Abstract
microRNAs (miRNAs) repress target transcripts through partial complementarity. By contrast, highly complementary miRNA-binding sites within viral and artificially engineered transcripts induce miRNA degradation in vitro and in cell lines. Here, we show that a genome-encoded transcript harboring a near-perfect and deeply conserved miRNA-binding site for miR-29 controls zebrafish and mouse behavior. This transcript originated in basal vertebrates as a long noncoding RNA (lncRNA) and evolved to the protein-coding gene NREP in mammals, where the miR-29-binding site is located within the 3′ UTR. We show that the near-perfect miRNA site selectively triggers miR-29b destabilization through 3′ trimming and restricts its spatial expression in the cerebellum. Genetic disruption of the miR-29 site within mouse Nrep results in ectopic expression of cerebellar miR-29b and impaired coordination and motor learning. Thus, we demonstrate an endogenous target-RNA-directed miRNA degradation event and its requirement for animal behavior.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Kim, V. N., Han, J. & Siomi, M. C. Biogenesis of small RNAs in animals. Nat. Rev. Mol. Cell Biol. 10, 126–139 (2009).
Meister, G. Argonaute proteins: functional insights and emerging roles. Nat. Rev. Genet. 14, 447–459 (2013).
Carrington, J. C. & Ambros, V. Role of microRNAs in plant and animal development. Science 301, 336–338 (2003).
Gurtan, A. M. & Sharp, P. A. The role of miRNAs in regulating gene expression networks. J. Mol. Biol. 425, 3582–3600 (2013).
Friedman, R. C., Farh, K. K., Burge, C. B. & Bartel, D. P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 19, 92–105 (2009).
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Filipowicz, W., Bhattacharyya, S. N. & Sonenberg, N. Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat. Rev. Genet. 9, 102–114 (2008).
Davis, E. et al. RNAi-mediated allelic trans-interaction at the imprinted Rtl1/Peg11 locus. Curr. Biol. 15, 743–749 (2005).
Yekta, S., Shih, I. H. & Bartel, D. P. MicroRNA-directed cleavage of HOXB8 mRNA. Science 304, 594–596 (2004).
Moran, Y. et al. Cnidarian microRNAs frequently regulate targets by cleavage. Genome Res. 24, 651–663 (2014).
Shin, C. et al. Expanding the microRNA targeting code: functional sites with centered pairing. Mol. Cell 38, 789–802 (2010).
Karginov, F. V. et al. Diverse endonucleolytic cleavage sites in the mammalian transcriptome depend upon microRNAs, Drosha, and additional nucleases. Mol. Cell 38, 781–788 (2010).
Park, J. H. et al. Degradome sequencing reveals an endogenous microRNA target in C. elegans. FEBS Lett. 587, 964–969 (2013).
Hutvágner, G. & Zamore, P. D. A microRNA in a multiple-turnover RNAi enzyme complex. Science 297, 2056–2060 (2002).
Ameres, S. L. et al. Target RNA-directed trimming and tailing of small silencing RNAs. Science 328, 1534–1539 (2010).
de la Mata, M. et al. Potent degradation of neuronal miRNAs induced by highly complementary targets. EMBO Rep. 16, 500–511 (2015).
Marcinowski, L. et al. Degradation of cellular mir-27 by a novel, highly abundant viral transcript is important for efficient virus replication in vivo. PLoS Pathog. 8, e1002510 (2012).
Baccarini, A. et al. Kinetic analysis reveals the fate of a microRNA following target regulation in mammalian cells. Curr. Biol. 21, 369–376 (2011).
Cazalla, D., Yario, T. & Steitz, J. A. Down-regulation of a host microRNA by a Herpesvirus saimiri noncoding RNA. Science 328, 1563–1566 (2010).
Libri, V. et al. Murine cytomegalovirus encodes a miR-27 inhibitor disguised as a target. Proc. Natl Acad. Sci. USA 109, 279–284 (2012).
Lee, S. et al. Selective degradation of host MicroRNAs by an intergenic HCMV noncoding RNA accelerates virus production. Cell Host Microbe 13, 678–690 (2013).
Ulitsky, I., Shkumatava, A., Jan, C. H., Sive, H. & Bartel, D. P. Conserved function of lincRNAs in vertebrate embryonic development despite rapid sequence evolution. Cell 147, 1537–1550 (2011).
Taylor, G. A. et al. Behavioral characterization of P311 knockout mice. Genes Brain Behav. 7, 786–795 (2008).
Taylor, G. A., Hudson, E., Resau, J. H. & Vande Woude, G. F. Regulation of P311 expression by Met-hepatocyte growth factor/scatter factor and the ubiquitin/proteasome system. J. Biol. Chem. 275, 4215–4219 (2000).
Studler, J. M., Glowinski, J. & Lévi-Strauss, M. An abundant mRNA of the embryonic brain persists at a high level in cerebellum, hippocampus and olfactory bulb during adulthood. Eur. J. Neurosci. 5, 614–623 (1993).
Norton, W. & Bally-Cuif, L. Adult zebrafish as a model organism for behavioural genetics. BMC Neurosci. 11, 90 (2010).
Stewart, A. M. et al. Molecular psychiatry of zebrafish. Mol. Psychiatry 20, 2–17 (2015).
Chodroff, R. A. et al. Long noncoding RNA genes: conservation of sequence and brain expression among diverse amniotes. Genome Biol. 11, R72 (2010).
Bencan, Z., Sledge, D. & Levin, E. D. Buspirone, chlordiazepoxide and diazepam effects in a zebrafish model of anxiety. Pharmacol. Biochem. Behav. 94, 75–80 (2009).
Norton, W. H. et al. Modulation of Fgfr1a signaling in zebrafish reveals a genetic basis for the aggression-boldness syndrome. J. Neurosci. 31, 13796–13807 (2011).
Egan, R. J. et al. Understanding behavioral and physiological phenotypes of stress and anxiety in zebrafish. Behav. Brain Res. 205, 38–44 (2009).
Kadotani, H. et al. Motor discoordination results from combined gene disruption of the NMDA receptor NR2A and NR2C subunits, but not from single disruption of the NR2A or NR2C subunit. J. Neurosci. 16, 7859–7867 (1996).
Galliano, E. et al. Silencing the majority of cerebellar granule cells uncovers their essential role in motor learning and consolidation. Cell Rep. 3, 1239–1251 (2013).
Watanabe, D. et al. Ablation of cerebellar Golgi cells disrupts synaptic integration involving GABA inhibition and NMDA receptor activation in motor coordination. Cell 95, 17–27 (1998).
Ceccarelli, M. et al. Altered cerebellum development and impaired motor coordination in mice lacking the Btg1 gene: Involvement of cyclin D1. Dev. Biol. 408, 109–125 (2015).
Crawley, J. N. Behavioral phenotyping of transgenic and knockout mice: experimental design and evaluation of general health, sensory functions, motor abilities, and specific behavioral tests. Brain Res. 835, 18–26 (1999).
Marazziti, D. et al. Precocious cerebellum development and improved motor functions in mice lacking the astrocyte cilium-, patched 1-associated Gpr37l1 receptor. Proc. Natl Acad. Sci. USA 110, 16486–16491 (2013).
Crawley, J. N. & Paylor, R. A proposed test battery and constellations of specific behavioral paradigms to investigate the behavioral phenotypes of transgenic and knockout mice. Horm. Behav. 31, 197–211 (1997).
Mandillo, S. et al. Mice lacking the Parkinson’s related GPR37/PAEL receptor show non-motor behavioral phenotypes: age and gender effect. Genes Brain Behav. 12, 465–477 (2013).
Conti, L. et al. Niche-independent symmetrical self-renewal of a mammalian tissue stem cell. PLoS Biol. 3, e283 (2005).
Splinter, E. et al. The inactive X chromosome adopts a unique three-dimensional conformation that is dependent on Xist RNA. Genes Dev. 25, 1371–1383 (2011).
Pinzón, N. et al. microRNA target prediction programs predict many false positives. Genome Res. 27, 234–245 (2017).
Papadopoulou, A. S. et al. Deficiency of the miR-29a/b-1 cluster leads to ataxic features and cerebellar alterations in mice. Neurobiol. Dis. 73, 275–288 (2015).
Gatfield, D. et al. Integration of microRNA miR-122 in hepatic circadian gene expression. Genes Dev. 23, 1313–1326 (2009).
Marzi, M. J. et al. Degradation dynamics of microRNAs revealed by a novel pulse-chase approach. Genome Res. 26, 554–565 (2016).
Rissland, O. S., Hong, S. J. & Bartel, D. P. MicroRNA destabilization enables dynamic regulation of the miR-16 family in response to cell-cycle changes. Mol. Cell 43, 993–1004 (2011).
Hwang, H. W., Wentzel, E. A. & Mendell, J. T. A hexanucleotide element directs microRNA nuclear import. Science 315, 97–100 (2007).
Griffiths-Jones, S., Saini, H. K., van Dongen, S. & Enright, A. J. miRBase: tools for microRNA genomics. Nucleic Acids Res. 36, D154–D158 (2008).
Auer, T. O., Duroure, K., De Cian, A., Concordet, J. P. & Del Bene, F. Highly efficient CRISPR/Cas9-mediated knock-in in zebrafish by homology-independent DNA repair. Genome Res. 24, 142–153 (2014).
Hwang, W. Y. et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat. Biotechnol. 31, 227–229 (2013).
Kimmel, C. B., Ballard, W. W., Kimmel, S. R., Ullmann, B. & Schilling, T. F. Stages of embryonic development of the zebrafish. Dev. Dyn. 203, 253–310 (1995).
Yang, H. et al. One-step generation of mice carrying reporter and conditional alleles by CRISPR/Cas-mediated genome engineering. Cell 154, 1370–1379 (2013).
Goujon, M. et al. A new bioinformatics analysis tools framework at EMBL-EBI. Nucleic Acids Res. 38, W695–W699 (2010).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
McWilliam, H. et al. Analysis tool web services from the EMBL-EBI. Nucleic Acids Res. 41, W597–W600 (2013).
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Pall, G. S. & Hamilton, A. J. Improved northern blot method for enhanced detection of small RNA. Nat. Protoc. 3, 1077–1084 (2008).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Kent, W. J., Zweig, A. S., Barber, G., Hinrichs, A. S. & Karolchik, D. BigWig and BigBed: enabling browsing of large distributed datasets. Bioinformatics 26, 2204–2207 (2010).
Davis, M. P., van Dongen, S., Abreu-Goodger, C., Bartonicek, N. & Enright, A. J. Kraken: a set of tools for quality control and analysis of high-throughput sequence data. Methods 63, 41–49 (2013).
Vitsios, D. M. & Enright, A. J. Chimira: analysis of small RNA sequencing data and microRNA modifications. Bioinformatics 31, 3365–3367 (2015).
Kozomara, A. & Griffiths-Jones, S. miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 42, D68–D73 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Gerlai, R., Lahav, M., Guo, S. & Rosenthal, A. Drinks like a fish: zebra fish (Danio rerio) as a behavior genetic model to study alcohol effects. Pharmacol. Biochem. Behav. 67, 773–782 (2000).
Curzon, P., Rustay, N.R. & Browman, K.E. Cued and contextual fear conditioning for rodents in Methods of Behavior Analysis in Neuroscience (ed. Buccafusco, J.J.) (CRC Press, Boca Raton, FL, 2009).
Simon, M. M. et al. A comparative phenotypic and genomic analysis of C57BL/6 J and C57BL/6 N mouse strains. Genome Biol. 14, R82 (2013).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Schulz, E. G. et al. The two active X chromosomes in female ESCs block exit from the pluripotent state by modulating the ESC signaling network. Cell Stem Cell 14, 203–216 (2014).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Acknowledgements
We thank R. Pillai and E. Heard for comments on this manuscript. We also thank all members of the Shkumatava lab for useful discussions and B. Foret and A.-V. Gendrel of the E. Heard lab for advice on ESC differentiation. This work was supported by grants from ERC (FLAME-337440), ATIP-Avenir, La Fondation Bettencourt Schueller and La Fondation pour la Recherche Médicale (FRM DBI201312285578) to A.S. The research leading to these results has received funding from the European Research Council under the European Union’s Seventh Framework Programme (FP7/2007-2013)/ERC grant agreement no. GA 310206 to D.O’C and the European Research Council under the European Union’s Seventh Framework Programme (FP7/2007-2013) grant agreement no. 602805 to W.H.J.N. High-throughput sequencing was performed by the ICGex NGS platform of the Institut Curie, supported by the grants ANR-10-EQPX-03 (Equipex) and ANR-10-INBS-09-08 (France Génomique Consortium) from the Agence Nationale de la Recherche (“Investissements d’Avenir” program), by the Canceropole Ile-de-France and by the SiRIC-Curie program - SiRIC Grant INCa-DGOS- 4654.
Author information
Authors and Affiliations
Contributions
A.B. and A.C.M. contributed to the design, execution and analysis of most of the experiments. C.C. contributed to the design, generation and execution of the NrepmiR-29scr allele. E.G. and S.M. performed and analyzed mouse behavioral experiments. H.C.G., W.H.J.N. and A.B. performed and analyzed zebrafish behavioral experiments. E.P. performed the in situ hybridization staining. Y.A.P.-R. performed the phylogenetic and sequence conservation analyses. G.P.T.-V. and D.O’C. thought of the NrepmiR-29scr allele. A.J.E. performed the bioinformatics analyses of small-RNA sequencing. A.S. conceived the study. D.O’C. and A.S. supervised the study. A.S. and A.C.M. wrote the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Conservation of the libra, Nrep and miR-29 sequences.
(a) Conservation of the miR-29 site and its flanking sequences. The sequence logo based on 30 homologous sequences is shown above representative examples from the indicated species. Asterisks indicate bases conserved in all of the representative examples. The miR-29 site is in blue and underlined. (b) Predicted Nrep pairing with the individual mouse miR-29 family members. Watson-Crick paired nucleotides are in black and connected by vertical lines, whereas wobbled paired nucleotides are in blue. (c) Multiple alignment of the NREP ORF across vertebrates. Amino acids conserved in all species are in blue. An asterisk indicates a putative stop codon. (d) Unrooted consensus Bayesian phylogenetic tree of zebrafish libra and its homologs in 24 vertebrate species. Branch lengths represent the number of substitutions per site as indicated in the scale bar. Posterior probabilities for all branch splits are displayed in the nodes. (e) The miR-29 family members of zebrafish (a and b) and human and mouse (a, b and c). The seed sequence is boxed, bases differing among the individual miR-29s are in red. (f) libra, miR-29a and miR-29b show partially overlapping expression detected by in situ hybridization on adult zebrafish brain sections. Representative sections for each probe are shown (wild-type animals, n=6). Telencephalon (Tel), Tectum opticum (TeO), Corpus Cerebelli (CCe), Medulla Oblongata (MO), Inferior Lobe (IL). (g) Control in situ hybridization on zebrafish adult brain sections using scrambled miRNA probes. (h) Control in situ hybridization on mouse adult brain sections using scrambled miRNA probes. The brain section is outlined with a dashed line. Representative sections for each probe are shown (wild-type zebrafish, n=6 and mouse, n=4).
Supplementary Figure 2 Generation and validation of the libradel and librainv zebrafish mutants.
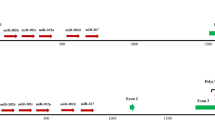
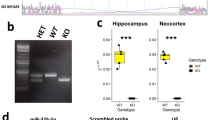
(a) The libradel zebrafish locus showing the positions of the sgRNA Guide 1 and Guide 2 used to generate the libradel mutant. The nucleotides defining the borders of the deleted genomic sequence block are shown in gray and delineated by red arrowheads. The short guide and PAM sequences are indicated with blue and pink blocks, respectively. (b) The librainv zebrafish locus showing the positions of the sgRNA Guide 2 and Guide 3 used to generate the librainv mutant and the inverted portion of exon 3 (hashed block). The nucleotides defining the borders of the inverted part of the libra transcript in the librainv mutant are indicated in lavender and delineated by red arrowheads. The short guide and PAM sequences are indicated with blue and pink blocks, respectively. (c) Generation of the libradel and librainv alleles was confirmed by RNA-Seq. The positions of the libra locus together with that of the adjacent annotated upstream and downstream loci are indicated, and the corresponding H3K4me3 ChIP-Seq, 3P-Seq and RNA-Seq tracks from wild type, libradel and librainv zebrafish are shown. The y-axis of the RNA-Seq tracks represents raw read counts. The inverted portion of the libra transcript in librainv mutant is indicated in lavender.
Supplementary Figure 3 Behavior of the libradel and librainv zebrafish mutants.
(a) The number (n) of entries, (b) time in seconds (s) spent near the novel object testing boldness and (c) latency to approach the novel object in seconds (s) of wild type and libra mutants. Each dot represents an individual adult fish from one experiment; wild-type animals, n=14; libradel animals, n=15; librainv animals, n=15. (d) Velocity in centimeters per second (cm/s), (e) total distance swum in centimeters (cm) and (f) time in seconds (s) spent at the aggressive display recorded in the aggression test for the indicated genotypes. Each dot represents an individual adult fish from one experiment; wild-type animals, n=12; libradel animals, n=15; librainv animals, n=14; **P < 0.01; n.s., not significant. Data are presented as mean ± s.e.m., One-way ANOVA and Dunnett’s multiple comparisons tests were performed for all analyses except for latency to approach the novel object where Kruskal-Wallis and Dunn’s multiple comparisons tests were performed. The novel-object boldness and aggression tests were each performed in one experiment. Detailed statistical analyses and source data are provided in Supplementary Table 1 and Supplementary Dataset 2.
Supplementary Figure 4 Generation and expression analyses of the mouse NrepmiR-29scr allele.
(a) The mouse Nrep locus showing the positions of the sgRNA Guide 4 and Guide 5 used to generate the NrepmiR-29scr mutant mice. The short guide and PAM sequences are indicated with blue and pink blocks, respectively. The wild-type and scrambled miR-29 site is highlighted in green, and the scrambled nucleotides are in red. A BamHI restriction site was introduced to facilitate mutant screening. (b) Hematoxylin and eosin staining of brain sections from wild-type (left) and NrepmiR-29scr (right) mice. Upper panel, whole brain; bottom panel, zoom-in of the cerebellum. Scale bars represent 50 µm. Representative sections for the indicated genotype are shown (wild-type animals, n=4; NrepmiR-29scr animals, n=4). Cortex (Cx), Striatum (Str), Hippocampus (Hip), Thalamus (Thal), Cerebellum (Cb). (c) qRT-PCR analysis of Nrep expression in the cerebellum of wild-type (blue) and NrepmiR-29scr (purple) mutant mice (wild-type animals, n=6; NrepmiR-29scr animals, n=5). Gapdh was used as a reference gene. Data are presented as mean ± s.e.m.; unpaired t-test: n.s., not significant at 95% confidence level. Detailed statistical analyses and source data for panel c are provided in Supplementary Table 1 and Supplementary Dataset 2.
Supplementary Figure 5 Behavior of NrepmiR-29scr mice.
(a) Work flow of the three phases of the contextual and cued fear conditioning test. (b) Number of activity counts before shock in the fear conditioning test; wild-type (WT) animals, n=16; NrepmiR-29scr animals, n=15. Data are presented as individual data points and mean ± s.e.m. (c-e) Number of freezing episodes of NrepmiR-29scr (purple) and wild type (blue) mice in the fear conditioning test presented in 1 min intervals during (c) the conditioning session (arrows indicate the time of the CS + foot-shock application); (d) the context session; (e) the cue session (CS presented during min 3-4 and 7-8). Wild-type animals, n=14; NrepmiR-29scr animals, n=12. Data are presented as mean ± s.e.m. (f-g) Elevated plus maze test to assess anxiety-related behavior in NrepmiR-29scr (purple) and wild-type (blue) mice. (f) frequency of total arm entries; wild-type animals, n=15; NrepmiR-29scr animals, n=15; and (g) percent time spent in open arms; wild-type animals, n=15; NrepmiR-29scr animals, n=13. Data are presented as individual data points and mean ± s.e.m. Detailed statistical analyses and source data for panels b-g are provided in Supplementary Table 1 and Supplementary Dataset 2.
Supplementary Figure 6 Molecular characterization of NrepmiR-29scr cell lines.
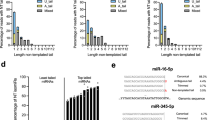
(a) Nrep and miR-29a, b and c expression in wild-type and NrepmiR-29scr ESCs and NPCs detected by RNA blots. The NPC portion of this blot is also shown in Fig. 5a. Three biological replicates are shown for each genotype, technical triplicates were performed for miR-29b and technical duplicates were performed for Nrep, miR-29a and miR-29c. 18S rRNA and U6 RNAs were used as loading references. Uncropped blot images are shown in Supplementary Dataset 1. (b) qRT-PCR analysis of Oct4 and Nestin expression in wild-type and NrepmiR-29scr ESCs and NPCs. Each dot represents an individual biological replicate ESC or NPC population. qRT-PCRs for each biological replicate were performed in technical triplicate. Gapdh was used as a reference gene. Data are presented as mean ± s.e.m.; unpaired t-test: n.s., not significant at 95% confidence level. (c) qRT-PCR analysis of Nrep expression in wild-type and NrepmiR-29scr NPCs. Each dot represents an individual biological replicate NPC population. qRT-PCRs for each biological replicate were performed in technical triplicate. Gapdh was used as a reference gene. Data are presented as mean ± s.e.m.; unpaired t-test: n.s., not significant at 95% confidence level. (d) qRT-PCR analysis of pri-miR-29 expression in wild type and NrepmiR-29scr NPCs. Each dot represents an individual biological replicate NPC population. β-actin was used as a reference gene. Data are presented as mean ± s.e.m.; unpaired t-test: n.s., not significant at 95% confidence level (e) Normalized expression of miR-29 star sequences from small RNA sequencing reads. Each bar represents an individual biological replicate NPC population; unpaired t-test: n.s., not significant. (f) The proportion of small RNA sequencing reads with coverage of at least n indicated nucleotides (nt) along the length of the top five most abundant NPC mmu-miRNAs (miR-21a, let-7i, let-7f, miR-148a and miR-9). For each miRNA, the canonical length and regions tested for trimming and tailing are delineated. Five nucleotides either side of the canonical length were used to test for significant differences in the mean values over these positions using a paired t-test for difference of means. Three biological replicates are shown for each genotype. n.s., not significant at 95% confidence level. Detailed statistical analyses and source data for panels b-f are provided in Supplementary Table 1 and Supplementary Dataset 2.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 2 and 3 and Supplementary Notes 1–3
Supplementary Dataset 1
Uncropped RNA blots
Rights and permissions
About this article
Cite this article
Bitetti, A., Mallory, A.C., Golini, E. et al. MicroRNA degradation by a conserved target RNA regulates animal behavior. Nat Struct Mol Biol 25, 244–251 (2018). https://doi.org/10.1038/s41594-018-0032-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0032-x
This article is cited by
-
Structural variants in the Epb41l4a locus: TAD disruption and Nrep gene misregulation as hypothetical drivers of neurodevelopmental outcomes
Scientific Reports (2024)
-
Pan-cancer onco-signatures reveal a novel mitochondrial subtype of luminal breast cancer with specific regulators
Journal of Translational Medicine (2023)
-
Screening of Drosophila microRNA-degradation sequences reveals Argonaute1 mRNA’s role in regulating miR-999
Nature Communications (2023)
-
LncRNA MIR200CHG inhibits EMT in gastric cancer by stabilizing miR-200c from target-directed miRNA degradation
Nature Communications (2023)
-
microRNAs in action: biogenesis, function and regulation
Nature Reviews Genetics (2023)