshland, D. E. & Greene, E. C. Single-molecule imaging reveals a collapsed conformational state for DNA-bound cohesin. Cell Rep. 15, 988–998 2016).
Kanke, M., Tahara, E., Huis In’t Veld, P. J. & Nishiyama, T. Cohesin acetylation and Wapl–Pds5 oppositely regulate translocation of cohesin along DNA. EMBO J. 35, 2686–2698 (2016).
Davidson, I. F. et al. Rapid movement and transcriptional re-localization of human cohesin on DNA. EMBO J. 35, 2671–2685 (2016).
Yoshimura, S. H. & Hirano, T. HEAT repeats—versatile arrays of amphiphilic helices working in crowded environments? J. Cell Sci. 129, 3963–3970 (2016).
Hirano, M. & Hirano, T. Opening closed arms: long-distance activation of SMC ATPase by hinge-DNA interactions. Mol. Cell 21, 175–186 (2006).
Chiu, A., Revenkova, E. & Jessberger, R. DNA interaction and dimerization of eukaryotic SMC hinge domains. J. Biol. Chem. 279, 26233–26242 (2004).
Kschonsak, M. et al. Structural basis for a safety-belt mechanism that anchors condensin to chromosomes. Cell 171, 588–600 (2017).
Diebold-Durand, M.-L. et al. Structure of full-length SMC and rearrangements required for chromosome organization. Mol. Cell 67, 334–347 (2017).
Marko, J. F., Rios, P. D. L., Barducci, A. & Gruber, S. DNA-segment-capture model for loop extrusion by structural maintenance of chromosome (SMC) protein complexes. Preprint at bioRxiv https://doi.org/10.1101/325373 (2018).
Srinivasan, M. et al. The cohesin ring uses its hinge to organize DNA using non-topological as well as topological mechanisms. Cell 173, 1508–1519 (2018).
Kappel, C., Zachariae, U., Dölker, N. & Grubmüller, H. An unusual hydrophobic core confers extreme flexibility to HEAT repeat proteins. Biophys. J. 99, 1596–1603 (2010).
Chao, W. C. H. et al. Structure of the cohesin loader Scc2. Nat. Commun. 8, 13952 (2017).
Wells, J. N., Gligoris, T. G., Nasmyth, K. A. & Marsh, J. A. Evolution of condensin and cohesin complexes driven by replacement of Kite by Hawk proteins. Curr. Biol. 27, R17–R18 (2017).
Zabrady, K. et al. Chromatin association of the SMC5/6 complex is dependent on binding of its NSE3 subunit to DNA. Nucleic Acids Res. 44, 1064–1079 (2016).
Kamada, K., Miyata, M. & Hirano, T. Molecular basis of SMC ATPase activation: role of internal structural changes of the regulatory subcomplex ScpAB. Structure 21, 581–594 (2013).
Keenholtz, R. A. et al. Oligomerization and ATP stimulate condensin-mediated DNA compaction. Sci. Rep. 7, 14279 (2017).
Eeftens, J. M. et al. Real-time detection of condensin-driven DNA compaction reveals a multistep binding mechanism. EMBO J. 36, 3448–3457 (2017).
Strick, T. R., Kawaguchi, T. & Hirano, T. Real-time detection of single-molecule DNA compaction by condensin I. Curr. Biol. 14, 874–880 (2004).
Sun, M., Nishino, T. & Marko, J. F. The SMC1–SMC3 cohesin heterodimer structures DNA through supercoiling-dependent loop formation. Nucleic Acids Res. 41, 6149–6160 (2013).
Kim, H. & Loparo, J. J. Multistep assembly of DNA condensation clusters by SMC. Nat. Commun. 7, 10200 (2016).
Hopfner, K. P. & Tainer, J. A. Rad50/SMC proteins and ABC transporters: unifying concepts from high-resolution structures. Curr. Opin. Struct. Biol. 13, 249–255 (2003).
Kamada, K., Su’etsugu, M., Takada, H., Miyata, M. & Hirano, T. Overall shapes of the SMC–ScpAB complex are determined by balance between constraint and relaxation of its structural parts. Structure 25, 603–616 (2017).
Eeftens, J. M. et al. Condensin Smc2–Smc4 dimers are flexible and dynamic. Cell Rep. 14, 1813–1818 (2016).
Nasmyth, K. & Haering, C. H. The structure and function of SMC and kleisin complexes. Annu. Rev. Biochem. 74, 595–648 (2005).
Gerlich, D., Koch, B., Dupeux, F., Peters, J. M. & Ellenberg, J. Live-cell imaging reveals a stable cohesin-chromatin interaction after but not before DNA replication. Curr. Biol. 16, 1571–1578 (2006).
Kikuchi, S., Borek, D. M., Otwinowski, Z., Tomchick, D. R. & Yu, H. Crystal structure of the cohesin loader Scc2 and insight into cohesinopathy. Proc. Natl. Acad. Sci. USA 113, 12444–12449 (2016).
Schwarzer, W. et al. Two independent modes of chromatin organization revealed by cohesin removal. Nature 551, 51–56 (2017).
Petela, N. et al. Multiple interactions between Scc1 and Scc2 activate cohesin’s DNA dependent ATPase and replace Pds5 during loading. Preprint at bioRxiv https://doi.org/10.1101/205914 (2017).
Rao, S. S. P. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Acknowledgements
Work in the authors’ lab is supported by US Public Health Service Award R01 GM035463 from the National Institutes of Health. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Nichols, M.H., Corces, V.G. A tethered-inchworm model of SMC DNA translocation. Nat Struct Mol Biol 25, 906–910 (2018). https://doi.org/10.1038/s41594-018-0135-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0135-4
This article is cited by
-
Single cohesin molecules generate force by two distinct mechanisms
Nature Communications (2023)
-
Genome folding through loop extrusion by SMC complexes
Nature Reviews Molecular Cell Biology (2021)
-
Tandem CTCF sites function as insulators to balance spatial chromatin contacts and topological enhancer-promoter selection
Genome Biology (2020)
-
The condensin holocomplex cycles dynamically between open and collapsed states
Nature Structural & Molecular Biology (2020)