Abstract
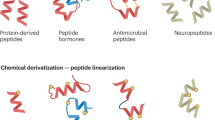
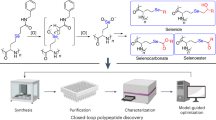
Peptides folded through interwoven disulfides display extreme biochemical properties and unique medicinal potential. However, their exploitation has been hampered by the limited amounts isolatable from natural sources and the expense of chemical synthesis. We developed reliable biological methods for high-throughput expression, screening and large-scale production of these peptides: 46 were successfully produced in multimilligram quantities, and >600 more were deemed expressible through stringent screening criteria. Many showed extreme resistance to temperature, proteolysis and/or reduction, and all displayed inhibitory activity against at least 1 of 20 ion channels tested, thus confirming their biological functionality. Crystal structures of 12 confirmed proper cystine topology and the utility of crystallography to study these molecules but also highlighted the need for rational classification. Previous categorization attempts have focused on limited subsets featuring distinct motifs. Here we present a global definition, classification and analysis of >700 structures of cystine-dense peptides, providing a unifying framework for these molecules.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Molesini, B., Treggiari, D., Dalbeni, A., Minuz, P. & Pandolfini, T. Plant cystine-knot peptides: pharmacological perspectives. Br. J. Clin. Pharmacol. 83, 63–70 (2017).
Herzig, V. & King, G. F. The cystine knot is responsible for the exceptional stability of the insecticidal spider toxin ω-hexatoxin-Hv1a. Toxins (Basel) 7, 4366–4380 (2015).
Reinwarth, M., Nasu, D., Kolmar, H. & Avrutina, O. Chemical synthesis, backbone cyclization and oxidative folding of cystine-knot peptides: promising scaffolds for applications in drug design. Molecules 17, 12533–12552 (2012).
Kolmar, H. Natural and engineered cystine knot miniproteins for diagnostic and therapeutic applications. Curr. Pharm. Des. 17, 4329–4336 (2011).
Postic, G., Gracy, J., Périn, C., Chiche, L. & Gelly, J. C. KNOTTIN: the database of inhibitor cystine knot scaffold after 10 years, toward a systematic structure modeling. Nucleic Acids Res. 46, D454–D458 (2018).
Kould, A., Ji, Y., Aboye, T. L. & Camarero, J. A. Cyclotides, a novel ultrastable polypeptide scaffold for drug discovery. Curr. Pharm. Des. 17, 4294–4307 (2011).
Schwarz, E. Cystine knot growth factors and their functionally versatile proregions. Biol. Chem. 398, 1295–1308 (2017).
Iyer, S. & Acharya, K. R. Tying the knot: the cystine signature and molecular-recognition processes of the vascular endothelial growth factor family of angiogenic cytokines. FEBS J. 278, 4304–4322 (2011).
Kintzing, J. R. & Cochran, J. R. Engineered knottin peptides as diagnostics, therapeutics, and drug delivery vehicles. Curr. Opin. Chem. Biol. 34, 143–150 (2016).
Al-Salama, Z. T. & Syed, Y. Y. Plecanatide: first global approval. Drugs 77, 593–598 (2017).
Veiseh, M. et al. Tumor paint: a chlorotoxin:Cy5.5 bioconjugate for intraoperative visualization of cancer foci. Cancer Res. 67, 6882–6888 (2007).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Bendtsen, J. D., Nielsen, H., von Heijne, G. & Brunak, S. Improved prediction of signal peptides: SignalP 3.0. J. Mol. Biol. 340, 783–795 (2004).
Liou, Y. C., Tocilj, A., Davies, P. L. & Jia, Z. Mimicry of ice structure by surface hydroxyls and water of a β-helix antifreeze protein. Nature 406, 322–324 (2000).
Liang, Z., Sottrup-Jensen, L., Aspán, A., Hall, M. & Söderhäll, K. Pacifastin, a novel 155-kDa heterodimeric proteinase inhibitor containing a unique transferrin chain. Proc. Natl. Acad. Sci. USA 94, 6682–6687 (1997).
Moura, A., Savageau, M. A. & Alves, R. Relative amino acid composition signatures of organisms and environments. PLoS One 8, e77319 (2013).
The UniProt Consortium. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 45, D158–D169 (2017).
Bandaranayake, A. D. et al. Daedalus: a robust, turnkey platform for rapid production of decigram quantities of active recombinant proteins in human cell lines using novel lentiviral vectors. Nucleic Acids Res. 39, e143 (2011).
Finton, K. A. et al. Autoreactivity and exceptional CDR plasticity (but not unusual polyspecificity) hinder elicitation of the anti-HIV antibody 4E10. PLoS Pathog. 9, e1003639 (2013).
Blommel, P. G. & Fox, B. G. A combined approach to improving large-scale production of tobacco etch virus protease. Protein Expr. Purif. 55, 53–68 (2007).
Cabrita, L. D. et al. Enhancing the stability and solubility of TEV protease using in silico design. Protein Sci. 16, 2360–2367 (2007).
Kapust, R. B. et al. Tobacco etch virus protease: mechanism of autolysis and rational design of stable mutants with wild-type catalytic proficiency. Protein Eng. 14, 993–1000 (2001).
Cesaratto, F., López-Requena, A., Burrone, O. R. & Petris, G. Engineered tobacco etch virus (TEV) protease active in the secretory pathway of mammalian cells. J. Biotechnol. 212, 159–166 (2015).
Wingerd, J. S. et al. The tarantula toxin β/δ-TRTX-Pre1a highlights the importance of the S1-S2 voltagesensor region for sodium channel subtype selectivity. Sci. Rep. 7, 974 (2017).
Liu, Q., Liu, Q. & Hendrickson, W. A. Robust structural analysis of native biological macromolecules from multi-crystal anomalous diffraction data. Acta Crystallogr. D Biol. Crystallogr. 69, 1314–1332 (2013).
Leaver-Fay, A. et al. ROSETTA3: an object-oriented software suite for the simulation and design of macromolecules. Methods Enzymol. 487, 545–574 (2011).
Kwong, P. D., McDonald, N. Q., Sigler, P. B. & Hendrickson, W. A. Structure of beta 2-bungarotoxin: potassium channel binding by Kunitz modules and targeted phospholipase action. Structure 3, 1109–1119 (1995).
Zhu, L. M., Gao, B. & Zhu, S. Y. Origin of neurotoxins from defensins. Sheng Li Xue Bao 67, 239–247 (2015).
Tarr, D. E. Establishing a reference array for the CS-αβ superfamily of defensive peptides. BMC Res. Notes 9, 490 (2016).
Wu, Y., Gao, B. & Zhu, S. New fungal defensin-like peptides provide evidence for fold change of proteins in evolution. Biosci. Rep. 37, BSR20160438 (2017).
Bhardwaj, G. et al. Accurate de novo design of hyperstable constrained peptides. Nature 538, 329–335 (2016).
Simonet, G., Claeys, I. & Broeck, J. V. Structural and functional properties of a novel serine protease inhibiting peptide family in arthropods. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 132, 247–255 (2002).
Lippens, G., Najib, J., Wodak, S. J. & Tartar, A. NMR sequential assignments and solution structure of chlorotoxin, a small scorpion toxin that blocks chloride channels. Biochemistry 34, 13–21 (1995).
Tsunemi, M., Matsuura, Y., Sakakibara, S. & Katsube, Y. Crystal structure of an elastase-specific inhibitor elafin complexed with porcine pancreatic elastase determined at 1.9 A resolution. Biochemistry 35, 11570–11576 (1996).
Francart, C., Dauchez, M., Alix, A. J. & Lippens, G. Solution structure of R-elafin, a specific inhibitor of elastase. J. Mol. Biol. 268, 666–677 (1997).
Rost, B. Twilight zone of protein sequence alignments. Protein Eng. 12, 85–94 (1999).
Baker, D. & Sali, A. Protein structure prediction and structural genomics. Science 294, 93–96 (2001).
Bailey, T. L. & Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 2, 28–36 (1994).
Schneider, T. D. & Stephens, R. M. Sequence logos: a new way to display consensus sequences. Nucleic Acids Res. 18, 6097–6100 (1990).
Janssen, B. J., Schirra, H. J., Lay, F. T., Anderson, M. A. & Craik, D. J. Structure of Petunia hybrida defensin 1, a novel plant defensin with five disulfide bonds. Biochemistry 42, 8214–8222 (2003).
Knappik, A. & Plückthun, A. An improved affinity tag based on the FLAG peptide for the detection and purification of recombinant antibody fragments. Biotechniques 17, 754–761 (1994).
Kim, Y. et al. High-throughput protein purification for X-ray crystallography and NMR. Adv. Protein Chem. Struct. Biol. 75, 85–105 (2008).
Wang, J., Yadav, V., Smart, A. L., Tajiri, S. & Basit, A. W. Toward oral delivery of biopharmaceuticals: an assessment of the gastrointestinal stability of 17 peptide drugs. Mol. Pharm. 12, 966–973 (2015).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Sheldrick, G. M. Experimental phasing with SHELXC/D/E: combining chain tracing with density modification. Acta Crystallogr. D Biol. Crystallogr. 66, 479–485 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Graef, J. D. et al. Validation of a high-throughput, automated electrophysiology platform for the screening of nicotinic agonists and antagonists. J. Biomol. Screen. 18, 116–127 (2013).
Gillie, D. J., Novick, S. J., Donovan, B. T., Payne, L. A. & Townsend, C. Development of a high-throughput electrophysiological assay for the human ether-à-go-go related potassium channel hERG. J. Pharmacol. Toxicol. Methods 67, 33–44 (2013).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Kearse, M. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Theobald, D. L. & Steindel, P. A. Optimal simultaneous superpositioning of multiple structures with missing data. Bioinformatics 28, 1972–1979 (2012).
Theobald, D. L. & Wuttke, D. S. THESEUS: maximum likelihood superpositioning and analysis of macromolecular structures. Bioinformatics 22, 2171–2172 (2006).
Pei, J., Kim, B. H. & Grishin, N. V. PROMALS3D: a tool for multiple protein sequence and structure alignments. Nucleic Acids Res. 36, 2295–2300 (2008).
Pei, J. & Grishin, N. V. PROMALS3D: multiple protein sequence alignment enhanced with evolutionary and three-dimensional structural information. Methods Mol. Biol. 1079, 263–271 (2014).
Acknowledgements
We thank J. Carter for substantial logistical support, and J. Simon, A. Mhyre and N. Nairn for helpful discussions. The project was supported by NIH grants R01CA135491-07 (J.M.O.) and T32-H600035 (C.D.B.), Project Violet, the Wissner-Slivka Foundation, the Kismet Foundation, the Sarah M. Hughes Foundation, Strong4Sam, Yahn Bernier and Beth McCaw, Len and Norma Klorfine, Anne Croco and Pocket Full of Hope.
Author information
Authors and Affiliations
Contributions
C.E.C., M.M.G., C.M., M.-Y.B., D.M., J.M.O. and R.K.S. designed experiments, analyzed data, administered the project and wrote the manuscript. C.E.C., C.M., A.D.B., W.A.J., W.d.v.d.S. and S.M.T. developed the CDP production and analysis pipeline. M.M.G., P.B.R. and R.K.S. performed crystallographic structure determinations and analyses. C.E.C., M.M.G., M.-Y.B., D.M. and S.E.B. performed phylogenetic and taxonomic analyses. C.E.C., A.D.B., W.A.J., M.C., S.E.B., W.d.v.d.S., S.M.T., A.W. and M.K.C. produced and analyzed CDPs. C.E.C., M.M.G., K.P., C.D.B. and R.K.S. designed and developed SuperTEV.
Corresponding authors
Ethics declarations
Competing interests
J.M.O. is a founder and shareholder of Blaze Bioscience, Inc.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 CDP-production workflow.
An overview of the CDP production pipeline is presented, showing the sequential steps in the process. Step 1 is large-scale suspension culture of transduced HEK293F cells. Step 2 is purification, as depicted by PAGE analysis of an intact Scn-CDP fusion protein immediately following IMAC elution (non-reducing PAGE shows a single, highly-pure fusion protein). Step 3 is preparative cleavage with TEV protease (non-reducing SDS-PAGE shows near complete liberation of the CDP from the fusion construct). Step 4 is preparative-scale separation of the CDP from the Scn fusion partner using RPC. Step 5 is bulk lyophilization of the retentate. Step 6 is final analysis by comparative reduced/non-reduced PAGE and RPC. Note the aberrant mobility typical of purified CDPs in the comparative reduced/non-reduced PAGE analysis (inset, lower right, Step 6), where typical proteins with intrachain cystines migrate faster under non-reducing loading conditions than reducing ones.
Supplementary Figure 2 SuperTEV design, production and characterization.
a, A cartoon representation of the structure of TEV protease (1LVB; Phan, J. et al., J Biol Chem. 277, 50564–50572, 2002) is shown, with the positions of cysteine residues colored yellow on the ribbon, the position of a modified valine residue colored magenta, and asparagine residues at potential N-glycosylation sites shown as orange spheres. A peptide substrate is shown in a licorice-stick representation, colored by atom type. Cysteine residues were substituted as indicated to stabilize the protein, eliminate the need for reducing agents in storage/cleavage buffers, and permit expression through eukaryotic secretion pathways. V219 was replaced to stabilize the protein, based on analyses with Rosetta (Leaver-Fay, A. et al., Methods Enzymol. 487, 545–574, 2011). In order to prevent deleterious cryptic N-glycosylation, the indicated mutations were made at N23, to prevent a folding-incompatible N-glycan, and T173, to prevent steric occlusion of the active site by a non-native glycan on N171. Two other potential N-glycan sites (N68, N52) were not modified, as N-glycans at these positions would not be predicted to affect folding or activity. Combinations and variations of these mutations have been proposed before (Blommel, P.G. et al., Protein Expr Purif. 55, 53–68, 2007; Cabrita, L.D. et al., Protein Sci. 16, 2360–2367, 2007; Kapust, R.B. et al., Protein Eng. 14, 993–1000, 2001; Cesaratto, F. et al., J Biotechnol. 212, 159–166, 2015), though not combined into a single construct. Unlike native TEV, SuperTEV, produced in bacterial or mammalian expression platforms, was monodisperse by SEC, stable in storage, and more thermostable in solution (TmTEV = 52° C; TmSuperTEV = 56° C, as determined by CD). When produced in mammalian cells, SuperTEV showed a molecular weight increase consistent with glycosylation at N52 and N68, can be readily prepared free of contaminating endotoxins, and can be functionally co-expressed in the mammalian secretion pathway. b, TEV and SuperTEV show identical activities on a Scn-TEV site-target protein (CD86 ectodomain) fusion construct by PAGE analysis over a time course of 4 h (uncleaved fusion protein: purple triangle; cleaved Scn: red triangle; cleaved partner protein: blue triangle; time points at 0, 5, 10, 15, 20, 30, 40, 50, 60, 90, 120, 180, and 240 m). c, At top are shown schematic representations of the Daedalus Lentivirus SuperTEV and blue fluorescent protein fusion constructs, incorporating Igκ leader peptide (white box), linker (gray lines), Scn fusion partner (red circle), TEV scission site (green diamond), hexa-histidine purification tag (blue diamond), SuperTEV (green oval), and blue fluorescent protein (BFP) sequences (Subach, O.M. et al., Chem Biol. 15, 1116–1124, 2008). The Scn fusion is designed to optimize expression of SuperTEV in the Daedalus system, and the BFP fusion was engineered to confirm activity of co-expressed SuperTEV in mammalian culture. When both viruses are co-transduced into HEK293F cells, the culture supernatant contains efficiently-cleaved BFP, as shown in the fluorescing samples at the bottom of the frame. Well 1 shows media from untransduced cells; well 2: IMAC flow-through of media from cells transduced with Scn-BFP; well 3: IMAC eluate of media from cells transduced with Scn-BFP; well 4: IMAC flow-through of media from cells co-transduced with Scn-BFP and Scn-SuperTEV fusion proteins; well 5: IMAC eluate of media from cells co-transduced with Scn-BFP and Scn-SuperTEV.
Supplementary Figure 3 Comparisons of CDP structures.
CDP crystal structures determined as part of this work (target number indicated) are shown superimposed on the closest related structure available (PDB accession code indicated) in a backbone representation, colored by atom type, with explicit disulfide bonds. Carbon atoms from the determined crystal structures are colored gray, and from the related PDB structure, green. All molecules in the crystallographic AU, or the sheaf of top NMR solutions available in the deposited structure, were independently aligned and are shown. Red arrows highlight some of the cystines with conformations that significantly deviated between NMR and crystal structures, eg., an inversion of handedness.
Supplementary Figure 4 Ion-channel inhibitory activity of selected CDPs.
The effect of 37 selected CDPs on 20 ion channels at two dosages (0.2 or 20 μM) is shown with triangular gnomons, with grayscale intensity indicating the degree of the effect. Percent inhibition is defined as: (1-(ICDP /Icontrol)) x 100%. Individual assays were run in duplicate, and only effects greater than 3σ are shown. Though these results are more extensive than previously reported for many of these molecules, they are largely consistent with prior reports of ion channel activity for specific examples. CDPs #9 and #11 were active on a subset of potassium channels, consistent with previous reports (Laraba-Djebari, F. et al., J Biol Chem. 269, 32835–32843, 1994; Kozminsky-Atias, A. et al., FEBS Lett. 581, 2478–2484, 2007) and #9 is related to Kaliotoxin-1, a known Kv channel inhibitor, though #11 shows additional activity against hERG (Kv11.1) and Nav1.7 channels (not previously tested). CDP #14 had been previously reported to inhibit hERG channels (Restano-Cassulini, R. et al., Neurochem Res. 33, 1525–1533, 2008), but showed activity against additional potassium channels. CDP #21, aka Hadrucalcin, and CDP #55, aka Opicalcin-2, had been reported to inhibit calcium channels (ryanodine receptors) (Schwartz, E.F. et al., Br J Pharmacol. 157, 392–403, 2009; Xiao, L. et al., J Gen Physiol. 147, 375–394, 2016), but also showed acitivity against additional potassium channels or serotonin receptors (eg., 5-HT3a). CDP #28, CTX, was a known inhibitor of chloride channels (DeBin, J.A. et al., Am J Physiol. 264, C361-369, 1993), but also showed activity against the α4β2 nicotinic receptor, and the hERG and Kv1.2 potassium channels. CDP #46 had been previously reported to inhibit Kv1.3 (Mao, X. et al., Biochem Biophys Res Commun. 360, 728–734, 2007), consistent with these results. CDP #47 had been reported to inhibit small conductance, calcium-activated K channels (Romi-Lebrun, R. et al., Eur J Biochem. 245, 457–464, 1997), but not Kv channels as observed here. CDP #48 had previously been demonstrated to bind to hERG in the patent literature (US7326772 B2), but this confirmed hERG inhibition. CDPs #49 and #53 have been reported to inhibit Kv1.3 (Romi-Lebrun, R. et al., Biochemistry. 36, 13473–13482, 1997; Abdel-Mottaleb, Y. et al., Toxicon. 51, 1424–1430, 2008), and showed strong Kv1.3 inhibition in these results. CDP #62 and #63 showed activity on various subsets of potassium channels (D'Suze, G. et al., Arch Biochem Biophys. 430, 256–263, 2004; Legros, C. et al., FEBS Lett. 390, 81–84, 1996), #67 on voltage-sensitive calcium channels (Lampe, R. A. et al., Mol Pharmacol. 44, 451–460, 1993), and #86 showed activity on tetrodotoxin-sensitive sodium channels (eg., Nav1.7) (Peng, K. et al., J Biol Chem. 277, 47564–47571, 2002).
Supplementary Figure 5 Taxonomic distribution of CDPs and knotted CDPs in the PDB.
The distribution of CDPs (left column) and knotted CDPs (right column) by phylum (top row), class (middle row), and order (bottom row) identified in the PDB is shown as pie charts. Taxonomic division and percentage of total is labeled.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Note
Supplementary Table 1
Classification of experimentally-determined CDP structures in the PDB
Supplementary Table 2
Targeted CDP sequences, expression outcomes, properties, and biochemical characteristics
Supplementary Table 3
CDPs expressed using the high-throughput platform
Supplementary Table 4
Crystallization conditions and crystallographic validation statistics for CDP crystal structures
Supplementary Dataset 1
Properties of CDPs
Rights and permissions
About this article
Cite this article
Correnti, C.E., Gewe, M.M., Mehlin, C. et al. Screening, large-scale production and structure-based classification of cystine-dense peptides. Nat Struct Mol Biol 25, 270–278 (2018). https://doi.org/10.1038/s41594-018-0033-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0033-9
This article is cited by
-
CysPresso: a classification model utilizing deep learning protein representations to predict recombinant expression of cysteine-dense peptides
BMC Bioinformatics (2023)
-
Atomic model for core modifying region of human fatty acid synthase in complex with Denifanstat
Nature Communications (2023)
-
C-Terminal Amidation of Chlorotoxin Does Not Affect Tumour Cell Proliferation and Has No Effect on Toxin Cytotoxicity
International Journal of Peptide Research and Therapeutics (2021)
-
Laboratory information management software for engineered mini-protein therapeutic workflow
BMC Bioinformatics (2019)