Abstract
Camelid single-domain antibody fragments (‘nanobodies’) provide the remarkable specificity of antibodies within a single 15-kDa immunoglobulin VHH domain. This unique feature has enabled applications ranging from use as biochemical tools to therapeutic agents. Nanobodies have emerged as especially useful tools in protein structural biology, facilitating studies of conformationally dynamic proteins such as G-protein-coupled receptors (GPCRs). Nearly all nanobodies available to date have been obtained by animal immunization, a bottleneck restricting many applications of this technology. To solve this problem, we report a fully in vitro platform for nanobody discovery based on yeast surface display. We provide a blueprint for identifying nanobodies, demonstrate the utility of the library by crystallizing a nanobody with its antigen, and most importantly, we utilize the platform to discover conformationally selective nanobodies to two distinct human GPCRs. To facilitate broad deployment of this platform, the library and associated protocols are freely available for nonprofit research.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hamers-Casterman, C. et al. Naturally occurring antibodies devoid of light chains. Nature 363, 446–448 (1993).
Muyldermans, S. Nanobodies: natural single-domain antibodies. Annu. Rev. Biochem. 82, 775–797 (2013).
Irannejad, R. et al. Conformational biosensors reveal GPCR signalling from endosomes. Nature 495, 534–538 (2013).
Rasmussen, S. G. et al. Structure of a nanobody-stabilized active state of the β(2) adrenoceptor. Nature 469, 175–180 (2011).
Staus, D. P. et al. Allosteric nanobodies reveal the dynamic range and diverse mechanisms of G-protein-coupled receptor activation. Nature 535, 448–452 (2016).
Manglik, A., Kobilka, B. K. & Steyaert, J. Nanobodies to study G protein-coupled receptor structure and function. Annu. Rev. Pharmacol. Toxicol. 57, 19–37 (2017).
Moutel, S. et al. NaLi-H1: A universal synthetic library of humanized nanobodies providing highly functional antibodies and intrabodies. eLife 5, e16228 (2016).
Gao, J., Sidhu, S. S. & Wells, J. A. Two-state selection of conformation-specific antibodies. Proc. Natl. Acad. Sci. USA 106, 3071–3076 (2009).
Rizk, S. S. et al. Allosteric control of ligand-binding affinity using engineered conformation-specific effector proteins. Nat. Struct. Mol. Biol. 18, 437–442 (2011).
Adams, J. J. & Sidhu, S. S. Synthetic antibody technologies. Curr. Opin. Struct. Biol. 24, 1–9 (2014).
Kayushin, A., Korosteleva, M. & Miroshnikov, A. Large-scale solid-phase preparation of 3′-unprotected trinucleotide phosphotriesters–precursors for synthesis of trinucleotide phosphoramidites. Nucleosides Nucleotides Nucleic Acids 19, 1967–1976 (2000).
Kayushin, A. L. et al. A convenient approach to the synthesis of trinucleotide phosphoramidites–synthons for the generation of oligonucleotide/peptide libraries. Nucleic Acids Res 24, 3748–3755 (1996).
Boder, E. T. & Wittrup, K. D. Yeast surface display for screening combinatorial polypeptide libraries. Nat. Biotechnol. 15, 553–557 (1997).
Kruse, A. C. et al. Activation and allosteric modulation of a muscarinic acetylcholine receptor. Nature 504, 101–106 (2013).
Rakestraw, J. A., Sazinsky, S. L., Piatesi, A., Antipov, E. & Wittrup, K. D. Directed evolution of a secretory leader for the improved expression of heterologous proteins and full-length antibodies in Saccharomyces cerevisiae. Biotechnol. Bioeng. 103, 1192–1201 (2009).
Orlean, P. Architecture and biosynthesis of the Saccharomyces cerevisiae cell wall. Genetics 192, 775–818 (2012).
Makrides, S. C. et al. Extended in vivo half-life of human soluble complement receptor type 1 fused to a serum albumin-binding receptor. J. Pharmacol. Exp. Ther. 277, 534–542 (1996).
Van Roy, M. et al. The preclinical pharmacology of the high affinity anti-IL-6R Nanobody ALX-0061 supports its clinical development in rheumatoid arthritis. Arthritis Res. Ther. 17, 135 (2015).
Tijink, B. M. et al. Improved tumor targeting of anti-epidermal growth factor receptor Nanobodies through albumin binding: taking advantage of modular Nanobody technology. Mol. Cancer Ther. 7, 2288–2297 (2008).
Kim, C. C., Wilson, E. B. & DeRisi, J. L. Improved methods for magnetic purification of malaria parasites and haemozoin. Malar. J. 9, 17 (2010).
Rasmussen, S. G. et al. Crystal structure of the β2 adrenergic receptor-Gs protein complex. Nature 477, 549–555 (2011).
Ring, A. M. et al. Adrenaline-activated structure of β2-adrenoceptor stabilized by an engineered nanobody. Nature 502, 575–579 (2013).
Manglik, A. & Kobilka, B. The role of protein dynamics in GPCR function: insights from the β2AR and rhodopsin. Curr. Opin. Cell Biol. 27, 136–143 (2014).
Rosenbaum, D. M. et al. Structure and function of an irreversible agonist-β(2) adrenoceptor complex. Nature 469, 236–240 (2011).
Staus, D. P. et al. Regulation of β2-adrenergic receptor function by conformationally selective single-domain intrabodies. Mol. Pharmacol. 85, 472–481 (2014).
Vijayan, D., Young, A., Teng, M. W. L. & Smyth, M. J. Targeting immunosuppressive adenosine in cancer. Nat. Rev. Cancer 17, 709–724 (2017).
Hino, T. et al. G-protein-coupled receptor inactivation by an allosteric inverse-agonist antibody. Nature 482, 237–240 (2012).
Weiskopf, K. et al. Engineered SIRPα variants as immunotherapeutic adjuvants to anticancer antibodies. Science 341, 88–91 (2013).
Manglik, A. et al. Structural insights into the dynamic process of β2-adrenergic receptor signaling. Cell 161, 1101–1111 (2015).
Zou, Y., Weis, W. I. & Kobilka, B. K. N-terminal T4 lysozyme fusion facilitates crystallization of a G protein coupled receptor. PLoS One 7, e46039 (2012).
Jaakola, V. P. et al. The 2.6 angstrom crystal structure of a human A2A adenosine receptor bound to an antagonist. Science 322, 1211–1217 (2008).
Whorton, M. R. et al. A monomeric G protein-coupled receptor isolated in a high-density lipoprotein particle efficiently activates its G protein. Proc. Natl. Acad. Sci. USA 104, 7682–7687 (2007).
Liberles, S. D. & Buck, L. B. A second class of chemosensory receptors in the olfactory epithelium. Nature 442, 645–650 (2006).
Caffrey, M. & Cherezov, V. Crystallizing membrane proteins using lipidic mesophases. Nat. Protoc. 4, 706–731 (2009).
Hein, K. L. et al. Crystallographic analysis reveals a unique lidocaine binding site on human serum albumin. J. Struct. Biol. 171, 353–360 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr 66, 213–221 (2010).
Acknowledgements
Financial support for this work was provided by the Vallee Foundation (A.C.K.), the Smith Family Foundation (A.C.K.), National Institutes of Health grants 5DP5OD021345 (A.C.K.), 1DP5OD023048 (A.M.), and 1DP5OD023088 (A.M.R.), the Lundbeck Foundation (grant no. R37-A3457 to S.G.F.R.), and the Danish Independent Research Council (grant no. 0602-02407B to S.G.F.R.).
Author information
Authors and Affiliations
Contributions
A.C.K., A.M., and C.M. designed and generated the nanobody library. C.M., A.S.B., and A.C.K. performed quality control of the library. C.M., R.P., S.Z., J.X.O., D.H., and A.M. prepared antigens, performed selections, and isolated nanobody binders. C.M., R.P., and S.C.E. characterized nanobodies. A.M.R., A.M., and A.C.K. developed the modified yeast display system and associated expression vectors. M.W. and S.G.F.R. purified the A2A adenosine receptor. C.M., A.M., and A.C.K. wrote the manuscript with assistance and input from all coauthors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Biochemical validation of nanobody clones
(a–k) Randomly chosen nanobodies were expressed and purified from E. coli, then analyzed by size exclusion chromatography to assess monodispersity. (l) SDS-PAGE analysis of nanobody purity following one-step nickel affinity purification.
Supplementary Figure 2 Design of display system
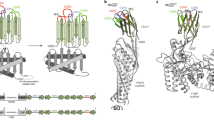
(a) The display system was engineered using the high affinity SIRPα variant CV1 as a test protein, and its ligand CD47 ectodomain as the staining reagent. A biotin tag is schematized as a glowing red circle. (b) Length of the stalk region determines accessibility of a displayed protein as a function of molecular weight. (c) Analytical flow cytometry plots showing length dependence for two staining reagents: CD47 biotin and α-HA antibody. The 649 amino acid long stalk was used in all nanobody display experiments.
Supplementary Figure 3 Analysis of HSA-targeted nanobodies
(a) Library design was assessed by monitoring the change in amino acid frequency in CDR3 throughout selection rounds with HSA as the antigen. Few changes were observed, with the only notable trend a modest increase in basic residue frequency and a decline in acidic residue frequency. (b) Assessment of Nb.b201 binding to human serum albumin by surface plasmon resonance, comparison with mouse serum albumin which shows no detectable binding. (c) 2Fo-Fc composite omit map contoured at 1.5 σ for antigen bound Nb.b201. The structure of both bound (yellow) and free (gray) forms of the nanobody are shown, highlighting structural divergence. (d) 2Fo-Fc composite omit map contoured at 1.5 σ for free Nb.b201.
Supplementary Figure 4 Discovery of nanobodies with nonpurified antigen
(a) Conditioned medium containing adiponectin (left lane) was used for selection of nanobodies. It shows a complex mixture of proteins as assessed by SDS-PAGE. For reference, purified adiponectin is shown in the right lane. Adiponectin exists as a mix of 16-mers, hexamers, and trimers. (b) Schematic of selection process. Fluorescent anti-FLAG antibody was used to specifically mark those yeast cells that display adiponectin-binding nanobodies. (c) Flow cytometry analysis of final clone pool, showing that the library was highly enriched in adiponectin-binding clones. (d) Sequences of five selected clones showed highly diverse CDR3 sequence composition and length. (e) Binding assessed using in vitro pull-down with purified adiponectin globular domain. (f) Binding to adiponectin was further confirmed in vitro using surface plasmon resonance. Kinetic fit is shown for clone Nb.AQ103.
Supplementary Figure 5 Affinity of β2AR-binding nanobodies
On-yeast titration to estimate affinity of β2AR binding nanobodies. EC50 values are summarized in the lower right. Bottom panel shows measurement of conformational selectivity for selected clones as assessed by flow cytometry.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1 and 2 and Supplementary Note 1
Rights and permissions
About this article
Cite this article
McMahon, C., Baier, A.S., Pascolutti, R. et al. Yeast surface display platform for rapid discovery of conformationally selective nanobodies. Nat Struct Mol Biol 25, 289–296 (2018). https://doi.org/10.1038/s41594-018-0028-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0028-6
This article is cited by
-
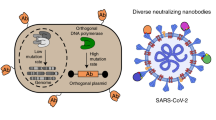
Single-domain antibodies against SARS-CoV-2 RBD from a two-stage phage screening of universal and focused synthetic libraries
BMC Infectious Diseases (2024)
-
Looking back at 30 years of Nature Structural & Molecular Biology
Nature Structural & Molecular Biology (2024)
-
Extracellular targeted protein degradation: an emerging modality for drug discovery
Nature Reviews Drug Discovery (2024)
-
Highly biased agonism for GPCR ligands via nanobody tethering
Nature Communications (2024)
-
Antibodies expand the scope of angiotensin receptor pharmacology
Nature Chemical Biology (2024)