Abstract
The C5a receptor (C5aR) is a G-protein-coupled receptor (GPCR) that can induce strong inflammatory response to the anaphylatoxin C5a. Targeting C5aR has emerged as a novel anti-inflammatory therapeutic method. However, developing potent C5aR antagonists as drugs has proven difficult. Here, we report two crystal structures of human C5aR in ternary complexes with the peptide antagonist PMX53 and a non-peptide antagonist, either avacopan or NDT9513727. The structures, together with other biophysical, computational docking and cell-based signaling data, reveal the orthosteric action of PMX53 and its effect of stabilizing the C5aR structure, as well as the allosteric action of chemically diverse non-peptide C5aR antagonists with different binding poses. Structural comparison analysis suggests the presence of similar allosteric sites in other GPCRs. We also discuss critical structural features of C5aR in activation, including a novel conformation of helix 8. On the basis of our results, we suggest novel strategies for developing C5aR-targeting drugs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Klos, A. et al. The role of the anaphylatoxins in health and disease. Mol. Immunol. 46, 2753–2766 (2009).
Klos, A., Wende, E., Wareham, K. J. & Monk, P. N. International union of basic and clinical pharmacology. [corrected]. LXXXVII. Complement peptide C5a, C4a, and C3a receptors. Pharmacol. Rev. 65, 500–543 (2013).
Bjarnadóttir, T. K. et al. Comprehensive repertoire and phylogenetic analysis of the G protein-coupled receptors in human and mouse. Genomics 88, 263–273 (2006).
Guo, R. F. & Ward, P. A. Role of C5a in inflammatory responses. Annu. Rev. Immunol. 23, 821–852 (2005).
Fonseca, M. I. et al. Contribution of complement activation pathways to neuropathology differs among mouse models of Alzheimer’s disease. J. Neuroinflammation 8, 4 (2011).
Landlinger, C. et al. Active immunization against complement factor C5a: a new therapeutic approach for Alzheimer’s disease. J. Neuroinflammation 12, 150 (2015).
Quadros, A. U. & Cunha, T. M. C5a and pain development: An old molecule, a new target. Pharmacol. Res. 112, 58–67 (2016).
Pandey, M. K. et al. Complement drives glucosylceramide accumulation and tissue inflammation in Gaucher disease. Nature 543, 108–112 (2017).
Markiewski, M. M. et al. Modulation of the antitumor immune response by complement. Nat. Immunol. 9, 1225–1235 (2008).
Nitta, H. et al. Enhancement of human cancer cell motility and invasiveness by anaphylatoxin C5a via aberrantly expressed C5a receptor (CD88). Clin. Cancer Res. 19, 2004–2013 (2013).
Vadrevu, S. K. et al. Complement c5a receptor facilitates cancer metastasis by altering T-cell responses in the metastatic niche. Cancer Res. 74, 3454–3465 (2014).
Darling, V. R., Hauke, R. J., Tarantolo, S. & Agrawal, D. K. Immunological effects and therapeutic role of C5a in cancer. Expert Rev. Clin. Immunol. 11, 255–263 (2015).
Woodruff, T. M., Nandakumar, K. S. & Tedesco, F. Inhibiting the C5-C5a receptor axis. Mol. Immunol. 48, 1631–1642 (2011).
Finch, A. M. et al. Low-molecular-weight peptidic and cyclic antagonists of the receptor for the complement factor C5a. J. Med. Chem. 42, 1965–1974 (1999).
March, D. R. et al. Potent cyclic antagonists of the complement C5a receptor on human polymorphonuclear leukocytes. Relationships between structures and activity. Mol. Pharmacol. 65, 868–879 (2004).
Monk, P. N., Scola, A. M., Madala, P. & Fairlie, D. P. Function, structure and therapeutic potential of complement C5a receptors. Br. J. Pharmacol. 152, 429–448 (2007).
Sumichika, H. et al. Identification of a potent and orally active non-peptide C5a receptor antagonist. J. Biol. Chem. 277, 49403–49407 (2002).
Brodbeck, R. M. et al. Identification and characterization of NDT 9513727 [N,N-bis(1,3-benzodioxol-5-ylmethyl)-1-butyl-2,4-diphenyl-1H-imidazole-5-methanamine], a novel, orally bioavailable C5a receptor inverse agonist. J. Pharmacol. Exp. Ther. 327, 898–909 (2008).
Bekker, P. et al. Characterization of pharmacologic and pharmacokinetic properties of CCX168, a potent and selective orally administered complement 5a receptor inhibitor, based on preclinical evaluation and randomized phase 1 clinical study. PLoS One 11, e0164646 (2016).
Seow, V. et al. Receptor residence time trumps drug-likeness and oral bioavailability in determining efficacy of complement C5a antagonists. Sci. Rep. 6, 24575 (2016).
Jayne, D. R. W. et al. Randomized trial of C5a receptor inhibitor avacopan in ANCA-associated vasculitis. J. Am. Soc. Nephrol. 28, 2756–2767 (2017).
Robertson, N. et al. Structure of the complement C5a receptor bound to the extra-helical antagonist NDT9513727. Nature 553, 111–114 (2018).
Lu, J. et al. Structural basis for the cooperative allosteric activation of the free fatty acid receptor GPR40. Nat. Struct. Mol. Biol. 24, 570–577 (2017).
Chun, E. et al. Fusion partner toolchest for the stabilization and crystallization of G protein-coupled receptors. Structure 20, 967–976 (2012).
Finch, A. M. et al. Biologically active conformer of the effector region of human C5a and modulatory effects of N-terminal receptor binding determinants on activity. J. Med. Chem. 40, 877–884 (1997).
Siciliano, S. J. et al. Two-site binding of C5a by its receptor: an alternative binding paradigm for G protein-coupled receptors. Proc. Natl. Acad. Sci. USA 91, 1214–1218 (1994).
Klco, J. M., Lassere, T. B. & Baranski, T. J. C5a receptor oligomerization. I. Disulfide trapping reveals oligomers and potential contact surfaces in a G protein-coupled receptor. J. Biol. Chem. 278, 35345–35353 (2003).
White, J. F. et al. Structure of the agonist-bound neurotensin receptor. Nature 490, 508–513 (2012).
Qin, L. et al. Structural biology. Crystal structure of the chemokine receptor CXCR4 in complex with a viral chemokine. Science 347, 1117–1122 (2015).
Shihoya, W. et al. Activation mechanism of endothelin ETB receptor by endothelin-1. Nature 537, 363–368 (2016).
Ma, Y. et al. Structural basis for apelin control of the human apelin receptor. Structure 25, 858–866.e854 (2017).
Cain, S. A., Coughlan, T. & Monk, P. N. Mapping the ligand-binding site on the C5a receptor: arginine74 of C5a contacts aspartate282 of the C5a receptor. Biochemistry 40, 14047–14052 (2001).
Baranski, T. J. et al. C5a receptor activation. Genetic identification of critical residues in four transmembrane helices. J. Biol. Chem. 274, 15757–15765 (1999).
Higginbottom, A. et al. Comparative agonist/antagonist responses in mutant human C5a receptors define the ligand binding site. J. Biol. Chem. 280, 17831–17840 (2005).
Nikiforovich, G. V., Marshall, G. R. & Baranski, T. J. Modeling molecular mechanisms of binding of the anaphylatoxin C5a to the C5a receptor. Biochemistry 47, 3117–3130 (2008).
Qu, H., Ricklin, D. & Lambris, J. D. Recent developments in low molecular weight complement inhibitors. Mol. Immunol. 47, 185–195 (2009).
Waters, S. M. et al. Molecular characterization of the gerbil C5a receptor and identification of a transmembrane domain V amino acid that is crucial for small molecule antagonist interaction. J. Biol. Chem. 280, 40617–40623 (2005).
Zhang, D. et al. Two disparate ligand-binding sites in the human P2Y1 receptor. Nature 520, 317–321 (2015).
Gerber, B. O., Meng, E. C., Dotsch, V., Baranski, T. J. & Bourne, H. R. An activation switch in the ligand binding pocket of the C5a receptor. J. Biol. Chem. 276, 3394–3400 (2001).
Wu, B. et al. Structures of the CXCR4 chemokine GPCR with small-molecule and cyclic peptide antagonists. Science 330, 1066–1071 (2010).
Konteatis, Z. D. et al. Development of C5a receptor antagonists. Differential loss of functional responses. J. Immunol. 153, 4200–4205 (1994).
Jia, N. et al. Pivotal advance: Interconversion between pure chemotactic ligands and chemoattractant/secretagogue ligands of neutrophil C5a receptor by a single amino acid substitution. J. Leukoc. Biol. 87, 965–975 (2010).
Rasmussen, S. G. et al. Structure of a nanobody-stabilized active state of the β(2) adrenoceptor. Nature 469, 175–180 (2011).
Huang, W. et al. Structural insights into µ-opioid receptor activation. Nature 524, 315–321 (2015).
Whistler, J. L. et al. Constitutive activation and endocytosis of the complement factor 5a receptor: evidence for multiple activated conformations of a G protein-coupled receptor. Traffic 3, 866–877 (2002).
Angel, T. E., Chance, M. R. & Palczewski, K. Conserved waters mediate structural and functional activation of family A (rhodopsin-like) G protein-coupled receptors. Proc. Natl. Acad. Sci. USA 106, 8555–8560 (2009).
Fenalti, G. et al. Molecular control of δ-opioid receptor signalling. Nature 506, 191–196 (2014).
Liu, W. et al. Structural basis for allosteric regulation of GPCRs by sodium ions. Science 337, 232–236 (2012).
Zhang, C. et al. High-resolution crystal structure of human protease-activated receptor 1. Nature 492, 387–392 (2012).
Li, S., Lee, S. Y. & Chung, K. Y. Conformational analysis of g protein-coupled receptor signaling by hydrogen/deuterium exchange mass spectrometry. Methods Enzymol 557, 261–278 (2015).
Konermann, L., Tong, X. & Pan, Y. Protein structure and dynamics studied by mass spectrometry: H/D exchange, hydroxyl radical labeling, and related approaches. J. Mass Spectrom. 43, 1021–1036 (2008).
Suvorova, E. S., Gripentrog, J. M., Oppermann, M. & Miettinen, H. M. Role of the carboxyl terminal di-leucine in phosphorylation and internalization of C5a receptor. Biochim. Biophys. Acta 1783, 1261–1270 (2008).
Rasmussen, S. G. et al. Crystal structure of the β2 adrenergic receptor-Gs protein complex. Nature 477, 549–555 (2011).
Zhou, X. E. et al. Identification of phosphorylation codes for arrestin recruitment by G protein-coupled receptors. Cell 170, 457–469.e413 (2017).
Gripentrog, J. M. & Miettinen, H. M. Formyl peptide receptor-mediated ERK1/2 activation occurs through G(i) and is not dependent on beta-arrestin1/2. Cell. Signal. 20, 424–431 (2008).
Moriconi, A. et al. Targeting the minor pocket of C5aR for the rational design of an oral allosteric inhibitor for inflammatory and neuropathic pain relief. Proc. Natl. Acad. Sci. USA 111, 16937–16942 (2014).
Zeng, Q. et al. Discovery of an irreversible HCV NS5B polymerase inhibitor. Bioorg. Med. Chem. Lett. 23, 6585–6587 (2013).
Li, R., Coulthard, L. G., Wu, M. C., Taylor, S. M. & Woodruff, T. M. C5L2: a controversial receptor of complement anaphylatoxin, C5a. FASEB J. 27, 855–864 (2013).
Van Lith, L. H., Oosterom, J., Van Elsas, A. & Zaman, G. J. C5a-stimulated recruitment of beta-arrestin2 to the nonsignaling 7-transmembrane decoy receptor C5L2. J. Biomol. Screen. 14, 1067–1075 (2009).
Zhang, H. et al. Structural basis for selectivity and diversity in angiotensin II receptors. Nature 544, 327–332 (2017).
Caffrey, M. Crystallizing membrane proteins for structure determination: use of lipidic mesophases. Annu. Rev. Biophys. 38, 29–51 (2009).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Williams, C. J. et al. MolProbity: More and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Mysinger, M. M. & Shoichet, B. K. Rapid context-dependent ligand desolvation in molecular docking. J. Chem. Inf. Model. 50, 1561–1573 (2010).
Duc, N. M. et al. Effective application of bicelles for conformational analysis of G protein-coupled receptors by hydrogen/deuterium exchange mass spectrometry. J. Am. Soc. Mass Spectrom. 26, 808–817 (2015).
Acknowledgements
We thank the staff at the GM/CA @ APS of Argonne National Laboratory at Chicago for their assistance with X-ray diffraction data collection. We acknowledge support from the University of Pittsburgh by the UPMC Competitive Medical Research Fund (CMRF) (C.Z.), the National Research Foundation of Korea funded by the Korean government (NFR-2015R1A1A1A05027473) (K.Y.C.), the funding support from the Biomedical Research Council, A*STAR (R.N.V.K.D. and H.F.) and the funding support from National Natural Science Foundation of China (31770791 and 315707410) (Z.W.). Z.W. is also supported by the Recruitment Program of Global Youth Experts of China.
Author information
Authors and Affiliations
Contributions
H.L. and C.Z. designed the research. H.L. performed all of the experiments for protein expression, purification, crystallization and functional characterization. H.R.K. and K.Y.C. performed HDX-MS experiments and data analysis. R.N.V.K.D. and H.F. performed computational docking studies. L.W. assisted with X-ray diffraction data collection. Z.W. determined the crystal structures. C.Z. processed the X-ray diffraction data and supervised the protein expression, purification, crystallization and functional studies. Z.W., K.Y.C. and C.Z. prepared the manuscript with the assistance from H.F.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated Supplementary Information
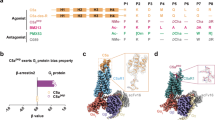
Supplementary Figure 1 Chemically diverse non-peptide C5aR antagonists and C5aR crystallization.
(a) Chemical structures of four C5aR non-peptide antagonists: avacopan (CCX168), NDT9513727, W54011 and CP-447697. The central tertiary nitrogen group is circled. (b) To aid the crystallization, we engineered human wtC5aR by replacing the N-terminal residues 1-29 with cytochrome b262 RIL (BRIL). We also introduced a TEV protease site between 331 and 332 and cut off the C-terminal segment 332-350 before crystallization by TEV protease. The construct is referred as C5aR-BRIL. Crystals of C5aR with PMX53 and avacopan in lipidic cubic phase are shown in the upper right corner. (c) The Fo-Fc omit electron-density maps for PMX53 (green), NDT9513727 (cyan) and avacopan (orange) contoured at 2σ are shown as purple mesh.
Supplementary Figure 2 Structural comparison with thermostable C5aR, crystal packing and ligand binding at the intracellular surface.
(a) Left: Superimposition of the C5aR-PMX53-NDT9513727 structure with the structure of thermostable C5aR-NDT95132727. Right: The surfaces of these two structures viewed from the extracellular side. The large differences are observed for ECL2 and the extracellular end of TM4. The unique conformation of helix 8 is observed in both structures. (b) Top: Lattice packing in C5aR-PMX53-avacopan crystals. Receptor is shown in blue and BRIL is shown in purple. PMX53 and avacopan are shown in green and orange respectively. In any two C5aR molecules that form crystal-packing interactions, the allosteric sites are far away from each other with a distance of ~27 Å (measured between two W255 residues in the sites). Bottom: In each layer of the crystal lattice, one receptor packs against the surrounding receptor molecules in an anti-parallel manner, excluding the possibility of forming physiological C5aR dimers in the crystals. (c) The malonate molecule (yellow) bound at the cytoplasmic surface of C5aR-PMX53-avacopan (blue). An Fo-Fc omit electron-density map for the malonate contoured at 2σ is shown as purple mesh. Residues N712.39, R1343.50 and T2406.33 that form hydrogen-bonding interactions with the malonate are shown as sticks.
Supplementary Figure 3 Comparison of C5aR structure to the structures of other GPCRs with peptide ligands.
Crystal structures of GPCRs bound to the peptide ligands: C5aR with PMX53; CXCR4 with CVX15 (PDB ID 3OE0); CXCR4 with mVIP-II (PDB ID 4RWS); neurotensin receptor 1 (NTSR) with neurotensin (8-13) (PDB ID 4GRV); endothelin receptor type-B (ETBR) with endothelin-1 (PDB ID 5GLH); apelin receptor (Apelin R) with an apelin mimetic peptide (PDB ID 5VBL). C5aR is colored in blue and other GPCRs are colored in grey. PMX53 is colored in green and other peptide ligands are colored in red. PMX53 interacts with ECL2 but not N-terminus of C5aR, while other peptide ligands all interact with the N-termini and ECL2s of their receptors.
Supplementary Figure 4 Allosteric site for NDT9513727.
(a) Allosteric binding site for NDT9513727 (cyan) in our structure. All the residues that interact with NDT9513727 are in the same conformation as observed in the structure of thermostable C5aR. However, the residue A1564.45 was mutated to leucine in the thermostable C5aR, which may provide additional hydrophobic interactions for the binding of NDT9513727. (b) The effects of 6 mutations V159A, V159H, L163W, L209W, W213L and W213G on the antagonistic action of NDT9513727 measured by an agonist YR10-induced Ca2 + release assay. The cellular calcium level was determined by measuring the fluorescence of fluo-4 in HEK293T cells stably expressing wtC5aR. 10 μM YR10 was used to stimulate the receptor. Data points are presented as the mean values + /- SEM, n = 3.
Supplementary Figure 5 Conservation of the allosteric site in other GPCRs.
Crystal structures of GPCRs with small compounds bound to the similar site as identified in C5aR and GPR40 for the allosteric modulators (PDB IDs shown in the brackets): (a) C5aR with avacopan; (b) δ-opioid receptor with 1-oleoyl-R-glycerol (DOR, 4N6H); (c) μ-opioid receptor with a PEG400 molecule (MOR, 5C1M); (d) N/OFQ opioid receptor with 1-oleoyl-R-glycerol (NOR, 4EA3); (e) viral GPCR US28 with a succinic acid molecule (US28, 4XT1); (f) A2a adenosine receptor with an oleic acid molecule (A2AR, 5K2D); (g) chemokine receptor CCR9 with an oleic acid molecule (CCR9, 5LWE); (i) cannabinoid receptor CB1 with an oleic acid molecule (CB1, 5TGZ). The cysteine residue at 5.57 position C5.57 is conserved in all of these GPCRs except A2AR and CB1.
Supplementary Figure 6 Different sodium coordination sites in four GPCRs.
The sodium coordination sites in four GPCRs: C5aR, protease-activated receptor 1(PAR1, PDB ID 3VW7), A2A adenosine receptor (A2AR, PDB ID 4EIY) and δ-opioid receptor (δOR, PDB ID 4N6H) are shown. The sodium ion (shown as a purple sphere) interacts with different sets of residues at the cytoplasmic side of these GPCRs and water molecules (shown as red spheres).
Supplementary Figure 7 HDX-MS sequence coverage map and deuterium uptake plots.
(a) Bars indicated the peptic peptides in different regions of C5aR for HDX analysis. (b) Deuterium uptake plots of those regions in C5aR are represented. The results of three samples, C5aR bound to NDT9513727 (NDT), C5aR bound to PMX53 (PMX53) and C5aR bound to both ligands (NDT + PMX53), are compared. There is no significant difference between the results of PMX53 and NDT + PMX53. Results are means ± S.E. for at least three independent experiments. *p < 0.05: NDT-bound state vs. PMX53-bound state, #p < 0.05: NDT-bound state vs. NDT and PMX53-bound state
Supplementary Figure 8 Unique helix 8 in C5aR.
(a) Hydrophobic interactions that stabilize the conformation of helix 8. In both C5aR structures, residues L315, L319, V322 and L323 on helix 8 form extensive hydrophobic interactions with the surrounding residues L571.52, V581.53, V611.56, F752.43, V2436.36, Y3007.53 and V3017.54 from TM1, TM2, TM6 and TM7, respectively. (b) The conformation of helix 8 in four GPCRs: C5aR, angiotensin II receptor 2 (AT2R, PDB ID 5UNF), δ-opioid receptor (δOR, PDB ID 4N6H) and protease-activated receptor 1(PAR1, PDB ID 3VW7). In nearly all solved structures of rhodopsin-like GPCRs, either there is no well-folded helix 8 as seen in PAR1 or the helix 8 adopts a similar conformation as that in δOR with the C-terminus extending away from the helical bundle. In AT2R, the helix 8 adopts a unique conformation to interact with the cytoplasmic surface, which can sterically block the binding of signaling effectors and result in the lacking of signaling responses for AT2R. In C5aR, the helix 8 is in a reversed orientation compared to that in δOR and inserts between TM1 and TM7. Such conformation has not been observed in other GPCRs so far.
Supplementary Figure 9 Extracellular sites for potential non-peptide C5aR antagonist.
(a) Alignment of C5aR structure with the structures of two angiotensin II receptors AT1R (pink, PDB ID 4ZUD) and AT2R (light pink, PDB ID 5UNF). PMX53 is colored in green and the antagonists of AT1R and AT2R are colored in red. The binding sites for angiotensin antagonists overlap with the bottom region of the PMX53 binding site, suggesting the feasibility of developing non-peptide C5aR antagonists that target this region. (b) Extracellular allosteric antagonist DF2593A. Two residues, D2827.35 and N1193.35, were proposed to form direct polar interactions with DF2593A. However, there is no enough space in our structures around these two residues to accommodate DF2593A, even with the absence of PMX53 (green sticks). It is likely that DF2593A binding is associated with large conformational changes of the extracellular region, resulting in a distinct binding pocket compared to PMX53.
Supplementary Figure 10 Sequence alignment of C5aR and C5L2.
The purple arrows point to the residues in C5aR that are involved in the interactions with PMX53, including F44, L92, I96, P113, I116, R178, E199, R206, Y178, T261, D282 and V286. Except for F44, R178 and T261, all other residues are conserved in C5L2. However, the substitution of G189 in C5aR, as indicated by the blue star, by V187 in C5L2 may introduce steric clash with PMX53. The right panel shows the position of G189 relative to PMX53 (green sticks) in the structure of C5aR (blue). The distance between G189 and the Pro residue in PMX53 is too short to tolerate any substitution of G189. The orange star indicates the residue W213 in C5aR, which is critical for the binding of non-peptide antagonists. It is substituted by G211 in C5L2.
Supplementary information
Rights and permissions
About this article
Cite this article
Liu, H., Kim, H.R., Deepak, R.N.V.K. et al. Orthosteric and allosteric action of the C5a receptor antagonists. Nat Struct Mol Biol 25, 472–481 (2018). https://doi.org/10.1038/s41594-018-0067-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0067-z
This article is cited by
-
G protein-coupled receptors (GPCRs): advances in structures, mechanisms, and drug discovery
Signal Transduction and Targeted Therapy (2024)
-
Structural insights into the activation and inhibition of CXC chemokine receptor 3
Nature Structural & Molecular Biology (2024)
-
Recent advances in targeting the “undruggable” proteins: from drug discovery to clinical trials
Signal Transduction and Targeted Therapy (2023)
-
Mechanism of activation and biased signaling in complement receptor C5aR1
Cell Research (2023)
-
Revealing the signaling of complement receptors C3aR and C5aR1 by anaphylatoxins
Nature Chemical Biology (2023)