Abstract
The mitochondrial genome is transcribed by a single-subunit DNA-dependent RNA polymerase (mtRNAP) and its auxiliary factors. Structural studies have elucidated how mtRNAP cooperates with its dedicated transcription factors to direct RNA synthesis: initiation factors TFAM and TFB2M assist in promoter-DNA binding and opening by mtRNAP while the elongation factor TEFM increases polymerase processivity to the levels required for synthesis of long polycistronic mtRNA transcripts. Here, we review the emerging body of structural and functional studies of human mitochondrial transcription, provide a molecular movie that can be used for teaching purposes and discuss the open questions to guide future directions of investigation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Ernster, L. & Schatz, G. Mitochondria: a historical review. J. Cell Biol. 91, 227s–255s (1981).
Hamanaka, R. B. & Chandel, N. S. Mitochondrial reactive oxygen species regulate cellular signaling and dictate biological outcomes. Trends Biochem. Sci. 35, 505–513 (2010).
Pozzan, T. & Rizzuto, R. High tide of calcium in mitochondria. Nat. Cell Biol. 2, E25–E27 (2000).
Kroemer, G. & Reed, J. C. Mitochondrial control of cell death. Nat. Med. 6, 513–519 (2000).
Sun, N., Youle, R. J. & Finkel, T. The mitochondrial basis of aging. Mol. Cell 61, 654–666 (2016).
Nunnari, J. & Suomalainen, A. Mitochondria: in sickness and in health. Cell 148, 1145–1159 (2012).
Gray, M. W., Burger, G. & Lang, B. F. Mitochondrial evolution. Science 283, 1476–1481 (1999).
Falkenberg, M., Larsson, N.-G. & Gustafsson, C. M. DNA replication and transcription in mammalian mitochondria. Annu. Rev. Biochem. 76, 679–699 (2007).
Masters, B. S., Stohl, L. L. & Clayton, D. A. Yeast mitochondrial RNA polymerase is homologous to those encoded by bacteriophages T3 and T7. Cell 51, 89–99 (1987).
Chang, D. D. & Clayton, D. A. Precise identification of individual promoters for transcription of each strand of human mitochondrial DNA. Cell 36, 635–643 (1984).
Aloni, Y. & Attardi, G. Expression of the mitochondrial genome in HeLa cells. II. Evidence for complete transcription of mitochondrial DNA. J. Mol. Biol. 55, 251–267 (1971).
Aloni, Y. & Attardi, G. Symmetrical in vivo transcription of mitochondrial DNA in HeLa cells. Proc. Natl Acad. Sci. USA 68, 1757–1761 (1971).
Ojala, D., Montoya, J. & Attardi, G. tRNA punctuation model of RNA processing in human mitochondria. Nature 290, 470–474 (1981).
Kasamatsu, H., Robberson, D. L. & Vinograd, J. A novel closed-circular mitochondrial DNA with properties of a replicating intermediate. Proc. Natl Acad. Sci. USA 68, 2252–2257 (1971).
Chang, D. D. & Clayton, D. A. Priming of human mitochondrial DNA replication occurs at the light-strand promoter. Proc. Natl Acad. Sci. USA 82, 351–355 (1985).
Wanrooij, S. et al. Human mitochondrial RNA polymerase primes lagging-strand DNA synthesis in vitro. Proc. Natl Acad. Sci. USA 105, 11122–11127 (2008).
Fusté, J. M. et al. Mitochondrial RNA polymerase is needed for activation of the origin of light-strand DNA replication. Mol. Cell 37, 67–78 (2010).
Kühl, I. et al. POLRMT regulates the switch between replication primer formation and gene expression of mammalian mtDNA. Sci. Adv. 2, e1600963 (2016).
Gustafsson, C. M., Falkenberg, M. & Larsson, N.-G. Maintenance and expression of mammalian mitochondrial DNA. Annu. Rev. Biochem. 85, 133–160 (2016).
Sousa, R. Structural and mechanistic relationships between nucleic acid polymerases. Trends Biochem. Sci. 21, 186–190 (1996).
McAllister, W. T. & Raskin, C. A. The phage RNA polymerases are related to DNA polymerases and reverse transcriptases. Mol. Microbiol. 10, 1–6 (1993).
Sousa, R., Chung, Y. J., Rose, J. P. & Wang, B. C. Crystal structure of bacteriophage T7 RNA polymerase at 3.3 A resolution. Nature 364, 593–599 (1993).
Steitz, T. A., Smerdon, S. J., Jäger, J. & Joyce, C. M. A unified polymerase mechanism for nonhomologous DNA and RNA polymerases. Science 266, 2022–2025 (1994).
Sosunov, V. et al. Unified two-metal mechanism of RNA synthesis and degradation by RNA polymerase. EMBO J. 22, 2234–2244 (2003).
Tahirov, T. H. et al. Structure of a T7 RNA polymerase elongation complex at 2.9 A resolution. Nature 420, 43–50 (2002).
Yin, Y. W. & Steitz, T. A. Structural basis for the transition from initiation to elongation transcription in T7 RNA polymerase. Science 298, 1387–1395 (2002).
Cramer, P. Common structural features of nucleic acid polymerases. BioEssays 24, 724–729 (2002).
Cheetham, G. M., Jeruzalmi, D. & Steitz, T. A. Structural basis for initiation of transcription from an RNA polymerase-promoter complex. Nature 399, 80–83 (1999).
Cheetham, G. M. & Steitz, T. A. Structure of a transcribing T7 RNA polymerase initiation complex. Science 286, 2305–2309 (1999).
Ringel, R. et al. Structure of human mitochondrial RNA polymerase. Nature 478, 269–273 (2011). The crystal structure of human mitochondrial RNA polymerase demonstrates a resemblance to phage RNAPs and highlights differences that may explain factor dependence of mtRNAP for initiation.
Lightowlers, R. N. & Chrzanowska-Lightowlers, Z. M. A. PPR (pentatricopeptide repeat) proteins in mammals: important aids to mitochondrial gene expression. Biochem. J. 416, e5–e6 (2008).
Filipovska, A. & Rackham, O. Pentatricopeptide repeats: modular blocks for building RNA-binding proteins. RNA Biol. 10, 1426–1432 (2013).
Schwinghammer, K. et al. Structure of human mitochondrial RNA polymerase elongation complex. Nat. Struct. Mol. Biol. 20, 1298–1303 (2013). The structure of mtRNAP bound to nucleic acids reveals the overall architecture of the EC and highlights conserved catalytic mechanisms and striking differences from phage RNAPs.
Gnatt, A. L., Cramer, P., Fu, J., Bushnell, D. A. & Kornberg, R. D. Structural basis of transcription: an RNA polymerase II elongation complex at 3.3 A resolution. Science 292, 1876–1882 (2001).
Vassylyev, D. G., Vassylyeva, M. N., Perederina, A., Tahirov, T. H. & Artsimovitch, I. Structural basis for transcription elongation by bacterial RNA polymerase. Nature 448, 157–162 (2007).
Temiakov, D. et al. Structural basis for substrate selection by t7 RNA polymerase. Cell 116, 381–391 (2004).
Gaspari, M., Falkenberg, M., Larsson, N.-G. & Gustafsson, C. M. The mitochondrial RNA polymerase contributes critically to promoter specificity in mammalian cells. EMBO J. 23, 4606–4614 (2004).
Morozov, Y. I. et al. A novel intermediate in transcription initiation by human mitochondrial RNA polymerase. Nucleic Acids Res. 42, 3884–3893 (2014). This study provides the first evidence of a direct interaction between mtRNAP and TFAM, and describes the preinitiation complex.
Morozov, Y. I. et al. A model for transcription initiation in human mitochondria. Nucleic Acids Res. 43, 3726–3735 (2015). This biochemical study maps the interactions between mtRNAP and initiation factors in the IC.
Posse, V. & Gustafsson, C. M. Human mitochondrial transcription factor B2 is required for promoter melting during initiation of transcription. J. Biol. Chem. 292, 2637–2645 (2016). This study provides biochemical evidence that TFB2M is required for the initial melting of promoter DNA.
Ramachandran, A., Basu, U., Sultana, S., Nandakumar, D. & Patel, S. S. Human mitochondrial transcription factors TFAM and TFB2M work synergistically in promoter melting during transcription initiation. Nucleic Acids Res. 45, 861–874 (2017).
Sologub, M., Litonin, D., Anikin, M., Mustaev, A. & Temiakov, D. TFB2 is a transient component of the catalytic site of the human mitochondrial RNA polymerase. Cell 139, 934–944 (2009).
Parisi, M. A. & Clayton, D. A. Similarity of human mitochondrial transcription factor 1 to high mobility group proteins. Science 252, 965–969 (1991).
Dairaghi, D. J., Shadel, G. S., & Clayton, D. A. Addition of a 29 residue carboxyl-terminal tail converts a simple HMG box-containing protein into a transcriptional activator.J. Mol. Biol. 249, 11–28 (1995).This study provides an elegant biochemical demonstration that the C-terminal tail of TFAM is required for transcriptional activation .
Fisher, R. P., Topper, J. N. & Clayton, D. A. Promoter selection in human mitochondria involves binding of a transcription factor to orientation-independent upstream regulatory elements. Cell 50, 247–258 (1987).
Fisher, R. P. & Clayton, D. A. Purification and characterization of human mitochondrial transcription factor 1. Mol. Cell. Biol. 8, 3496–3509 (1988).
Alam, T. I. et al. Human mitochondrial DNA is packaged with TFAM. Nucleic Acids Res. 31, 1640–1645 (2003).
Kanki, T. et al. Architectural role of mitochondrial transcription factor A in maintenance of human mitochondrial DNA. Mol. Cell. Biol. 24, 9823–9834 (2004).
Kaufman, B. A. et al. The mitochondrial transcription factor TFAM coordinates the assembly of multiple DNA molecules into nucleoid-like structures. Mol. Biol. Cell 18, 3225–3236 (2007).
Kukat, C. et al. Cross-strand binding of TFAM to a single mtDNA molecule forms the mitochondrial nucleoid. Proc. Natl Acad. Sci. USA 112, 11288–11293 (2015).
Diffley, J. F. & Stillman, B. A close relative of the nuclear, chromosomal high-mobility group protein HMG1 in yeast mitochondria. Proc. Natl Acad. Sci. USA 88, 7864–7868 (1991).
Fisher, R. P., Lisowsky, T., Breen, G. A. & Clayton, D. A. A rapid, efficient method for purifying DNA-binding proteins: denaturation-renaturation chromatography of human and yeast mitochondrial extracts. J. Biol. Chem. 266, 9153–9160 (1991).
Rubio-Cosials, A. et al. Human mitochondrial transcription factor A induces a U-turn structure in the light strand promoter. Nat. Struct. Mol. Biol. 18, 1281–1289 (2011). A structure of human TFAM bound to a DNA stretch encompassing its binding site at LSP reveals that TFAM induces a 180° turn in the DNA (additional structure in ref. 54 ).
Ngo, H. B., Kaiser, J. T. & Chan, D. C. The mitochondrial transcription and packaging factor Tfam imposes a U-turn on mitochondrial DNA. Nat. Struct. Mol. Biol. 18, 1290–1296 (2011). A structure of human TFAM bound to a DNA stretch encompassing its binding site at LSP reveals that TFAM induces a 180° turn in the DNA (additional structure in ref. 53 ).
Ngo, H. B., Lovely, G. A., Phillips, R. & Chan, D. C. Distinct structural features of TFAM drive mitochondrial DNA packaging versus transcriptional activation. Nat. Commun. 5, 3077 (2014).
Morozov, Y. I. & Temiakov, D. Human mitochondrial transcription initiation complexes have similar topology on the light and heavy strand promoters. J. Biol. Chem. 291, 13432–13435 (2016).
Falkenberg, M. et al. Mitochondrial transcription factors B1 and B2 activate transcription of human mtDNA. Nat. Genet. 31, 289–294 (2002).
McCulloch, V., Seidel-Rogol, B. L. & Shadel, G. S. A human mitochondrial transcription factor is related to RNA adenine methyltransferases and binds S-adenosylmethionine. Mol. Cell. Biol. 22, 1116–1125 (2002).
Xu, B. & Clayton, D. A. Assignment of a yeast protein necessary for mitochondrial transcription initiation. Nucleic Acids Res. 20, 1053–1059 (1992).
Jang, S. H. & Jaehning, J. A. The yeast mitochondrial RNA polymerase specificity factor, MTF1, is similar to bacterial sigma factors. J. Biol. Chem. 266, 22671–22677 (1991).
Shadel, G. S. & Clayton, D. A. A. Saccharomyces cerevisiae mitochondrial transcription factor, sc-mtTFB, shares features with sigma factors but is functionally distinct. Mol. Cell. Biol. 15, 2101–2108 (1995).
Schubot, F. D. et al. Crystal structure of the transcription factor sc-mtTFB offers insights into mitochondrial transcription. Protein Sci. 10, 1980–1988 (2001). The structure of the yeast mitochondrial transcription initiation factor Mtf1 demonstrates its similarity to bacterial rRNA methyltransferases.
Seidel-Rogol, B. L., McCulloch, V. & Shadel, G. S. Human mitochondrial transcription factor B1 methylates ribosomal RNA at a conserved stem-loop. Nat. Genet. 33, 23–24 (2003).
Metodiev, M. D. et al. Methylation of 12S rRNA is necessary for in vivo stability of the small subunit of the mammalian mitochondrial ribosome. Cell Metab. 9, 386–397 (2009).
Litonin, D. et al. Human mitochondrial transcription revisited: only TFAM and TFB2M are required for transcription of the mitochondrial genes in vitro. J. Biol. Chem. 285, 18129–18133 (2010).
Hillen, H. S., Morozov, Y. I., Sarfallah, A., Temiakov, D. & Cramer, P. Structural basis of mitochondrial transcription initiation. Cell 171, 1072–1081.e10 (2017). Structures of TFB2M and the mitochondrial transcription IC elucidate how the initiation factors interact with mtRNAP to facilitate transcription initiation.
Cliften, P. F., Park, J. Y., Davis, B. P., Jang, S. H. & Jaehning, J. A. Identification of three regions essential for interaction between a sigma-like factor and core RNA polymerase. Genes Dev. 11, 2897–2909 (1997).
Guja, K. E. et al. Structural basis for S-adenosylmethionine binding and methyltransferase activity by mitochondrial transcription factor B1. Nucleic Acids Res. 41, 7947–7959 (2013).
Posse, V. et al. The amino terminal extension of mammalian mitochondrial RNA polymerase ensures promoter specific transcription initiation. Nucleic Acids Res. 42, 3638–3647 (2014).
Dairaghi, D. J., Shadel, G. S. & Clayton, D. A. Human mitochondrial transcription factor A and promoter spacing integrity are required for transcription initiation. Biochim. Biophys. Acta 1271, 127–134 (1995).
Yakubovskaya, E. et al. Organization of the human mitochondrial transcription initiation complex. Nucleic Acids Res. 42, 4100–4112
Feklistov, A. & Darst, S. A. Structural basis for promoter-10 element recognition by the bacterial RNA polymerase σ subunit. Cell 147, 1257–1269 (2011).
Zhang, Y. et al. Structural basis of transcription initiation. Science 338, 1076–1080 (2012).
Murakami, K. S. & Darst, S. A. Bacterial RNA polymerases: the wholo story. Curr. Opin. Struct. Biol. 13, 31–39 (2003).
Nayak, D., Guo, Q. & Sousa, R. A promoter recognition mechanism common to yeast mitochondrial and phage t7 RNA polymerases. J. Biol. Chem. 284, 13641–13647 (2009).
Minczuk, M. et al. TEFM (c17orf42) is necessary for transcription of human mtDNA. Nucleic Acids Res. 39, 4284–4299 (2011). This study identifies TEFM as a mitochondrial transcription elongation factor required for processive transcription of mtDNA.
Agaronyan, K., Morozov, Y. I., Anikin, M. & Temiakov, D. Mitochondrial biology: replication-transcription switch in human mitochondria. Science 347, 548–551 (2015).
Posse, V., Shahzad, S., Falkenberg, M., Hällberg, B. M. & Gustafsson, C. M. TEFM is a potent stimulator of mitochondrial transcription elongation in vitro. Nucleic Acids Res. 43, 2615–2624 (2015).
Wanrooij, P. H., Uhler, J. P., Simonsson, T., Falkenberg, M. & Gustafsson, C. M. G-quadruplex structures in RNA stimulate mitochondrial transcription termination and primer formation. Proc. Natl Acad. Sci. USA 107, 16072–16077 (2010). This study provides in vitro evidence that transcription termination at CSBII is mediated by formation of a G quadruplex in nascent RNA.
Kang, D., Miyako, K., Kai, Y., Irie, T. & Takeshige, K. In vivo determination of replication origins of human mitochondrial DNA by ligation-mediated polymerase chain reaction. J. Biol. Chem. 272, 15275–15279 (1997).
Hillen, H. S. et al. Mechanism of transcription anti-termination in human mitochondria. Cell 171, 1082–1093.e13 (2017). Biochemical and structural characterization of TEFM domains and of the processive antitermination complex reveals how TEFM confers processivity to mtRNAP.
Lockshon, D. et al. A role for recombination junctions in the segregation of mitochondrial DNA in yeast. Cell 81, 947–955 (1995).
Ceschini, S. et al. Crystal structure of the fission yeast mitochondrial Holliday junction resolvase Ydc2. EMBO J. 20, 6601–6611 (2001).
Górecka, K. M., Komorowska, W. & Nowotny, M. Crystal structure of RuvC resolvase in complex with Holliday junction substrate. Nucleic Acids Res. 41, 9945–9955 (2013).
Liu, B. & Steitz, T. A. Structural insights into NusG regulating transcription elongation. Nucleic Acids Res. 45, 968–974 (2017).
Martinez-Rucobo, F. W., Sainsbury, S., Cheung, A. C. M. & Cramer, P. Architecture of the RNA polymerase-Spt4/5 complex and basis of universal transcription processivity. EMBO J. 30, 1302–1310 (2011).
Bernecky, C., Plitzko, J. M. & Cramer, P. Structure of a transcribing RNA polymerase II–DSIF complex reveals a multidentate DNA–RNA clamp. Nat. Struct. Mol. Biol. 24, 809–815 (2017).
Werner, F. & Werner, F. A nexus for gene expression-molecular mechanisms of Spt5 and NusG in the three domains of life. J. Mol. Biol. 417, 13–27 (2012).
Grohmann, D. et al. The initiation factor TFE and the elongation factor Spt4/5 compete for the RNAP clamp during transcription initiation and elongation. Mol. Cell 43, 263–274 (2011).
Plaschka, C. et al. Transcription initiation complex structures elucidate DNA opening. Nature 533, 353–358 (2016).
Montoya, J., Gaines, G. L. & Attardi, G. The pattern of transcription of the human mitochondrial rRNA genes reveals two overlapping transcription units. Cell 34, 151–159 (1983).
Kruse, B., Narasimhan, N. & Attardi, G. Termination of transcription in human mitochondria: identification and purification of a DNA binding protein factor that promotes termination. Cell 58, 391–397 (1989).
Yakubovskaya, E., Mejia, E., Byrnes, J., Hambardjieva, E. & Garcia-Diaz, M. Helix unwinding and base flipping enable human MTERF1 to terminate mitochondrial transcription. Cell 141, 982–993 (2010). The crystal structure of MTERF1 bound to a sequence from the gene encoding tRNALeu reveals how MTERF1 wraps around the DNA and flips out bases from the DNA duplex (additional structure in ref. 100 ).
Asin-Cayuela, J., Schwend, T., Farge, G. & Gustafsson, C. M. The human mitochondrial transcription termination factor (mTERF) is fully active in vitro in the non-phosphorylated form. J. Biol. Chem. 280, 25499–25505 (2005).
Shang, J. & Clayton, D. A. Human mitochondrial transcription termination exhibits RNA polymerase independence and biased bipolarity in vitro. J. Biol. Chem. 269, 29112–29120 (1994).
Christianson, T. W. & Clayton, D. A. In vitro transcription of human mitochondrial DNA: accurate termination requires a region of DNA sequence that can function bidirectionally. Proc. Natl Acad. Sci. USA 83, 6277–6281 (1986).
Terzioglu, M. et al. MTERF1 binds mtDNA to prevent transcriptional interference at the light-strand promoter but is dispensable for rRNA gene transcription regulation. Cell Metab. 17, 618–626 (2013).
Shi, Y. et al. Mitochondrial transcription termination factor 1 directs polar replication fork pausing. Nucleic Acids Res. 44, 5732–5742 (2016).
Linder, T. et al. A family of putative transcription termination factors shared amongst metazoans and plants. Curr. Genet. 48, 265–269 (2005).
Jiménez-Menéndez, N. et al. Human mitochondrial mTERF wraps around DNA through a left-handed superhelical tandem repeat. Nat. Struct. Mol. Biol. 17, 891–893 (2010). This study reports a crystal structure of MTERF1 bound to a double-stranded DNA segment (additional structure in ref. 93 ).
Jemt, E. et al. Regulation of DNA replication at the end of the mitochondrial D-loop involves the helicase TWINKLE and a conserved sequence element. Nucleic Acids Res. 43, 9262–9275 (2015).
Park, C. B. et al. MTERF3 is a negative regulator of mammalian mtDNA transcription. Cell 130, 273–285 (2007).
Wredenberg, A. et al. MTERF3 regulates mitochondrial ribosome biogenesis in invertebrates and mammals. PLoS Genet. 9, e1003178 (2013).
Cámara, Y. et al. MTERF4 regulates translation by targeting the methyltransferase NSUN4 to the mammalian mitochondrial ribosome. Cell Metab. 13, 527–539 (2011).
Pellegrini, M. et al. MTERF2 is a nucleoid component in mammalian mitochondria. Biochim. Biophys. Acta 1787, 296–302 (2009).
Farge, G. et al. Protein sliding and DNA denaturation are essential for DNA organization by human mitochondrial transcription factor A. Nat Commun. 3, 1013 (2012).
Cotney, J. & Shadel, G. S. Evidence for an early gene duplication event in the evolution of the mitochondrial transcription factor B family and maintenance of rRNA methyltransferase activity in human mtTFB1 and mtTFB2. J. Mol. Evol. 63, 707–717 (2006).
Cermakian, N., Ikeda, T. M., Cedergren, R. & Gray, M. W. Sequences homologous to yeast mitochondrial and bacteriophage T3 and T7 RNA polymerases are widespread throughout the eukaryotic lineage. Nucleic Acids Res. 24, 648–654 (1996).
Cermakian, N. et al. On the evolution of the single-subunit RNA polymerases. J. Mol. Evol. 45, 671–681 (1997).
Shutt, T. E. & Gray, M. W. Bacteriophage origins of mitochondrial replication and transcription proteins. Trends Genet. 22, 90–95 (2006).
Lang, B. F. et al. An ancestral mitochondrial DNA resembling a eubacterial genome in miniature. Nature 387, 493–497 (1997).
Burger, G., Gray, M. W., Forget, L. & Lang, B. F. Strikingly bacteria-like and gene-rich mitochondrial genomes throughout jakobid protists. Genome Biol. Evol. 5, 418–438 (2013).
Zhelyazkova, P. et al. The primary transcriptome of barley chloroplasts: numerous noncoding RNAs and the dominating role of the plastid-encoded RNA polymerase. Plant Cell 24, 123–136 (2012).
Ropp, P. A. & Copeland, W. C. Cloning and characterization of the human mitochondrial DNA polymerase, DNA polymerase gamma. Genomics 36, 449–458 (1996).
Korhonen, J. A., Gaspari, M. & Falkenberg, M. TWINKLE has 5′ → 3′ DNA helicase activity and is specifically stimulated by mitochondrial single-stranded DNA-binding protein. J. Biol. Chem. 278, 48627–48632 (2003).
Maier, D. et al. Mitochondrial single-stranded DNA-binding protein is required for mitochondrial DNA replication and development in Drosophila melanogaster. Mol. Biol. Cell 12, 821–830 (2001).
Schilbach, S. et al. Structures of transcription pre-initiation complex with TFIIH and Mediator. Nature 551, 204–209 (2017).
Engel, C. et al. Structural basis of RNA Polymerase I transcription initiation. Cell 169, 120–131.e22 (2017).
Spåhr, H. et al. SLIRP stabilizes LRPPRC via an RRM-PPR protein interface. Nucleic Acids Res. 44, 6868–6882 (2016).
Rackham, O. et al. Hierarchical RNA processing is required for mitochondrial ribosome assembly. Cell Rep. 16, 1874–1890 (2016).
Pham, X. H. et al. Conserved sequence box II directs transcription termination and primer formation in mitochondria. J. Biol. Chem. 281, 24647–24652 (2006).
Tan, B. G., Wellesley, F. C., Savery, N. J. & Szczelkun, M. D. Length heterogeneity at conserved sequence block 2 in human mitochondrial DNA acts as a rheostat for RNA polymerase POLRMT activity. Nucleic Acids Res. 44, 7817–7829 (2016).
Gangelhoff, T. A., Mungalachetty, P. S., Nix, J. C. & Churchill, M. E. A. Structural analysis and DNA binding of the HMG domains of the human mitochondrial transcription factor A. Nucleic Acids Res. 37, 3153–3164 (2009).
Spåhr, H., Samuelsson, T., Hällberg, B. M. & Gustafsson, C. M. Structure of mitochondrial transcription termination factor 3 reveals a novel nucleic acid-binding domain. Biochem. Biophys. Res. Commun. 397, 386–390 (2010).
Murphy, W. I., Attardi, B., Tu, C. & Attardi, G. Evidence for complete symmetrical transcription in vivo of mitochondrial DNA in HeLa cells. J. Mol. Biol. 99, 809–814 (1975).
Hixson, J. E. & Clayton, D. A. Initiation of transcription from each of the two human mitochondrial promoters requires unique nucleotides at the transcriptional start sites. Proc. Natl Acad. Sci. USA 82, 2660–2664 (1985).
Fisher, R. P. & Clayton, D. A. A transcription factor required for promoter recognition by human mitochondrial RNA polymerase: accurate initiation at the heavy- and light-strand promoters dissected and reconstituted in vitro. J. Biol. Chem. 260, 11330–11338 (1985).
Topper, J. N. & Clayton, D. A. Identification of transcriptional regulatory elements in human mitochondrial DNA by linker substitution analysis. Mol. Cell. Biol. 9, 1200–1211 (1989).
Wang, Y. & Shadel, G. S. Stability of the mitochondrial genome requires an amino-terminal domain of yeast mitochondrial RNA polymerase. Proc. Natl Acad. Sci. USA 96, 8046–8051 (1999).
Acknowledgements
We wish to thank current and past members of the laboratories of P.C. and D.T. for critical discussions and valuable comments. H.S.H. was supported by a Boehringer Ingelheim Fonds PhD fellowship. P.C. was supported by the Deutsche Forschungsgemeinschaft (SFB860 and SPP1935), the European Research Council Advanced Investigator Grant TRANSREGULON (grant agreement 693023) and the Volkswagen Foundation. D.T. was supported by NIH RO1 GM104231 and R01 GM118941.
Author information
Authors and Affiliations
Contributions
H.S.H. prepared figures and Supplementary Video 1. H.S.H., D.T. and P.C. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Video 1
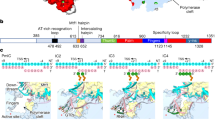
Structural basis of human mitochondrial transcription. A movie depicting events in the human mitochondrial transcription cycle based on known molecular structures. To initiate transcription, the mitochondrial RNA polymerase (PDB ID: 3SPA) is recruited to the promoter by TFAM (PDB ID: 3TMM and 3TQ6) via the tether helix in the mtRNAP N-terminal extension, to form the closed pre-initiation complex. Binding of TFB2M (PDB ID: 6ERO) then induces open initiation complex formation (PDB ID: 6ERP). Note that formation of the pre-initiation complex is not depicted due to a lack of structural information for this transient complex. During transition to elongation phase, the initiation factors are lost and the upstream DNA undergoes re-arrangement to occupy the site on mtRNAP previously bound by TFB2M, forming the elongation complex (PDB ID: 4BOC). Faithful transcription of long, polycistronic mitochondrial transcripts requires elongation factor TEFM, whose binding (only the active C-terminal domain is shown: PDB ID: 5OL8) results in formation of a processive anti-termination complex (PDB ID: 5OLA). Transcription termination is not shown due to lack of structural data on the interaction between mtRNAP and termination factor(s).
Rights and permissions
About this article
Cite this article
Hillen, H.S., Temiakov, D. & Cramer, P. Structural basis of mitochondrial transcription. Nat Struct Mol Biol 25, 754–765 (2018). https://doi.org/10.1038/s41594-018-0122-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0122-9
This article is cited by
-
Mitochondrially mediated RNA interference, a retrograde signaling system affecting nuclear gene expression
Heredity (2024)
-
Mechanisms and regulation of human mitochondrial transcription
Nature Reviews Molecular Cell Biology (2024)
-
Mitochondria transcription and cancer
Cell Death Discovery (2024)
-
Dynamic mitochondrial transcription and translation in B cells control germinal center entry and lymphomagenesis
Nature Immunology (2023)
-
Structures illustrate step-by-step mitochondrial transcription initiation
Nature (2023)