Abstract
The shelterin protein TRF2 assembles protective T loops at chromosome ends by stimulating intramolecular invasion of the telomeric G-rich single-stranded DNA (ssDNA) overhang into the duplex telomeric array. The other shelterin factor, TRF1, is thought to mainly facilitate telomeric dsDNA replication without directly participating in end protection. Here we show that in vitro human TRF2 stimulates invasion of G-rich TERRA-like RNA into telomeric dsDNA, leading to formation of telomeric RNA–DNA hybrids (telR loops). The N-terminal basic domain of TRF2 binds to TERRA-like RNA and enables TRF2 to promote efficient RNA invasion. TRF1, through its N-terminal acidic domain, counteracts TRF2-mediated RNA invasion but not ssDNA invasion. In vivo, when TRF1 is depleted or replaced with a variant lacking the acidic domain, TRF2 induces formation of telR loops, which in turn cause telomere loss. Hence, uncontrolled TRF2 threatens telomere integrity, and TRF1 directly supports end protection by suppressing harmful telR loops.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Martínez, P. & Blasco, M. A. Replicating through telomeres: a means to an end. Trends Biochem. Sci. 40, 504–515 (2015).
Azzalin, C. M. & Lingner, J. Telomere functions grounding on TERRA firma. Trends Cell. Biol. 25, 29–36 (2015).
Azzalin, C. M., Reichenbach, P., Khoriauli, L., Giulotto, E. & Lingner, J. Telomeric repeat containing RNA and RNA surveillance factors at mammalian chromosome ends. Science 318, 798–801 (2007).
Nergadze, S. G. et al. CpG-island promoters drive transcription of human telomeres. RNA 15, 2186–2194 (2009).
Schoeftner, S. & Blasco, M. A. Developmentally regulated transcription of mammalian telomeres by DNA-dependent RNA polymerase II. Nat. Cell. Biol. 10, 228–236 (2008).
Arnoult, N. & Karlseder, J. Complex interactions between the DNA-damage response and mammalian telomeres. Nat. Struct. Mol. Biol. 22, 859–866 (2015).
de Lange, T. How shelterin solves the telomere end-protection problem. Cold Spring Harb. Symp. Quant. Biol. 75, 167–177 (2010).
Sfeir, A. & de Lange, T. Removal of shelterin reveals the telomere end-protection problem. Science 336, 593–597 (2012).
Broccoli, D., Smogorzewska, A., Chong, L. & de Lange, T. Human telomeres contain two distinct Myb-related proteins, TRF1 and TRF2. Nat. Genet. 17, 231–235 (1997).
Sfeir, A. et al. Mammalian telomeres resemble fragile sites and require TRF1 for efficient replication. Cell 138, 90–103 (2009).
Zimmermann, M., Kibe, T., Kabir, S. & de Lange, T. TRF1 negotiates TTAGGG repeat-associated replication problems by recruiting the BLM helicase and the TPP1/POT1 repressor of ATR signaling. Genes. Dev. 28, 2477–2491 (2014).
Martínez, P. et al. Increased telomere fragility and fusions resulting from TRF1 deficiency lead to degenerative pathologies and increased cancer in mice. Genes. Dev. 23, 2060–2075 (2009).
Denchi, E. L. & de Lange, T. Protection of telomeres through independent control of ATM and ATR by TRF2 and POT1. Nature 448, 1068–1071 (2007).
Griffith, J. D. et al. Mammalian telomeres end in a large duplex loop. Cell 97, 503–514 (1999).
Doksani, Y., Wu, J. Y., de Lange, T. & Zhuang, X. Super-resolution fluorescence imaging of telomeres reveals TRF2-dependent T-loop formation. Cell 155, 345–356 (2013).
Amiard, S. et al. A topological mechanism for TRF2-enhanced strand invasion. Nat. Struct. Mol. Biol. 14, 147–154 (2007).
Benarroch-Popivker, D. et al. TRF2-mediated control of telomere DNA topology as a mechanism for chromosome-end protection. Mol. Cell. 61, 274–286 (2016).
Court, R., Chapman, L., Fairall, L. & Rhodes, D. How the human telomeric proteins TRF1 and TRF2 recognize telomeric DNA: a view from high-resolution crystal structures. EMBO Rep. 6, 39–45 (2005).
Wu, H., Lima, W. F. & Crooke, S. T. Investigating the structure of human RNase H1 by site-directed mutagenesis. J. Biol. Chem. 276, 23547–23553 (2001).
Deng, Z., Norseen, J., Wiedmer, A., Riethman, H. & Lieberman, P. M. TERRA RNA binding to TRF2 facilitates heterochromatin formation and ORC recruitment at telomeres. Mol. Cell. 35, 403–413 (2009).
Apte, M. S. & Cooper, J. P. Life and cancer without telomerase: ALT and other strategies for making sure ends (don’t) meet. Crit. Rev. Biochem. Mol. Biol. 52, 57–73 (2017).
Arora, R. et al. RNaseH1 regulates TERRA-telomeric DNA hybrids and telomere maintenance in ALT tumour cells. Nat. Commun. 5, 5220 (2014).
Boguslawski, S. J. et al. Characterization of monoclonal antibody to DNA.RNA and its application to immunodetection of hybrids. J. Immunol. Methods 89, 123–130 (1986).
Smith, S., Giriat, I., Schmitt, A. & de Lange, T. Tankyrase, a poly(ADP-ribose) polymerase at human telomeres. Science 282, 1484–1487 (1998).
Dynek, J. N. & Smith, S. Resolution of sister telomere association is required for progression through mitosis. Science 304, 97–100 (2004).
Nguyen, H. D. et al. Functions of replication protein A as a sensor of R loops and a regulator of RNaseH1. Mol. Cell. 65, 832–847.e4. (2017).
Muñoz, P., Blanco, R., Flores, J. M. & Blasco, M. A. XPF nuclease-dependent telomere loss and increased DNA damage in mice overexpressing TRF2 result in premature aging and cancer. Nat. Genet. 37, 1063–1071 (2005).
Nera, B., Huang, H. S., Lai, T. & Xu, L. Elevated levels of TRF2 induce telomeric ultrafine anaphase bridges and rapid telomere deletions. Nat. Commun. 6, 10132 (2015).
Tong, A. S. et al. ATM and ATR signaling regulate the recruitment of human telomerase to telomeres. Cell. Rep. 13, 1633–1646 (2015).
Balk, B. et al. Telomeric RNA-DNA hybrids affect telomere-length dynamics and senescence. Nat. Struct. Mol. Biol. 20, 1199–1205 (2013).
Graf, M. et al. Telomere length determines TERRA and R-Loop regulation through the cell cycle. Cell 170, 72–85.e14 (2017).
Fouché, N. et al. The basic domain of TRF2 directs binding to DNA junctions irrespective of the presence of TTAGGG repeats. J. Biol. Chem. 281, 37486–37495 (2006).
Poulet, A. et al. TRF2 promotes, remodels and protects telomeric Holliday junctions. EMBO J. 28, 641–651 (2009).
Schmutz, I., Timashev, L., Xie, W., Patel, D. J. & de Lange, T. TRF2 binds branched DNA to safeguard telomere integrity. Nat. Struct. Mol. Biol. 24, 734–742 (2017).
Santos-Pereira, J. M. & Aguilera, A. R loops: new modulators of genome dynamics and function. Nat. Rev. Genet. 16, 583–597 (2015).
Lee, Y. W. & Kim, W. T. Telomerase-dependent 3′ G-strand overhang maintenance facilitates GTBP1-mediated telomere protection from misplaced homologous recombination. Plant Cell. 25, 1329–1342 (2013).
Acknowledgements
We thank T. de Lange (The Rockefeller University, New York), J. Karlseder (The Salk Institute for Biological Studies, La Jolla, CA, USA), E. Gilson (IRCAN, Nice, France), S. Smith (The Skirball Institute of Biomolecular Medicine, New York), S. Leppla (NIAID, Bethesda, MD, USA) and J. Lingner (ISREC, Lausanne, Switzerland) for reagents, the Bioimaging facility of iMM Lisboa and the Scientific Center for Optical and Electron Microscopy of ETHZ for microscopy services, members of the Azzalin laboratory for discussions and J. Lingner for critical reading of the manuscript. This work was initiated at the Institute of Biochemistry of ETH Zürich and supported by grants awarded to C.M.A. by the Swiss National Science Foundation (31003A_160338), the European Research Council (BFTERRA), EMBO (IG3576) and Fundação para a Ciência e a Tecnologia (IF/01269/2015). Y.W.L. was supported by an EMBO long-term fellowship (ALTF 395-2014) and the National Research Foundation of Korea (NRF-2013R1A6A3A03063846). Publication costs were supported through LISBOA-01-0145-FEDER-007391, a project cofunded by FEDER through POR Lisboa 2020 (Programa Operacional Regional de Lisboa, PORTUGAL 2020) and Fundação para a Ciência e a Tecnologia.
Author information
Authors and Affiliations
Contributions
Y.W.L. and C.M.A. conceived and supervised the project; Y.W.L., R.A., H.W. and C.M.A. performed and analyzed the experiments; Y.W.L., R.A. and C.M.A. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Control strand-invasion assays.
(a) Strand invasion assays with 0.6 nM pcDNA6 empty vector or pcDNA6 containing a ~800 bp long telomeric array (p-Tel) in combination with increasing amounts of the indicated RNA and DNA oligonucleotides. (b) Invaded plasmids in experiments as in a were quantified and graphed as the fold increase compared to reactions performed with 2 nM oligonucleotides. Values are means ± SD (n = 3 independent experiments). (c) Reaction products from invasion assays with the indicated oligonucleotides and the p-Tel plasmid were incubated with RNaseH (RH) before electrophoresis. (d) Strand invasion assays with TERRA-like oligonucleotides, the p-Tel plasmid and the indicated recombinant proteins were incubated with RNaseH before electrophoresis. GST-TRF2 concentration was 40 nM, GST-TRF1 and GST-TRF1ΔA concentrations were 0, 10, 20, 40 and 80 nM. inv, invaded plasmid; ss, single stranded RNA or DNA oligonucleotides; *, wells.
Supplementary Figure 2 Recombinant proteins used in this study and their binding to telomeric dsDNA.
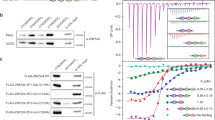
(a) Structural domains of the recombinant proteins used in strand invasion and EMSA assays. GST, glutathione S-transferase; B, basic; A, acidic; TRFH, TRF homodimerization; M, Myb; RH1HBD, RNA:DNA hybrid binding domain of human RNaseH1. Black dots indicate single amino acid substitutions within the TRFH domain of the Topless mutant. (b) 200 ng of recombinant proteins were size-fractionated by SDS-PAGE and stained with EZBlue reagent. Molecular weights of a size marker (m) are on the left in kDa. (c) The indicated proteins (80 nM) were incubated with G-rich RNA or DNA oligonucleotides (10 nM) in the same conditions as for invasion assays, followed by electrophoresis in a denaturing polyacrylamide gel and radioactive signal detection. Note that incubation with proteins did not affect the signal associated to the oligonucleotides, when compared to samples incubated in absence of proteins, thus excluding DNase, RNase and phosphatase contaminations. (d) EMSAs with a radiolabeled dsDNA fragment containing 24 TTAGGG repeats (0.26 nM) and increasing amounts of the indicated recombinant proteins. (e) EMSAs with a radiolabeled dsDNA fragment comprising a ~0.8 kb long (TTAGGG)n array (0.03 nM) and increasing amounts of the indicated recombinant proteins. Bound probes were quantified and graphed as fraction of the total signal within each lane. Values are means ± SD (n = 3 independent experiments). (f) EMSAs as in d using GST fused to A, B or A and B (AB) domains. *, wells.
Supplementary Figure 3 Control in vitro experiments.
(a) Strand invasion assays with (UUAGGG)5 TERRA-like oligonucleotides (10 nM), p-Tel plasmid (0.6 nM) and increasing amounts of the indicated recombinant proteins. Invaded plasmids (inv) were quantified and graphed as the fold increase compared with samples lacking proteins. Values are means ± SD (n = 3 independent experiments). A curve for GST-TRF2 was included as a reference. ss, single stranded oligonucleotides; *, wells. (b) EMSAs with a radiolabeled dsDNA fragment comprising a ~0.8 kb long (TTAGGG)n array (0.03 nM) and increasing amounts of GST-TRF1 or GST-TRF1ΔA, in presence or absence of GST-TRF2. Note that the mobility of GST-TRF2-bound probes is further retarded when GST-TRF1 or GST-TRF1ΔA are included in the reactions. The table on the right is to facilitate direct comparison of DNA and protein concentrations used in EMSAs and strand invasion assays. (c) Strand invasion assays were performed with 0.6 nM p-Tel plasmid, cold (UUAGGG)5 TERRA-like oligonucleotides (10 nM), and increasing amounts of GST-TRF1 or GST-TRF1ΔA, in presence or absence of GST-TRF2. Reaction products were fractionated in agarose gels, transferred to nitrocellulose membranes and subjected to western blot analysis with anti-TRF1 and anti-TRF2 antibodies. TRF2 signals were quantified and graphed as the fold increase compared to reactions without TRF1 proteins. Values are means ± SD (n = 5 independent experiments). (d) EMSAs with a radiolabeled RNA:DNA duplex comprising 5 TERRA-like RNA repeats and 5 complementary C-rich telomeric DNA repeats. A GST-fusion with the hybrid-binding domain of human RNaseH1 (GST-RH1HBD) was used as a positive control. (e) EMSAs performed with the indicated recombinant proteins (40 nM) and radiolabeled RNA or DNA oligonucleotides (0.25 nM). Bound and free oligonucleotide positions are indicated. *, wells.
Supplementary Figure 4 TelR loops and telomere instability in HeLa cells depleted for TRF1 and/or expressing TRF1ΔA.
(a) Western blot analysis of HeLa cells transfected with siRNAs against TRF1 (siT1c or siT1e) and infected with retroviruses expressing fl-TRF1, fl-ΔA, RH1wt or RH1CD, or with empty vector (ev) retroviruses. TRF1, fl-TRF1 and fl-ΔA are detected using an anti-TRF1 antibody. Endogenous RNaseH1 and Myc-tagged RNaseH1 variants are detected using an anti-RNaseH1 antibody. Actin serves as a loading control. (b) Telomeric dot-blots of S9.6 DRIP experiments. Nucleic acids were collected 4 days after siRNA transfections and expression of ectopic proteins. In, input; Bd, beads only control; IP, S9.6 immunoprecipitated material. (c) pSer33 immunostaining (green) combined with telomeric DNA FISH (red) on cells transfected with siRNAs against TRF1 and infected with retroviruses expressing fl-TRF1 or fl-ΔA. Arrowheads point to examples of pSer33 TIFs. (d) Partial metaphases hybridized in situ with telomeric probes. Telomeres are in red, DNA in blue. Filled arrowheads point to TFEs, unfilled arrowheads to FTs. (e) pSer33 immunostaining combined with telomeric DNA FISH (red) on cells transfected with siRNAs against TRF1 and infected with retroviruses expressing myc-tagged RH1wt or RH1CD. Arrowheads point to pSer33 TIFs. (f) Western blot analysis of retrovirus-infected HeLa cells harvested 4 days after infections. (g) Telomeric dot-blot hybridization of S9.6 DRIPs of cells as in f. Signals are graphed as the fraction of input found in IPs after normalization to ev/ev cells. Values are means ± SD (n = 3 independent experiments). (h) Examples of pSer33 immunostaining on cells as in f. Arrowheads point to pSer33 TIFs. (i) Percentage of cells as in h with at least 3 pSer33 TIFs graphed as means ± SD (n = 3 independent experiments). *P < 0.05, **P < 0.005, ***P < 0.0001 (two-tailed Student’s t-test). (l) Quantification of TFEs in cells as in f harvested 7 days after infections. At least 4500 chromosomes from 3 independent experiments were analyzed for each condition. Dots are percentages of TFEs per chromosome end in one metaphase. Black bars are means. **P < 0.005, ***P < 0.0001 (Mann-Whitney U test). Scale bar in panels c, d, e and h, 5 μm.
Supplementary Figure 5 TRF1 A domain suppresses telR-loop formation and telomeric pSer33 accumulation in U2OS cells.
(a) Western blot analysis of U2OS cells transfected with siRNAs against TRF1 (T1c and T1e) or control siRNAs (Ct), or infected with lentiviruses expressing doxycycline inducible fl-TRF1 or fl-ΔA, or with empty vector (ev) control lentiviruses. TRF1, fl-TRF1 and fl-ΔA are detected using an anti-TRF1 antibody. Actin serves as a loading control. (b) Telomeric and Alu repeat dot-blot hybridization of S9.6 DRIPs of cells as in a. Nucleic acids were collected 3 days after siRNA transfections or 4 days of ectopic protein expression. In, input (1 or 5% as indicated); Bd, beads only control (100%); IP, S9.6 immunoprecipitated material (100%). RH: RNaseH treatment of nucleic acids prior to immunoprecipitation. (c) Quantifications of experiments as in b. Signals are graphed as the fraction of In DNA found in IPs after subtraction of Bd-associated signal. Values are means ± SD (n = 6 independent experiments for the graph on the left, 1 experiment for the graph in the center, 3 independent experiments for the graph on the right). (d) Examples of pSer33 immunostaining (green) combined with telomeric DNA FISH (red) on cells as in a. Arrowheads point to pSer33 TIFs. Scale bar, 5 μm. An example of extensive colocalization between pSer33 and telomeric signals is shown. The percentages of cells with at least 5 TIFs were graphed as means ± SD (n = 3 independent experiments). **P < 0.005, ***P < 0.0001 (two-tailed Student’s t-test).
Supplementary Figure 6 Ectopic fl-TRF1 and fl-ΔA localize at telomeres.
(a) Flag immunostaining (green) combined with telomeric DNA FISH (red) on HeLa cells infected with pLPC-derived retroviruses expressing fl-TRF1 and fl-ΔA from a constitutive promoter. Numbers are percentages of cells expressing the ectopic proteins. (b) Flag (green) and TRF2 (red) immunostaining on U2OS cells infected with pLVX-TetOne-Puro-derived lentiviruses expressing fl-TRF1 or fl-ΔA from a doxycycline-inducible promoter. Cells were treated with doxycycline for 24 hours (+dox) or left untreated (-dox). Numbers are percentages of cells expressing the ectopic proteins. (c) Immunostaining using an anti-Flag antibody (green) combined with an antibody raised against a peptide comprising the acidic domain of TRF1 (red) on HeLa and U2OS cells infected with pLVX-TetOne-Puro-derived lentiviruses expressing fl-TRF1 or fl-ΔA. Cells were treated with doxycycline for 24 hours (+dox). Note that expression of fl-ΔA substantially reduces the signal generated by the TRF1 A domain antibody within the same cells indicating displacement of endogenous TRF1 from telomeres. Scale bars, 5 μm.
Supplementary Figure 7 Tankyrase1 depletion does not affect telR-loop levels.
(a) Western blot analysis of HeLa and U2OS cells transfected with an siRNA against Tankyrase 1 (siTNK1) or a control siRNA (siCt) using anti-TNK1 and Lamin A and C (LMNA and LMNC; loading control) antibodies. Cells were harvested 4 days after transfections. (b) Cells as in a were stained with DAPI and mitotic cells were counted and graphed as means ± SD (n = 3 independent experiments). Examples of DAPI-stained mitotic cells (white arrowheads) are shown on the right. Scale bar, 30 μm. (c) Telomeric dot-blot hybridization of S9.6 DRIPs of cells as in a. In, input; Bd, beads only control; IP, S9.6 immunoprecipitated material. S9.6 DRIP quantifications are shown on the right. Signals are graphed as the fraction of In DNA found in IPs after subtraction of Bd-associated signal and normalization over siCt-transfected cells. Values are means ± SD (n = 4 independent experiments).
Supplementary Figure 8 TelR-loop accumulation and telomeric defects in TRF1-compromised HeLa cells depend on TRF2.
(a) Western blot analysis of HeLa cells transfected with siRNAs against TRF1 (siT1) or TRF2 (siT2), either alone or in combination, or with a control siRNA (siCt). Actin serves as a loading control. (b) Telomeric dot-blot hybridization of S9.6 DRIPs in cells as in a harvested 4 days after transfections. In, input; Bd, beads only control; IP, S9.6 immunoprecipitated material. Signals are graphed on the right as the fraction of In DNA found in IPs after subtraction of Bd-associated signal and normalization over siCt-transfected cells. Values are means ± SD (n = 8 independent experiments). *P < 0.05, **P < 0.005 (two-tailed Student’s t-test). (c) Western blot analysis of HeLa cells infected with retroviruses expressing fl-TRF1 or fl-ΔA and transfected with siT2 or siCt. TRF1, fl-TRF1 and fl-ΔA are detected using an anti-TRF1 antibody. (d) Telomeric dot-blot hybridization of S9.6 DRIPs in cells as in c harvested 4 days after infections/transfections. In, input; Bd, beads only control; IP, S9.6 immunoprecipitated material. (e) pSer33 immunostaining (green) combined with telomeric DNA FISH (red). Cells were harvested 4 days after infections/transfections. Arrowheads point to examples of pSer33 TIFs. (f) Partial metaphases from cells as in c hybridized in situ with telomeric probes. Cells were harvested 7 days after infections/transfections. Filled arrowheads point to TFEs, unfilled arrowheads to FTs. Plots are quantification of TFEs and FTs. At least 4500 chromosomes from three independent experiments were analyzed for each condition. Dots are percentages of TFEs and FTs per chromosome end in one metaphase. Black bars are means. **P < 0.005, ***P < 0.0001 (vs ev/siCt; Mann-Whitney U test). Scale bar in panels e and f, 5 μm.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Table 1
Supplementary Data Set 1
Uncropped images for gels used in main figures
Rights and permissions
About this article
Cite this article
Lee, Y.W., Arora, R., Wischnewski, H. et al. TRF1 participates in chromosome end protection by averting TRF2-dependent telomeric R loops. Nat Struct Mol Biol 25, 147–153 (2018). https://doi.org/10.1038/s41594-017-0021-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-017-0021-5
This article is cited by
-
Homology directed telomere clustering, ultrabright telomere formation and nuclear envelope rupture in cells lacking TRF2B and RAP1
Nature Communications (2023)
-
Exploring the Causal Relationship Between Telomere Biology and Alzheimer’s Disease
Molecular Neurobiology (2023)
-
The landscape of aging
Science China Life Sciences (2022)
-
TERRA transcription destabilizes telomere integrity to initiate break-induced replication in human ALT cells
Nature Communications (2021)
-
ADAR1 RNA editing enzyme regulates R-loop formation and genome stability at telomeres in cancer cells
Nature Communications (2021)