Abstract
mRNAs transmit the genetic information that dictates protein production and are a nexus for numerous pathways that regulate gene expression. The prevailing view of canonical mRNA decay is that it is mediated by deadenylation and decapping followed by exonucleolysis from the 3′ and 5′ ends. By developing Akron-seq, a novel approach that captures the native 3′ and 5′ ends of capped and polyadenylated RNAs, respectively, we show that canonical human mRNAs are subject to repeated cotranslational and ribosome-phased endonucleolytic cuts at the exit site of the mRNA ribosome channel, in a process that we term ribothrypsis. We uncovered RNA G quadruplexes among likely ribothrypsis triggers and show that ribothrypsis is a conserved process. Strikingly, we found that mRNA fragments are abundant in living cells and thus have important implications for the interpretation of experiments, such as RNA-seq, that rely on the assumption that mRNAs exist largely as full-length molecules in vivo.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Dreyfuss, G., Kim, V. N. & Kataoka, N. Messenger-RNA-binding proteins and the messages they carry. Nat. Rev. Mol. Cell Biol. 3, 195–205 (2002).
Singh, G., Pratt, G., Yeo, G. W. & Moore, M. J. The clothes make the mRNA: past and present trends in mRNP fashion. Annu. Rev. Biochem. 84, 325–354 (2015).
Green, R. & Noller, H. F. Ribosomes and translation. Annu. Rev. Biochem. 66, 679–716 (1997).
Schoenberg, D. R. & Maquat, L. E. Regulation of cytoplasmic mRNA decay. Nat. Rev. Genet. 13, 246–259 (2012).
Houseley, J. & Tollervey, D. The many pathways of RNA degradation. Cell 136, 763–776 (2009).
Lykke-Andersen, S., Tomecki, R., Jensen, T. H. & Dziembowski, A. The eukaryotic RNA exosome: same scaffold but variable catalytic subunits. RNA Biol. 8, 61–66 (2011).
Coller, J. & Parker, R. Eukaryotic mRNA decapping. Annu. Rev. Biochem. 73, 861–890 (2004).
Grudzien-Nogalska, E. & Kiledjian, M. New insights into decapping enzymes and selective mRNA decay. Wiley Interdiscip. Rev. RNA 8, e1379 (2017).
Jens, M. & Rajewsky, N. Competition between target sites of regulators shapes post-transcriptional gene regulation. Nat. Rev. Genet. 16, 113–126 (2015).
Vlachos, I. S. & Hatzigeorgiou, A. G. Functional analysis of miRNAs using the DIANA tools online suite. Methods Mol. Biol. 1517, 25–50 (2017).
Richter, J. D. & Coller, J. Pausing on polyribosomes: make way for elongation in translational control. Cell 163, 292–300 (2015).
Hu, W., Sweet, T. J., Chamnongpol, S., Baker, K. E. & Coller, J. Co-translational mRNA decay in Saccharomyces cerevisiae. Nature 461, 225–229 (2009).
Pelechano, V., Wei, W. & Steinmetz, L. M. Widespread co-translational RNA decay reveals ribosome dynamics. Cell 161, 1400–1412 (2015).
Yu, X., Willmann, M. R., Anderson, S. J. & Gregory, B. D. Genome-wide mapping of uncapped and cleaved transcripts reveals a role for the nuclear mRNA cap-binding complex in cotranslational RNA decay in Arabidopsis. Plant Cell 28, 2385–2397 (2016).
Isken, O. & Maquat, L. E. Quality control of eukaryotic mRNA: safeguarding cells from abnormal mRNA function. Genes Dev. 21, 1833–1856 (2007).
Shoemaker, C. J. & Green, R. Translation drives mRNA quality control. Nat. Struct. Mol. Biol. 19, 594–601 (2012).
Brandman, O. & Hegde, R. S. Ribosome-associated protein quality control. Nat. Struct. Mol. Biol. 23, 7–15 (2016).
Inada, T. The ribosome as a platform for mRNA and nascent polypeptide quality control. Trends Biochem. Sci. 42, 5–15 (2017).
Mendell, J. T., Sharifi, N. A., Meyers, J. L., Martinez-Murillo, F. & Dietz, H. C. Nonsense surveillance regulates expression of diverse classes of mammalian transcripts and mutes genomic noise. Nat. Genet. 36, 1073–1078 (2004).
Doma, M. K. & Parker, R. Endonucleolytic cleavage of eukaryotic mRNAs with stalls in translation elongation. Nature 440, 561–564 (2006).
Frischmeyer, P. A. et al. An mRNA surveillance mechanism that eliminates transcripts lacking termination codons. Science 295, 2258–2261 (2002).
Tsuboi, T. et al. Dom34:hbs1 plays a general role in quality-control systems by dissociation of a stalled ribosome at the 3′ end of aberrant mRNA. Mol. Cell 46, 518–529 (2012).
Ikeuchi, K. & Inada, T. Ribosome-associated Asc1/RACK1 is required for endonucleolytic cleavage induced by stalled ribosome at the 3′ end of nonstop mRNA. Sci. Rep. 6, 28234 (2016).
Guydosh, N. R. & Green, R. Translation of poly(A) tails leads to precise mRNA cleavage. RNA 23, 749–761 (2017).
Shoemaker, C. J., Eyler, D. E. & Green, R. Dom34:Hbs1 promotes subunit dissociation and peptidyl-tRNA drop-off to initiate no-go decay. Science 330, 369–372 (2010).
Shoemaker, C. J. & Green, R. Kinetic analysis reveals the ordered coupling of translation termination and ribosome recycling in yeast. Proc. Natl. Acad. Sci. USA 108, E1392–E1398 (2011).
Pisareva, V. P., Skabkin, M. A., Hellen, C. U., Pestova, T. V. & Pisarev, A. V. Dissociation by Pelota, Hbs1 and ABCE1 of mammalian vacant 80S ribosomes and stalled elongation complexes. EMBO J. 30, 1804–1817 (2011).
Becker, T. et al. Structure of the no-go mRNA decay complex Dom34-Hbs1 bound to a stalled 80S ribosome. Nat. Struct. Mol. Biol. 18, 715–720 (2011).
Bengtson, M. H. & Joazeiro, C. A. Role of a ribosome-associated E3 ubiquitin ligase in protein quality control. Nature 467, 470–473 (2010).
Brandman, O. et al. A ribosome-bound quality control complex triggers degradation of nascent peptides and signals translation stress. Cell 151, 1042–1054 (2012).
Defenouillère, Q. et al. Cdc48-associated complex bound to 60S particles is required for the clearance of aberrant translation products. Proc. Natl. Acad. Sci. USA 110, 5046–5051 (2013).
Verma, R., Oania, R. S., Kolawa, N. J. & Deshaies, R. J. Cdc48/p97 promotes degradation of aberrant nascent polypeptides bound to the ribosome. eLife 2, e00308 (2013).
German, M. A. et al. Global identification of microRNA-target RNA pairs by parallel analysis of RNA ends. Nat. Biotechnol. 26, 941–946 (2008).
Schmidt, S. A. et al. Identification of SMG6 cleavage sites and a preferred RNA cleavage motif by global analysis of endogenous NMD targets in human cells. Nucleic Acids Res. 43, 309–323 (2015).
Addo-Quaye, C., Eshoo, T. W., Bartel, D. P. & Axtell, M. J. Endogenous siRNA and miRNA targets identified by sequencing of the Arabidopsis degradome. Curr. Biol. 18, 758–762 (2008).
Gregory, B. D. et al. A link between RNA metabolism and silencing affecting Arabidopsis development. Dev. Cell 14, 854–866 (2008).
Chang, H., Lim, J., Ha, M. & Kim, V. N. TAIL-seq: genome-wide determination of poly(A) tail length and 3′ end modifications. Mol. Cell 53, 1044–1052 (2014).
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
Park, J. E., Yi, H., Kim, Y., Chang, H. & Kim, V. N. Regulation of poly(A) tail and translation during the somatic cell cycle. Mol. Cell 62, 462–471 (2016).
Carlevaro-Fita, J., Rahim, A., Guigó, R., Vardy, L. A. & Johnson, R. Cytoplasmic long noncoding RNAs are frequently bound to and degraded at ribosomes in human cells. RNA 22, 867–882 (2016).
Zhuang, F., Fuchs, R. T., Sun, Z., Zheng, Y. & Robb, G. B. Structural bias in T4 RNA ligase-mediated 3′-adapter ligation. Nucleic Acids Res. 40, e54 (2012).
Guydosh, N. R. & Green, R. Dom34 rescues ribosomes in 3′ untranslated regions. Cell 156, 950–962 (2014).
Lubas, M. et al. The human nuclear exosome targeting complex is loaded onto newly synthesized RNA to direct early ribonucleolysis. Cell Rep. 10, 178–192 (2015).
Schmidt, C. et al. The cryo-EM structure of a ribosome-Ski2-Ski3-Ski8 helicase complex. Science 354, 1431–1433 (2016).
Martinez, J. & Tuschl, T. RISC is a 5′ phosphomonoester-producing RNA endonuclease. Genes Dev. 18, 975–980 (2004).
Orban, T. I. & Izaurralde, E. Decay of mRNAs targeted by RISC requires XRN1, the Ski complex, and the exosome. RNA 11, 459–469 (2005).
Tani, H. et al. Genome-wide determination of RNA stability reveals hundreds of short-lived noncoding transcripts in mammals. Genome Res. 22, 947–956 (2012).
Ingolia, N. T., Lareau, L. F. & Weissman, J. S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802 (2011).
Spitale, R. C. et al. Structural imprints in vivo decode RNA regulatory mechanisms. Nature 519, 486–490 (2015).
Silverman, I. M., Berkowitz, N. D., Gosai, S. J. & Gregory, B. D. Genome-wide approaches for RNA structure probing. Adv. Exp. Med. Biol. 907, 29–59 (2016).
Rouskin, S., Zubradt, M., Washietl, S., Kellis, M. & Weissman, J. S. Genome-wide probing of RNA structure reveals active unfolding of mRNA structures in vivo. Nature 505, 701–705 (2014).
Song, J., Perreault, J. P., Topisirovic, I. & Richard, S. RNA G-quadruplexes and their potential regulatory roles in translation. Translation (Austin) 4, e1244031 (2016).
Kwok, C. K., Marsico, G., Sahakyan, A. B., Chambers, V. S. & Balasubramanian, S. rG4-seq reveals widespread formation of G-quadruplex structures in the human transcriptome. Nat. Methods 13, 841–844 (2016).
Guo, J. U. & Bartel, D. P. RNA G-quadruplexes are globally unfolded in eukaryotic cells and depleted in bacteria. Science 353, aaf5371 (2016).
Vourekas, A. et al. The RNA helicase MOV10L1 binds piRNA precursors to initiate piRNA processing. Genes Dev. 29, 617–629 (2015).
Endoh, T. & Sugimoto, N. Mechanical insights into ribosomal progression overcoming RNA G-quadruplex from periodical translation suppression in cells. Sci. Rep. 6, 22719 (2016).
Guydosh, N. R., Kimmig, P., Walter, P. & Green, R. Regulated Ire1-dependent mRNA decay requires no-go mRNA degradation to maintain endoplasmic reticulum homeostasis in S. pombe. eLife 6, e29216 (2017).
Simms, C. L., Yan, L. L. & Zaher, H. S. Ribosome collision is critical for quality control during no-go decay. Mol. Cell 68, 361–373 e5 (2017).
Simms, C. L., Hudson, B. H., Mosior, J. W., Rangwala, A. S. & Zaher, H. S. An active role for the ribosome in determining the fate of oxidized mRNA. Cell Rep. 9, 1256–1264 (2014).
Peach, S. E., York, K. & Hesselberth, J. R. Global analysis of RNA cleavage by 5′-hydroxyl RNA sequencing. Nucleic Acids Res. 43, e108 (2015).
Ibrahim, F. et al. Identification of in vivo, conserved, TAF15 RNA binding sites reveals the impact of TAF15 on the neuronal transcriptome. Cell Rep. 3, 301–308 (2013).
Subtelny, A. O., Eichhorn, S. W., Chen, G. R., Sive, H. & Bartel, D. P. Poly(A)-tail profiling reveals an embryonic switch in translational control. Nature 508, 66–71 (2014).
Maragkakis, M., Alexiou, P., Nakaya, T. & Mourelatos, Z. CLIPSeqTools: a novel bioinformatics CLIP-seq analysis suite. RNA 22, 1–9 (2016).
Acknowledgements
We thank the members of the laboratory of Z.M. for discussions; X. Ji and S. Liebhaber for assistance with ISCO polysome fractionation; and J. Schug (University of Pennsylvania) and D. Pouchnik (Washington State University) for technical help with Illumina and PacBio sequencing, respectively. We thank the University of Pennsylvania Diabetes Research Center (DRC) for the use of the Functional Genomics Core (P30-DK19525) for Illumina sequencing, and the Washington State University Molecular Biology and Genomics Core for the use of Pacific Biosciences sequencing. This work was supported by a Brody family fellowship to M.M. and by grants from the ALS Therapy Alliance and the NIH (GM072777) to Z.M.
Author information
Authors and Affiliations
Contributions
F.I., M.M. and Z.M. conceived the study. F.I. designed, performed and interpreted all wet-lab experiments and developed Akron-SMRT and Akron-seq. M.M. designed, performed and interpreted all computational analyses and ribothrypsis modeling, to which P.A. substantially contributed. All authors analyzed the data. F.I., M.M. and Z.M. wrote the manuscript, and P.A. provided input and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Akron-SMRT and high-resolution northern blots reveal cotranslational mRNA decay and persistence of mRNA fragments.
(a) Schematic of Akron-SMRT. Capped RNAs were enriched by treating RNA that was isolated after immediate cell lysis and protein denaturation with Trizol (total), or isolated from polysomal fractions after sucrose gradient sedimentation of cytoplasmic HeLa lysates (polysomal), with the Terminator 5’-phosphate (P)-dependent exonuclease. The 3’ ends of mRNA molecules were marked by ligating a biotinylated adapter; the ligated products were captured on streptavidin beads and gene-specific libraries were generated by reverse transcription-polymerase chain reaction (RT-PCR), and PCR amplification using forward primers (For) corresponding to the 5’ Untranslated Region (5’ UTR) and start codon (to ensure that the amplified products originated from mRNAs with intact 5’ ends) and reverse primer (Rev) corresponding to the 3’ adapter. Libraries were sequenced with PacBio Single-Molecule Real-Time (SMRT) technology and only circular consensus (CCS) reads were used for analyses. (b) Cumulative size distribution of Akron-SMRT CCS reads of capped cyclin-dependent kinase 4 (CDK4) and tubulin beta class I (TUBB) mRNAs from Terminator-treated, total and polysomal HeLa RNAs. CDK4 and TUBB mRNAs were randomly selected for Akron-SMRT. We found SMRT quantification quite inaccurate but nevertheless useful for qualitative investigation of mRNA decay from the 3’ end. (c) Cumulative size distribution of Akron-SMRT CCS reads of capped CDK4 and TUBB mRNAs from Terminator-treated, total and polysomal HeLa RNAs, divided depending whether they harbor untemplated adenines (poly(A)) or not (no-poly(A)). (d,e) Cotranslational degradation of capped mRNAs detected by high-resolution northern blots. Terminator-treated RNAs isolated from input (10%) and indicated fractions after sucrose gradient sedimentation of cytoplasmic HeLa lysate (d), or after anti-RPL7A (60S ribosomal protein L7a) immunoprecipitation of pooled polysomal fractions (e), were resolved by denaturing (UREA) polyacrylamide gel electrophoresis (PAGE); probed with radiolabeled probes straddling the 5' UTR and coding sequence (CDS) of indicated mRNAs; and visualized by storage-phosphor autoradiography. Brackets show prominent smears in RNA extracted from pooled polysome fractions, indicative of 3'-trimming of capped mRNAs, running faster than full length mRNAs, and corresponding, to ~73%, 75%, 78% and 82% of the total CDK4, TUBB, ACTB and RPL19 signal, respectively. Association of CDK4 mRNA fragments with ribosomes that were immunoprecipitated (IP) from polysomal fractions (fragments correspond to ~64% of total signal in input, and ~70% in polysome IP) confirms that the mRNA fragments associate with elongating ribosomes and not with cosedimenting "heavy" mRNPs (e). Non-immune rabbit serum (NRS) served as negative control; immunoprecipitates were also analyzed by Western blotting with anti-RPL7A. (f) Northern blot of capped CDK4 mRNA from Terminator-treated RNA from two independent total RNA isolations, probed with a radiolabeled probe straddling 5’ UTR and CDS of CDK4, reveals the presence of capped mRNA fragments in total RNA (fragments corresponding to ~80% of total signal). (g) Autoradiography of radiolabeled, in vitro synthesized, capped, Renilla luciferase mRNA, spiked in to input and polysome pool (fractions 6-14) prior to total RNA isolation and Terminator digestion, confirms absence of exogenous RNA degradation from experimental manipulations or from possible contaminating nuclease(s) in the Terminator enzyme preparation.
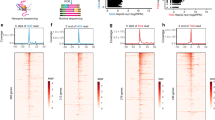
Supplementary Figure 2 Akron-seq.
(a) Capped RNAs represent the majority of HeLa RNAs remaining after Terminator treatment. Top: schematic of assay. After treating total RNA, which had been depleted of ribosomal RNAs (rRNAs) and small capped RNAs, with Terminator to degrade RNAs harboring 5’-P, the sample was divided into two equal aliquots. To detect 5’-OH RNAs, the first aliquot was 5’ end radiolabeled with T4 PolyNucleotide Kinase (PNK) and γ32P-ATP. To detect capped RNAs, the second aliquot was first treated with T4 PNK and cold ATP, to phosphorylate any RNAs bearing 5’-OH, followed by Terminator treatment to remove them. The sample was then treated by Tobacco Acid Pyrophosphatase (TAP) to remove the 5’ cap, leaving a residual 5’-P, which was removed with Calf Intestinal Phosphatase (CIP) treatment; the RNA was then 5’ end radiolabeled with T4 PNK and γ32P-ATP. Samples were resolved by UREA-PAGE and visualized by autoradiography (bottom). Capped RNAs are abundant but RNAs bearing 5’-OH are present in negligible amounts in total HeLa RNA. (b) Schematic of assay for Akron3 library generation from 5’-OH bearing RNAs; specific PCR product could not be generated due to low amounts of 5’-OH RNAs. (c) Genomic distribution of Akron5 and Akron3 ends for all libraries. UTR; untranslated region, CDS; coding sequence, tRNA; transfer RNA, rRNA; ribosomal RNA, rmsk; repeat masker, snRNA; small nuclear RNA. (d) Pearson’s correlation values for biological replicates of Akron5 and Akron3.
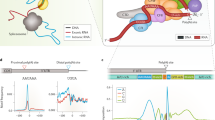
Supplementary Figure 3 Long noncoding RNAs are subjected to ribothrypsis; TAIL-seq reveals cotranslational generation of 3′ ends of mRNAs; and ribothrypsis simulation.
(a) TAIL-Seq 3’ end density and Discrete Fourier Transform (DFT) around RPFs 5’ ends with corresponding random control. (b) Density and DFT of Akron5 5’ ends of long noncoding RNAs (lncRNAs) around RPFs’ 5’ ends with corresponding random control. (c) Density of Akron5 and Akron3 ends around RPFs’ 5’ ends with corresponding control. Control corresponds to down-sampled Akron5 and Akron3 libraries such that the nucleotide (nt) content at the cut site is 25% for all nucleotides. (d) Density of simulated Akron5 5’ ends around simulated RPFs’ 5’ ends. We simulated 100 transcripts of uniformly random length between 200 and 5200 nt and uniformly random expression level between 1 and 1000 copies. In each transcript of length L, we introduced L/200 randomly located ribosome slowdown regions of 6-nt in length. We simulated actively translating ribosomes that move along the transcript with a 3-nt movement pattern. To simulate this pattern, we constrained ribosomes to move 30% faster in Open Reading Frames (ORFs) +2 and +3 than in +1. Ribosome speed over nucleotide is defined at their 3’ end and their RPF size is set at 29-nt. Speed in slowdown regions is reduced compared to background based on a normal distribution (mean: 0, standard deviation: 1) allowing for the maximum slowdown factor to be at maximum ~3 times slower than the background speed. The central 3- nt of the slowdown region is designated as an area 'protected' from ribothrypsis cuts. Ribosomes moving along a simulated transcript have 0.01 probability per unit time to produce an RPF at their current location. Also, ribosomes have 0.60 probability to create a ribothrypsis cut at their 5’ end per unit time. Each time a primary ribothrypsis cut is created, we define 0.01 probability of an incoming ribosome being captured approximately 15-nt upstream.
Supplementary Figure 4 5PSeq reanalysis indicates that ribothrypsis probably operates in S. cerevisiae.
(a–c) 5’ end density of RPFs (a), 5PSeq from wild-type (YPD) (b), and Xrn1 knockout (Xrn1Δ) (c) yeast strains, upstream and downstream of stop codon (last nucleotide of stop codon set as position 0) and DFT upstream and downstream of position -18 (marks the terminating ribosome 5’ end, shown with green dotted lines). (d,e) 5PSeq 5' end density and DFT upstream and downstream of RPF 5’ ends for YPD (d) and Xrn1Δ (e) with corresponding random controls.
Supplementary Figure 5 The exosome is not ribothrypsin; final ribothrypsis products (FRTPs) are evident in sRNA-seq libraries.
(a,b) End density and DFT around RPF 5’ ends for small RNA (sRNA) in RRP40 knockdown (KD) 5’ ends (a), and small RNA in combined DIS3, DIS3L, RRP6 KD 5’ ends (b). (c) Density of 5' and 3' ends of sRNA (<17-nt) around non-collapsed Akron5 ends. Dashed lines correspond to upper confidence intervals. A schematic of the analysis is shown at the top.
Supplementary Figure 6 Codon distribution around Akron5 ends.
Plots show the percentage of ends with marked codon at each position around Akron5 ends along with corresponding controls designed to maintain nucleotide content at the cut site. Codon and corresponding amino acid is marked at the top of each plot in the form amino acid:codon.
Supplementary Figure 7 Codon distribution around Akron3 ends.
Plots show the percentage of ends with marked codon at each position around Akron3 ends along with corresponding controls designed to maintain nucleotide content at the cut site. Codon and corresponding amino acid is marked at the top of each plot in the form amino acid:codon.
Supplementary Figure 8 rG4s are enriched downstream of yeast 5PSeq ends independently of Xrn1.
Centered coverage of predicted rG4s upstream and downstream of 5PSeq ends in wild-type (YPD, a) and Xrn1 knockout (Xrn1Δ, b) yeast strains along with random and shuffled controls.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 1 and 2
Rights and permissions
About this article
Cite this article
Ibrahim, F., Maragkakis, M., Alexiou, P. et al. Ribothrypsis, a novel process of canonical mRNA decay, mediates ribosome-phased mRNA endonucleolysis. Nat Struct Mol Biol 25, 302–310 (2018). https://doi.org/10.1038/s41594-018-0042-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0042-8
This article is cited by
-
Boric acid intercepts 80S ribosome migration from AUG-stop by stabilizing eRF1
Nature Chemical Biology (2024)
-
CD95/Fas ligand mRNA is toxic to cells through more than one mechanism
Molecular Biomedicine (2023)
-
Feature selection for RNA cleavage efficiency at specific sites using the LASSO regression model in Arabidopsis thaliana
BMC Bioinformatics (2021)
-
Translation and codon usage regulate Argonaute slicer activity to trigger small RNA biogenesis
Nature Communications (2021)
-
Ribosome-guided piRNA production
Nature Cell Biology (2020)