Abstract
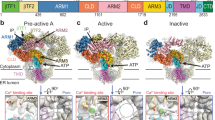
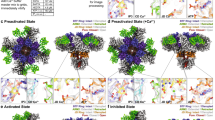
Inositol trisphosphate receptors (IP3Rs) are ubiquitous Ca2+-permeable channels that mediate release of Ca2+ from the endoplasmic reticulum, thereby regulating numerous processes including cell division, cell death, differentiation and fertilization. IP3Rs are jointly activated by inositol trisphosphate (IP3) and their permeant ion, Ca2+. At high concentrations, however, Ca2+ inhibits activity, ensuring precise spatiotemporal control over intracellular Ca2+. Despite extensive characterization of IP3R, the mechanisms through which these molecules control channel gating have remained elusive. Here, we present structures of full-length human type 3 IP3Rs in ligand-bound and ligand-free states. Multiple IP3-bound structures demonstrate that the large cytoplasmic domain provides a platform for propagation of long-range conformational changes to the ion-conduction gate. Structures in the presence of Ca2+ reveal two Ca2+-binding sites that induce the disruption of numerous interactions between subunits, thereby inhibiting IP3R. These structures thus provide a mechanistic basis for beginning to understand the regulation of IP3R.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Change history
08 August 2018
In the version of this article initially published, one of the PDB codes associated with the study was listed in the data availability statement as 6DBC but should have been 6DRC. The error has been corrected in the HTML and PDF versions of the article.
References
Burgess, G. M., McKinney, J. S., Fabiato, A., Leslie, B. A. & Putney, J. W. Jr. Calcium pools in saponin-permeabilized guinea pig hepatocytes. J. Biol. Chem. 258, 15336–15345 (1983).
Ehrlich, B. E. & Watras, J. Inositol 1,4,5-trisphosphate activates a channel from smooth muscle sarcoplasmic reticulum. Nature 336, 583–586 (1988).
Furuichi, T. et al. Primary structure and functional expression of the inositol 1,4,5-trisphosphate-binding protein P400. Nature 342, 32–38 (1989).
Mignery, G. A., Südhof, T. C., Takei, K. & De Camilli, P. Putative receptor for inositol 1,4,5-trisphosphate similar to ryanodine receptor. Nature 342, 192–195 (1989).
Streb, H., Irvine, R. F., Berridge, M. J. & Schulz, I. Release of Ca2+ from a nonmitochondrial intracellular store in pancreatic acinar cells by inositol-1,4,5-trisphosphate. Nature 306, 67–69 (1983).
Foskett, J. K., White, C., Cheung, K.-H. & Mak, D.-O. D. Inositol trisphosphate receptor Ca2+ release channels. Physiol. Rev. 87, 593–658 (2007).
Vervloessem, T., Yule, D. I., Bultynck, G. & Parys, J. B. The type 2 inositol 1,4,5-trisphosphate receptor, emerging functions for an intriguing Ca2+-release channel. Biochim. Biophys. Acta 1853, 1992–2005 (2015).
Ferris, C. D., Huganir, R. L., Supattapone, S. & Snyder, S. H. Purified inositol 1,4,5-trisphosphate receptor mediates calcium flux in reconstituted lipid vesicles. Nature 342, 87–89 (1989).
Finch, E. A., Turner, T. J. & Goldin, S. M. Calcium as a coagonist of inositol 1,4,5-trisphosphate-induced calcium release. Science 252, 443–446 (1991).
Jean, T. & Klee, C. B. Calcium modulation of inositol 1,4,5-trisphosphate-induced calcium release from neuroblastoma x glioma hybrid (NG108-15) microsomes. J. Biol. Chem. 261, 16414–16420 (1986).
Suematsu, E., Hirata, M., Hashimoto, T. & Kuriyama, H. Inositol 1,4,5-trisphosphate releases Ca2+ from intracellular store sites in skinned single cells of porcine coronary artery. Biochem. Biophys. Res. Commun. 120, 481–485 (1984).
Armant, D. R. Intracellular Ca2+ signaling and preimplantation development. Adv. Exp. Med. Biol. 843, 151–171 (2015).
Berridge, M. J. Calcium signalling remodelling and disease. Biochem. Soc. Trans. 40, 297–309 (2012).
Kania, E., Roest, G., Vervliet, T., Parys, J. B. & Bultynck, G. IP3 receptor-mediated calcium signaling and its role in autophagy in cancer. Front. Oncol. 7, 140 (2017).
Ando, H., Kawaai, K., Bonneau, B. & Mikoshiba, K. Remodeling of Ca2+signaling in cancer: regulation of inositol 1,4,5-trisphosphate receptors through oncogenes and tumor suppressors. Adv. Biol. Regul. 68, 64–76 (2018).
Berridge, M. J. The inositol trisphosphate/calcium signaling pathway in health and disease. Physiol. Rev. 96, 1261–1296 (2016).
Egorova, P.A. & Bezprozvanny, I.B. Inositol 1,4,5-trisphosphate receptors and neurodegenerative disorders. FEBS J. https://doi.org/10.1111/febs.14366 (2017).
Bezprozvanny, I., Watras, J. & Ehrlich, B. E. Bell-shaped calcium-response curves of Ins(1,4,5)P3- and calcium-gated channels from endoplasmic reticulum of cerebellum. Nature 351, 751–754 (1991).
Mikoshiba, K. Role of IP3 receptor signaling in cell functions and diseases. Adv. Biol. Regul. 57, 217–227 (2015).
Gerasimenko, O. V., Gerasimenko, J. V., Belan, P. V. & Petersen, O. H. Inositol trisphosphate and cyclic ADP-ribose-mediated release of Ca2+ from single isolated pancreatic zymogen granules. Cell 84, 473–480 (1996).
Iino, M. Biphasic Ca2+ dependence of inositol 1,4,5-trisphosphate-induced Ca release in smooth muscle cells of the guinea pig taenia caeci. J. Gen. Physiol. 95, 1103–1122 (1990).
Stehno-Bittel, L., Lückhoff, A. & Clapham, D. E. Calcium release from the nucleus by InsP3 receptor channels. Neuron 14, 163–167 (1995).
Bootman, M. D., Berridge, M. J. & Lipp, P. Cooking with calcium: the recipes for composing global signals from elementary events. Cell 91, 367–373 (1997).
Marchant, J. S. & Parker, I. Role of elementary Ca2+ puffs in generating repetitive Ca2+ oscillations. EMBO J. 20, 65–76 (2001).
Parker, I., Choi, J. & Yao, Y. Elementary events of InsP3-induced Ca2+ liberation in Xenopus oocytes: hot spots, puffs and blips. Cell Calcium 20, 105–121 (1996).
Hajnóczky, G. & Thomas, A. P. Minimal requirements for calcium oscillations driven by the IP3 receptor. EMBO J. 16, 3533–3543 (1997).
Marshall, I. C. & Taylor, C. W. Two calcium-binding sites mediate the interconversion of liver inositol 1,4,5-trisphosphate receptors between three conformational states. Biochem. J. 301, 591–598 (1994).
Sienaert, I. et al. Molecular and functional evidence for multiple Ca2+-binding domains in the type 1 inositol 1,4,5-trisphosphate receptor. J. Biol. Chem. 272, 25899–25906 (1997).
Striggow, F. & Ehrlich, B. E. The inositol 1,4,5-trisphosphate receptor of cerebellum. Mn2+ permeability and regulation by cytosolic Mn2+. J. Gen. Physiol. 108, 115–124 (1996).
Hamada, K., Terauchi, A. & Mikoshiba, K. Three-dimensional rearrangements within inositol 1,4,5-trisphosphate receptor by calcium. J. Biol. Chem. 278, 52881–52889 (2003).
Shinohara, T. et al. Mechanistic basis of bell-shaped dependence of inositol 1,4,5-trisphosphate receptor gating on cytosolic calcium. Proc. Natl. Acad. Sci. USA 108, 15486–15491 (2011).
Mak, D.-O. D., McBride, S. & Foskett, J. K. Inositol 1,4,5-tris-phosphate activation of inositol tris-phosphate receptor Ca2+ channel by ligand tuning of Ca2+ inhibition. Proc. Natl. Acad. Sci. USA 95, 15821–15825 (1998).
Maranto, A. R. Primary structure, ligand binding, and localization of the human type 3 inositol 1,4,5-trisphosphate receptor expressed in intestinal epithelium. J. Biol. Chem. 269, 1222–1230 (1994).
Bosanac, I. et al. Structure of the inositol 1,4,5-trisphosphate receptor binding core in complex with its ligand. Nature 420, 696–700 (2002).
Bosanac, I. et al. Crystal structure of the ligand binding suppressor domain of type 1 inositol 1,4,5-trisphosphate receptor. Mol. Cell 17, 193–203 (2005).
Lin, C.-C., Baek, K. & Lu, Z. Apo and InsP3-bound crystal structures of the ligand-binding domain of an InsP3 receptor. Nat. Struct. Mol. Biol. 18, 1172–1174 (2011).
Seo, M.-D. et al. Structural and functional conservation of key domains in InsP3 and ryanodine receptors. Nature 483, 108–112 (2012).
Fan, G. et al. Gating machinery of InsP3R channels revealed by electron cryomicroscopy. Nature 527, 336–341 (2015).
Scheres, S. H. W. Processing of structurally heterogeneous cryo-EM data in RELION. Methods Enzymol. 579, 125–157 (2016).
des Georges, A. et al. Structural basis for gating and activation of RyR1. Cell 167, 145–157.e17 (2016).
Prole, D. L. & Taylor, C. W. Inositol 1,4,5-trisphosphate receptors and their protein partners as signalling hubs. J. Physiol. 594, 2849–2866 (2016).
Boehning, D. & Joseph, S. K. Direct association of ligand-binding and pore domains in homo- and heterotetrameric inositol 1,4,5-trisphosphate receptors. EMBO J. 19, 5450–5459 (2000).
Chan, J. et al. Structural studies of inositol 1,4,5-trisphosphate receptor: coupling ligand binding to channel gating. J. Biol. Chem. 285, 36092–36099 (2010).
Uchida, K., Miyauchi, H., Furuichi, T., Michikawa, T. & Mikoshiba, K. Critical regions for activation gating of the inositol 1,4,5-trisphosphate receptor. J. Biol. Chem. 278, 16551–16560 (2003).
Yamazaki, H., Chan, J., Ikura, M., Michikawa, T. & Mikoshiba, K. Tyr-167/Trp-168 in type 1/3 inositol 1,4,5-trisphosphate receptor mediates functional coupling between ligand binding and channel opening. J. Biol. Chem. 285, 36081–36091 (2010).
McGoldrick, L. L. et al. Opening of the human epithelial calcium channel TRPV6. Nature 553, 233–237 (2018).
Ionescu, L. et al. Mode switching is the major mechanism of ligand regulation of InsP3 receptor calcium release channels. J. Gen. Physiol. 130, 631–645 (2007).
Goehring, A. et al. Screening and large-scale expression of membrane proteins in mammalian cells for structural studies. Nat. Protoc. 9, 2574–2585 (2014).
Whicher, J. R. & MacKinnon, R. Structure of the voltage-gated K+ channel Eag1 reveals an alternative voltage sensing mechanism. Science 353, 664–669 (2016).
Kirchhofer, A. et al. Modulation of protein properties in living cells using nanobodies. Nat. Struct. Mol. Biol. 17, 133–138 (2010).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Grigorieff, N., Grant, T. & Rohou, A. cisTEM: user-friendly software for single-particle image processing. elife. 7, e35383 (2018).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Pettersen, E. F. et al. UCSF Chimera: a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
Drozdetskiy, A., Cole, C., Procter, J. & Barton, G. J. JPred4: a protein secondary structure prediction server. Nucleic Acids Res .43, W389–W394 (2015).
Wang, R. Y. R. et al. Automated structure refinement of macromolecular assemblies from cryo-EM maps using Rosetta. eLife 5, e17219 (2016).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Morin, A. et al. Collaboration gets the most out of software. eLife 2, e01456 (2013).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Acknowledgements
We thank M. de le Cruz at the Memorial Sloan Kettering Cancer Center Cryo-EM facility and staff at the New York Structural Biology Simons Electron Microscopy Center for help with data collection, and S.B. Long and T. Walz for comments on the manuscript. This work was supported in part by NIH-NCI Cancer Center Support Grant (P30 CA008748), the Josie Robertson Investigators Program (to R.K.H.) and the Searle Scholars Program (to R.K.H).
Author information
Authors and Affiliations
Contributions
N.P. and R.K.H. designed, performed and analyzed the experiments. N.P. and R.K.H. prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Cryo-EM analysis of apo-hIP3R3.
a, Representative raw image and 2D averages of apo hIP3R3. b, Particle selection and focused refinement procedures used to obtain full-channel and focused refinement structures of apo hIP3R3. c, Full-channel reconstruction colored by local resolution estimation from ResMap. d, CD focused refinement reconstruction colored by local resolution estimation from ResMap. e, S1-S4 focused refinement reconstruction colored by local resolution estimation from ResMap. f, FSC plot of full channel (red), CD focused refinement (blue) and S1-S4 focused refinement reconstruction (green). g, FSC plot of full channel reconstruction of all particles compared with the refined model (FSC sum, black), FSC plot of half-map1 compared with the refined model (FSC work, red) and FSC plot of half-map2 compared with the refined model (FSC free, blue) generated by phenix.mtriage.
Supplementary Figure 2 C-terminal domain in apo-hIP3R3.
Cryo-EM density map of apo hIP3R3 lowpass filtered to (a) 3.5 Å and (b) 5.5 Å. Structure of apo hIP3R3 is colored by domain. Boxed region highlights caboxy-terminal domain, which is less well resolved than neighboring regions due to its extensive conformational flexibility.
Supplementary Figure 3 S1′ and S1′′ transmembrane helices in apo-hIP3R3.
a, S1-S4 focused refinement density for S1’, S1” and ordered lipids. One subunit is colored by domain. S1-S4 domain is red, S1’ and S1” are blue and lipid molecules are yellow. Density is contoured at 8σ. b, c, Structural diversity of S1-S2 linker in 6 TM ions channels40,65–68. S1 and S2 colored red and S1-S2 linker colored in blue. Unregistered transmembrane helix in the ryanodine receptor structure is colored magenta. d, Sequence alignment of S1-S2 linker domain of hIP3R1, hIP3R2, hIP3R3 and human type 1 and type 2 ryanodine receptor. 65. Long, S. B., Tao, X., Campbell, E. B. & MacKinnon, R. Atomic structure of a voltage-dependent K+channel in a lipid membrane-like environment. Nature 450, 376–382 (2007). 66. Tang, L. et al. Structural basis for inhibition of a voltage-gated Ca2+ channel by Ca2+ antagonist drugs. Nature 537, 117–121 (2016). 67. Hite, R. K. & MacKinnon, R. Structural Titration of Slo2.2, a Na+-Dependent K+ Channel. Cell 168, 390–399.e11 (2017). 68. Hirschi, M. et al. Cryo-electron microscopy structure of the lysosomal calcium-permeable channel TRPML3. Nature 550, 411–414 (2017).
Supplementary Figure 4 Cryo-EM analysis of IP3-bound hIP3R3.
a, Representative raw image and 2D averages of IP3-bound hIP3R3. b, Initial reference for 3D classification lacking the ARM2 domains and low-pass filtered to 60 Å. c, Particle selection and iterative refinement procedures used to obtain structures of IP3 classes 1, 2, 3, 4 and 5. d, Reconstructions colored by local resolution estimation from ResMap. e, FSC plots of full channel (red) and CD focused refinement (blue) reconstructions. f, FSC plots of all particles compared with the refined model (FSC sum, black), FSC plot of half-map1 compared with the refined model (FSC work, red) and FSC plot of half-map2 compared with the refined model (FSC free, blue) generated by phenix.mtriage.
Supplementary Figure 5 IP3-binding domain.
a-f, IP3-binding site in (a) apo, (b) IP3 class 1, and (c) IP3 class 2, (d) IP3 class 3, (e) IP3 class 4 and (f) IP3 class 5. Coordinating side chains and IP3 are shown as sticks and colored by element. h, Superposition of ligand-free IP3-binding domain structures aligned by BTF1 and BTF2. g, Superposition of IP3-binding domain structures aligned by BTF1 and BTF2. IP3 is shown in sticks.
Supplementary Figure 6 Cryo-EM analysis of Ca2+-bound hIP3R3.
a, Representative size exclusion chromatography traces of hIP3R3 in 5 mM EGTA (red) and 1 mM Ca2+ (blue). b, Representative raw image and 2D averages of Ca2+-bound hIP3R3. c, Particle selection and focused refinement procedures used to obtain full-channel and focused refinement structures of Ca2+-bound hIP3R3. d, Full-channel reconstruction colored by local resolution estimation from ResMap. e, CD focused refinement reconstruction colored by local resolution estimation from ResMap. f, FSC plot of full channel (red), TMD focused refinement (blue) and CD focused refinement reconstructions (black). g, FSC plot of full channel reconstruction of all particles compared with the refined model (FSC sum, black), FSC plot of half-map1 compared with the refined model (FSC work, red) and FSC plot of half-map2 compared with the refined model (FSC free, blue) generated by phenix.mtriage.
Supplementary Figure 7 Ca2+-binding sites in Ca2+-bound hIP3R3.
a-c, Ca2+-binding sites in (a) full channel Ca2+-bound map, (b) CD focused refinement Ca2+-bound map and TMD focused refinement Ca2+-bound map. Coordinating side chains and IP3 are shown as sticks and colored by element. JD Ca2+ is shown as a green sphere and CD Ca2+ is shown as a magenta sphere. d, Superposition of the JD Ca2+ binding domain of Ca2+-bound hIP3R3 (colored by domain) with the Ca2+-binding site of rabbit type 1 ryanodine receptor (grey, PDB: 5T15). The structures are aligned by their Ca2+-coordinating residues. Coordinating residues are shown as sticks and Ca2+ ions are shown as green spheres. Top numbers correspond to hIP3R3 numbering and bottom residues correspond to rabbit type 1 ryanodine receptor numbering. e, Empty Ca2+-binding sites in apo full channel reconstruction.
Supplementary Figure 8 Cryo-EM analysis of low IP3–Ca2+ hIP3R3.
a, Representative raw image and 2D averages of low IP3-Ca2+ hIP3R3. b, Particle selection and focused refinement procedures used to obtain full-channel and focused refinement structures of low IP3-Ca2+ hIP3R3. c, Full-channel reconstruction colored by local resolution estimation from ResMap. d, CD focused refinement reconstruction colored by local resolution estimation from ResMap. e, FSC plot of full channel (red), TMD focused refinement (blue) and CD focused refinement reconstructions (black). f, FSC plot of full channel reconstruction of all particles compared with the refined model (FSC sum, black), FSC plot of half-map1 compared with the refined model (FSC work, red) and FSC plot of half-map2 compared with the refined model (FSC free, blue) generated by phenix.mtriage.
Supplementary Figure 9 Cryo-EM analysis of high IP3–Ca2+ hIP3R3.
a, Representative raw image and 2D averages of high IP3-Ca2+ hIP3R3. b, Particle selection and focused refinement procedures used to obtain full-channel and focused refinement structures of high IP3-Ca2+ hIP3R3. c, Full-channel reconstruction colored by local resolution estimation from ResMap. d, CD focused refinement reconstruction colored by local resolution estimation from ResMap. e, FSC plot of full channel (red), TMD focused refinement (blue) and CD focused refinement reconstructions (black). f, FSC plot of full channel reconstruction of all particles compared with the refined model (FSC sum, black), FSC plot of half-map1 compared with the refined model (FSC work, red) and FSC plot of half-map2 compared with the refined model (FSC free, blue) generated by phenix.mtriage.
Supplementary Figure 10 Features of the high IP3–Ca2+ and low IP3–Ca2+ structures.
Ca2+ and IP3-binding sites in (a) full channel high IP3-Ca2+, (b) TMD focused refinement high IP3-Ca2+, (c) CD focused refinement high IP3-Ca2+, (d) full channel low IP3-Ca2+, (e) TMD focused refinement low IP3-Ca2+ and (f) CD focused refinement low IP3-Ca2+ density maps. Coordinating side chains and IP3 are shown as sticks and colored by element. JD Ca2+ is shown as a green sphere and CD Ca2+ is shown as a magenta sphere.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Note and Supplementary Table 1
Supplementary Video 1
Conformational diversity of IP3-bound states in hIP3R3
Supplementary Video 2
Conformational changes between apo state, IP3 class 1 and IP3 class 2
Supplementary Video 3
Conformational changes between apo state, Ca2+-bound state and high IP3-Ca2+ state
Rights and permissions
About this article
Cite this article
Paknejad, N., Hite, R.K. Structural basis for the regulation of inositol trisphosphate receptors by Ca2+ and IP3. Nat Struct Mol Biol 25, 660–668 (2018). https://doi.org/10.1038/s41594-018-0089-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0089-6
This article is cited by
-
Structural titration reveals Ca2+-dependent conformational landscape of the IP3 receptor
Nature Communications (2023)
-
Structure and transport mechanism of the human calcium pump SPCA1
Cell Research (2023)
-
RCN2 promotes Nasopharyngeal carcinoma progression by curbing Calcium flow and Mitochondrial apoptosis
Cellular Oncology (2023)
-
Structural basis for activation and gating of IP3 receptors
Nature Communications (2022)
-
Conformational motions and ligand-binding underlying gating and regulation in IP3R channel
Nature Communications (2022)