Abstract
Transient receptor potential vanilloid subfamily member 3 (TRPV3) channel plays a crucial role in skin physiology and pathophysiology. Mutations in TRPV3 are associated with various skin diseases, including Olmsted syndrome, atopic dermatitis, and rosacea. Here we present the cryo-electron microscopy structures of full-length mouse TRPV3 in the closed apo and agonist-bound open states. The agonist binds three allosteric sites distal to the pore. Channel opening is accompanied by conformational changes in both the outer pore and the intracellular gate. The gate is formed by the pore-lining S6 helices that undergo local α-to-π helical transitions, elongate, rotate, and splay apart in the open state. In the closed state, the shorter S6 segments are entirely α-helical, expose their nonpolar surfaces to the pore, and hydrophobically seal the ion permeation pathway. These findings further illuminate TRP channel activation and can aid in the design of drugs for the treatment of inflammatory skin conditions, itch, and pain.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Clapham, D. E. TRP channels as cellular sensors. Nature 426, 517–524 (2003).
Xu, H. X. et al. TRPV3 is a calcium-permeable temperature-sensitive cation channel. Nature 418, 181–186 (2002).
Peier, A. M. et al. A heat-sensitive TRP channel expressed in keratinocytes. Science 296, 2046–2049 (2002).
Smith, G. D. et al. TRPV3 is a temperature-sensitive vanilloid receptor-like protein. Nature 418, 186–190 (2002).
Cheng, X. et al. TRP channel regulates EGFR signaling in hair morphogenesis and skin barrier formation. Cell 141, 331–343 (2010).
Aijima, R. et al. The thermosensitive TRPV3 channel contributes to rapid wound healing in oral epithelia. FASEB J. 29, 182–192 (2015).
Imura, K. et al. Influence of TRPV3 mutation on hair growth cycle in mice. Biochem. Biophys. Res. Commun. 363, 479–483 (2007).
Liu, B., Yao, J., Zhu, M. X. & Qin, F. Hysteresis of gating underlines sensitization of TRPV3 channels. J. Gen. Physiol. 138, 509–520 (2011).
Ferrer-Montiel, A. et al. Advances in modulating thermosensory TRP channels. Expert Opin. Ther. Pat. 22, 999–1017 (2012).
Sherkheli, M. A., Vogt-Eisele, A. K., Weber, K. & Hatt, H. Camphor modulates TRPV3 cation channels activity by interacting with critical pore-region cysteine residues. Pak. J. Pharm. Sci. 26, 431–438 (2013).
Vogt-Eisele, A. K. et al. Monoterpenoid agonists of TRPV3. Br. J. Pharmacol. 151, 530–540 (2007).
Broad, L. M. et al. TRPV3 in drug development. Pharmaceuticals (Basel) 9, E55 (2016).
Doerner, J. F., Hatt, H. & Ramsey, I. S. Voltage- and temperature-dependent activation of TRPV3 channels is potentiated by receptor-mediated PI(4,5)P2 hydrolysis. J. Gen. Physiol. 137, 271–288 (2011).
Chung, M. K., Lee, H., Mizuno, A., Suzuki, M. & Caterina, M. J. 2-aminoethoxydiphenyl borate activates and sensitizes the heat-gated ion channel TRPV3. J. Neurosci. 24, 5177–5182 (2004).
Moqrich, A. et al. Impaired thermosensation in mice lacking TRPV3, a heat and camphor sensor in the skin. Science 307, 1468–1472 (2005).
Wang, G. & Wang, K. The Ca2+-permeable cation transient receptor potential TRPV3 channel: an emerging pivotal target for itch and skin diseases. Mol. Pharmacol. 92, 193–200 (2017).
Nelson, A. M. et al. 2-APB arrests human keratinocyte proliferation and inhibits cutaneous squamous cell carcinoma in vitro. Preprint at https://www.biorxiv.org/content/early/2018/01/23/249821 (2018).
Hu, H. Z. et al. 2-aminoethoxydiphenyl borate is a common activator of TRPV1, TRPV2, and TRPV3. J. Biol. Chem. 279, 35741–35748 (2004).
Hinman, A., Chuang, H. H., Bautista, D. M. & Julius, D. TRP channel activation by reversible covalent modification. Proc. Natl Acad. Sci. USA 103, 19564–19568 (2006).
Li, M., Jiang, J. & Yue, L. Functional characterization of homo- and heteromeric channel kinases TRPM6 and TRPM7. J. Gen. Physiol. 127, 525–537 (2006).
Kühn, F. J. P., Mathis, W., Cornelia, K., Hoffmann, D. C. & Lückhoff, A. Modulation of activation and inactivation by Ca2+ and 2-APB in the pore of an archetypal TRPM channel from Nematostella vectensis. Sci. Rep. 7, 7245 (2017).
Togashi, K., Inada, H. & Tominaga, M. Inhibition of the transient receptor potential cation channel TRPM2 by 2-aminoethoxydiphenyl borate (2-APB). Br. J. Pharmacol. 153, 1324–1330 (2008).
Lievremont, J. P., Bird, G. S. & Putney, J. W. Jr. Mechanism of inhibition of TRPC cation channels by 2-aminoethoxydiphenylborane. Mol. Pharmacol. 68, 758–762 (2005).
Chokshi, R., Fruasaha, P. & Kozak, J. A. 2-aminoethyl diphenyl borinate (2-APB) inhibits TRPM7 channels through an intracellular acidification mechanism. Channels (Austin) 6, 362–369 (2012).
Singh, A. K., Saotome, K., McGoldrick, L. L. & Sobolevsky, A. I. Structural bases of TRP channel TRPV6 allosteric modulation by 2-APB. Nat. Commun. 9, 2465 (2018).
Chuang, H. H. et al. Bradykinin and nerve growth factor release the capsaicin receptor from PtdIns(4,5)P2-mediated inhibition. Nature 411, 957–62 (2001).
Mercado, J., Gordon-Shaag, A., Zagotta, W. N. & Gordon, S. E. Ca2+-dependent desensitization of TRPV2 channels is mediated by hydrolysis of phosphatidylinositol 4,5-bisphosphate. J. Neurosci. 30, 13338–13347 (2010).
Garcia-Elias, A. et al. Phosphatidylinositol-4,5-biphosphate-dependent rearrangement of TRPV4 cytosolic tails enables channel activation by physiological stimuli. Proc. Natl Acad. Sci. USA 110, 9553–9558 (2013).
Lee, J., Cha, S. K., Sun, T. J. & Huang, C. L. PIP2 activates TRPV5 and releases its inhibition by intracellular Mg2+. J. Gen. Physiol. 126, 439–451 (2005).
Thyagarajan, B., Lukacs, V. & Rohacs, T. Hydrolysis of phosphatidylinositol 4,5-bisphosphate mediates calcium-induced inactivation of TRPV6 channels. J. Biol. Chem. 283, 14980–14987 (2008).
Xiao, R. et al. Calcium plays a central role in the sensitization of TRPV3 channel to repetitive stimulations. J. Biol. Chem. 283, 6162–6174 (2008).
Lin, Z. et al. Exome sequencing reveals mutations in TRPV3 as a cause of Olmsted syndrome. Am. J. Hum. Genet. 90, 558–564 (2012).
Ni, C. et al. A novel mutation in TRPV3 gene causes atypical familial Olmsted syndrome. Sci. Rep. 6, 21815 (2016).
Li, X. et al. Overexpression of TRPV3 correlates with tumor progression in non-small cell lung cancer. Int. J. Mol. Sci. 17, 437 (2016).
Huang, S. M. & Chung, M. K. Targeting TRPV3 for the development of novel analgesics. Open Pain J. 6, 119–126 (2013).
Reilly, R. M. & Kym, P. R. Analgesic potential of TRPV3 antagonists. Curr. Top. Med. Chem. 11, 2210–2215 (2011).
Goehring, A. et al. Screening and large-scale expression of membrane proteins in mammalian cells for structural studies. Nat. Protoc. 9, 2574–2585 (2014).
Shi, D. J., Ye, S., Cao, X., Zhang, R. & Wang, K. Crystal structure of the N-terminal ankyrin repeat domain of TRPV3 reveals unique conformation of finger 3 loop critical for channel function. Protein Cell 4, 942–950 (2013).
Gao, Y., Cao, E., Julius, D. & Cheng, Y. TRPV1 structures in nanodiscs reveal mechanisms of ligand and lipid action. Nature 534, 347–351 (2016).
Zubcevic, L. et al. Cryo-electron microscopy structure of the TRPV2 ion channel. Nat. Struct. Mol. Biol. 23, 180–186 (2016).
McGoldrick, L. L. et al. Opening of the human epithelial calcium channel TRPV6. Nature 553, 233–237 (2018).
Deng, Z. et al. Cryo-EM and X-ray structures of TRPV4 reveal insight into ion permeation and gating mechanisms. Nat. Struct. Mol. Biol. 25, 252–260 (2018).
Hughes, T. E. T. et al. Structural basis of TRPV5 channel inhibition by econazole revealed by cryo-EM. Nat. Struct. Mol. Biol. 25, 53–60 (2018).
Huynh, K. W. et al. Structure of the full-length TRPV2 channel by cryo-EM. Nat. Commun. 7, 11130 (2016).
Yin, Y. et al. Structure of the cold- and menthol-sensing ion channel TRPM8. Science 359, 237–241 (2018).
Grandl, J. et al. Pore region of TRPV3 ion channel is specifically required for heat activation. Nat. Neurosci. 11, 1007–1013 (2008).
Saotome, K., Singh, A. K., Yelshanskaya, M. V. & Sobolevsky, A. I. Crystal structure of the epithelial calcium channel TRPV6. Nature 534, 506–511 (2016).
Cao, E., Liao, M., Cheng, Y. & Julius, D. TRPV1 structures in distinct conformations reveal activation mechanisms. Nature 504, 113–118 (2013).
Luo, J., Stewart, R., Berdeaux, R. & Hu, H. Tonic inhibition of TRPV3 by Mg2+ in mouse epidermal keratinocytes. J. Invest. Dermatol. 132, 2158–2165 (2012).
Hu, H. Z., Grandl, J., Bandell, M., Petrus, M. & Patapoutian, A. Two amino acid residues determine 2-APB sensitivity of the ion channels TRPV3 and TRPV4. Proc. Natl Acad. Sci. USA 106, 1626–1631 (2009).
Hughes, T. E. T. et al. Structural basis of TRPV5 channel inhibition by econazole revealed by cryo-EM. Nat. Struct. Mol. Biol. 25, 53–60 (2018).
Jiang, Y. et al. X-ray structure of a voltage-dependent K+ channel. Nature 423, 33–41 (2003).
Jiang, Y. et al. Crystal structure and mechanism of a calcium-gated potassium channel. Nature 417, 515–522 (2002).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360 (1996).
Singh, A. K., Saotome, K. & Sobolevsky, A. I. Swapping of transmembrane domains in the epithelial calcium channel TRPV6. Sci. Rep. 7, 10669 (2017).
Russo, C. J. & Passmore, L. A. Electron microscopy: ultrastable gold substrates for electron cryomicroscopy. Science 346, 1377–1380 (2014).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Kimanius, D., Forsberg, B. O., Scheres, S. H. W. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016).
Scheres, S. H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012).
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D Biol. Crystallogr. 68, 352–367 (2012).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Acknowledgements
We thank H. Kao for computational support, R. Grassucci and F. Acosta-Reyes for assistance with microscope operation, and U. Baxa and T. Edwards for help with data collection. L.L.M. is supported by a National Institutes of Health grant (T32 GM008224). A.I.S. is supported by two NIH grants (R01 CA206573 and R01 NS083660), the Amgen Young Investigator Award, and the Irma T. Hirschl Career Scientist Award. Data were collected at the Frederick National Laboratory for Cancer Research National Cryo-EM Facility (NIH) and at the Simons Electron Microscopy Center and National Resource for Automated Molecular Microscopy (New York Structural Biology Center), supported by grants from the Simons Foundation (349247), NYSTAR, and the NIH (GM103310).
Author information
Authors and Affiliations
Contributions
A.K.S., L.L.M., and A.I.S. designed the project. A.K.S. and L.L.M. carried out protein expression, purification, cryo-EM sample preparation, and data collection. A.K.S., L.L.M., and A.I.S. processed and analyzed the cryo-EM data. A.K.S., L.L.M., and A.I.S. built the models and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Functional characterization of wild-type and mutant TRPV3.
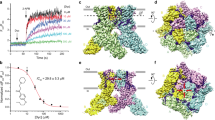
a–c, Representative Fura-2 AM ratiometric fluorescence measurements of calcium uptake for HEK cells expressing wild-type TRPV3 (a), TRPV3(Y564A) (b) and TRPV3(H426A) (c). Changes in fluorescence were recorded in response to applications of 2-APB (a–c) at different concentrations (indicated) or 5 mM camphor (c). Arrows indicate the time at which 2-APB or camphor was added. d, Dose–response curves for wild-type and mutant TRPV3 channel activation by 2-APB. For TRPV3(H426A) and TRPV3(R487A), the changes in F340/F380 elicited by the addition of 2-APB were normalized to the maximal change in F340/F380 in response to application of 5 mM camphor. For wild-type and other TRPV3 mutants, the changes in F340/F380 were normalized to their approximated maximal values at saturating concentrations of 2-APB. Curves through the points are fits with the logistic equation, with the mean ± s.e.m. values of the half-maximal effective concentration (EC50), 27.4 ± 4.5 μM (n = 4) for TRPV3, 1.35 ± 0.07 μM (n = 3) for TRPV3(Y564A), 33.1 ± 2.6 μM (n = 3) for TRPV3(H426A-Y564A), 97.8 ± 14.7 μM (n = 3) for TRPV3(Y540A), 147 ± 29 μM (n = 3) for TRPV3(Q483A) and 460 ± 20 μM (n = 3) for TRPV3(R487A). Each data point represents the average and s.e.m. of three independent measurements; for TRPV3(Y564A), the error bars were smaller than the symbol size
Supplementary Figure 2 Overview of single-particle cryo-EM for TRPV3.
a, Example cryo-EM micrograph for TRPV3 with example particles circled in red. b, Reference-free 2D class averages illustrating different particle orientations. c, Local resolution mapped on TRPV3 density at a 0.010 threshold level (UCSF Chimera) calculated using ResMap (Bioinformatics 21, 3327–3328, 2005) and two unfiltered half maps, with the highest resolution observed for the channel core. d, Orientation distribution of particles that contributed to the final 4.3-Å reconstruction; longer red rods represent orientations that comprise more particles. e, FSC curve calculated between half maps. e, Cross-validation FSC curves for the refined model versus unfiltered half maps (only half map 1 was used for refinement with PHENIX (Acta Crystallogr. D Biol. Crystallogr. 68, 352–367, 2012)) and the unfiltered summed map
Supplementary Figure 3 Cryo-EM density of TRPV3.
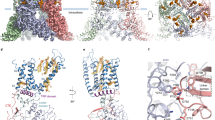
a, Cryo-EM density (blue mesh) at 4σ for a single TRPV3 subunit; the structure, shown as a ribbon, is colored according to domain. b–h, Fragments of the TRPV3 transmembrane and C-terminal domains with the corresponding cryo-EM density shown as a blue mesh at 4σ
Supplementary Figure 4 Structural comparison of TRPV channels.
a–j, Comparison of the structures of mouse TRPV3 in the closed state (yellow) and rat TRPV1 in the closed state (red; PDB 3J5P) (a,f), rabbit TRPV2 in a putative desensitized state (teal; PDB 5AN8) (b,g), Xenopus tropicalis TRPV4 in the closed state (purple; PDB 6BBJ) (c,h), rabbit TRPV5 in the closed state (green; PDB 6B5V) (d,i) and human TRPV6 in the open state (pink; PDB 6BO8) (e,j) viewed intracellularly (a–e) or parallel to the membrane (f–j). In f–j, only the TMD and the TRP helix from one subunit are shown. The structures are aligned based on their pore domains
Supplementary Figure 5 Sequence alignment of mouse TRPV channels.
Above the sequences, α-helices and β-strands are depicted as cylinders and arrows, respectively. The *, ¥ and Ω symbols indicate residues contributing to sites 2, 3 and 4, respectively. The location of the selectivity filter (S.F.) is indicated by a red box
Supplementary Figure 6 Comparison of the pore region density in different cryo-EM reconstructions.
a–c, Fragments of cryo-EM density for the S6 and TRP helices in TRPV3 (a), TRPV3(Y564A)2-APB (b) and TRPV32-APB (c) viewed intracellularly. The density for M677, which forms the gate in the closed state, is indicated by arrows. d–f, Fragments of cryo-EM density for the S5, S6 and TRP helices in TRPV3 (d), TRPV3(Y564A)2-APB (e) and TRPV3(Y564A) (f), all filtered to the same (6.5–Å) resolution
Supplementary Figure 7 Overview of single-particle cryo-EM for TRPV3(Y564A)2-APB.
a, Example cryo-EM micrograph for TRPV3(Y564A)2-APB with example particles circled in red. b, Reference-free 2D class averages illustrating different particle orientations. c, Local resolution mapped on TRPV3(Y564A)2-APB density at a 0.012 threshold level (UCSF Chimera) calculated using ResMap (Bioinformatics 21, 3327–3328, 2005) and two unfiltered half maps, with the highest resolution observed for the channel core. d, Orientation distribution of particles that contributed to the final 4.1-Å reconstruction; longer red rods represent orientations that comprise more particles. e, FSC curve calculated between half maps. e, Cross-validation FSC curves for the refined model versus unfiltered half maps (only half map 1 was used for refinement with PHENIX (Acta Crystallogr. D Biol. Crystallogr. 68, 352–367, 2012)) and the unfiltered summed map
Supplementary Figure 8 Comparison of sites 2–4 in TRPV3 and TRPV3(Y564A)2-APB.
a,b, Extracellular part of the S1–S4 domain in TRPV3 (a) and TRPV3(Y564A)2-APB (b) viewed parallel to the membrane with cryo-EM density shown as blue mesh at 4σ. Note that the S1–S2 loop occupies site 4 in TRPV3 (a). In TRPV3(Y564A)2-APB (b), this loop is displaced upward and site 4 is occupied by a nonprotein density into which a 2-APB molecule (green sticks) can be easily fit. c, Superposition of the region in one subunit encompassing sites 2 and 3 of TRPV3 (blue) and TRPV3(Y564A)2-APB (orange). 2-APB molecules and residues forming sites 2 and 3 are shown as sticks. Note that, while S5 and S6 show significant rearrangements during gating, sites 2 and 3 remain nearly intact
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8
Rights and permissions
About this article
Cite this article
Singh, A.K., McGoldrick, L.L. & Sobolevsky, A.I. Structure and gating mechanism of the transient receptor potential channel TRPV3. Nat Struct Mol Biol 25, 805–813 (2018). https://doi.org/10.1038/s41594-018-0108-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0108-7
This article is cited by
-
Structural basis of TRPV3 inhibition by an antagonist
Nature Chemical Biology (2023)
-
A pentameric TRPV3 channel with a dilated pore
Nature (2023)
-
Structural basis of ligand activation and inhibition in a mammalian TRPV4 ion channel
Cell Discovery (2023)
-
Structural mechanisms of TRPV2 modulation by endogenous and exogenous ligands
Nature Chemical Biology (2023)
-
Structure of human TRPV4 in complex with GTPase RhoA
Nature Communications (2023)