Abstract
BiP is the endoplasmic member of the Hsp70 family. BiP is regulated by several co-chaperones including the nucleotide-exchange factor (NEF) Bap (Sil1 in yeast). Bap is a two-domain protein. The interaction of the Bap C-terminal domain with the BiP ATPase domain is sufficient for its weak NEF activity. However, stimulation of the BiP ATPase activity requires full-length Bap, suggesting a complex interplay of these two factors. Here, single-molecule FRET experiments with mammalian proteins reveal that Bap affects the conformation of both BiP domains, including the lid subdomain, which is important for substrate binding. The largely unstructured Bap N-terminal domain promotes the substrate release from BiP. Thus, Bap is a conformational regulator affecting both nucleotide and substrate interactions. The preferential interaction with BiP in its ADP state places Bap at a late stage of the chaperone cycle, in which it coordinates release of substrate and ADP, thereby resetting BiP for ATP and substrate binding.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Kim, Y. E., Hipp, M. S., Bracher, A., Hayer-Hartl, M. & Hartl, F. U. Molecular chaperone functions in protein folding and proteostasis. Annu. Rev. Biochem. 82, 323–355 (2013).
Kampinga, H. H. & Craig, E. A. The HSP70 chaperone machinery: J proteins as drivers of functional specificity. Nat. Rev. Mol. Cell Biol. 11, 579–592 (2010).
Otero, J. H., Lizák, B. & Hendershot, L. M. Life and death of a BiP substrate. Semin. Cell Dev. Biol. 21, 472–478 (2010).
Qi, R. et al. Allosteric opening of the polypeptide-binding site when an Hsp70 binds ATP. Nat. Struct. Mol. Biol. 20, 900–907 (2013).
Marcinowski, M. et al. Substrate discrimination of the chaperone BiP by autonomous and cochaperone-regulated conformational transitions. Nat. Struct. Mol. Biol. 18, 150–158 (2011).
Buck, T. M., Wright, C. M. & Brodsky, J. L. The activities and function of molecular chaperones in the endoplasmic reticulum. Semin. Cell Dev. Biol. 18, 751–761 (2007).
Behnke, J., Feige, M. J. & Hendershot, L. M. BiP and its nucleotide exchange factors Grp170 and Sil1: mechanisms of action and biological functions. J. Mol. Biol. 427, 1589–1608 (2015).
Mapa, K. et al. The conformational dynamics of the mitochondrial Hsp70 chaperone. Mol. Cell 38, 89–100 (2010).
Bukau, B. & Horwich, A. L. The Hsp70 and Hsp60 chaperone machines. Cell 92, 351–366 (1998).
Mayer, M. P. & Bukau, B. Hsp70 chaperones: cellular functions and molecular mechanism. Cell. Mol. Life Sci. 62, 670–684 (2005).
Theyssen, H., Schuster, H. P., Packschies, L., Bukau, B. & Reinstein, J. The second step of ATP binding to DnaK induces peptide release. J. Mol. Biol. 263, 657–670 (1996).
Cyr, D. M. Swapping nucleotides, tuning Hsp70. Cell 133, 945–947 (2008).
Chung, K. T., Shen, Y. & Hendershot, L. M. BAP, a mammalian BiP-associated protein, is a nucleotide exchange factor that regulates the ATPase activity of BiP. J. Biol. Chem. 277, 47557–47563 (2002).
Yan, M., Li, J. & Sha, B. Structural analysis of the Sil1-Bip complex reveals the mechanism for Sil1 to function as a nucleotide-exchange factor. Biochem. J. 438, 447–455 (2011).
Tyson, J. R. & Stirling, C. J. LHS1 and SIL1 provide a lumenal function that is essential for protein translocation into the endoplasmic reticulum. EMBO J. 19, 6440–6452 (2000).
Shomura, Y. et al. Regulation of Hsp70 function by HspBP1: structural analysis reveals an alternate mechanism for Hsp70 nucleotide exchange. Mol. Cell 17, 367–379 (2005).
Anttonen, A. K. et al. The gene disrupted in Marinesco-Sjögren syndrome encodes SIL1, an HSPA5 cochaperone. Nat. Genet. 37, 1309–1311 (2005).
Senderek, J. et al. Mutations in SIL1 cause Marinesco-Sjögren syndrome, a cerebellar ataxia with cataract and myopathy. Nat. Genet. 37, 1312–1314 (2005).
Howes, J., Shimizu, Y., Feige, M. J. & Hendershot, L. M. C-terminal mutations destabilize SIL1/BAP and can cause Marinesco-Sjögren syndrome. J. Biol. Chem. 287, 8552–8560 (2012).
Siegenthaler, K. D., Pareja, K. A., Wang, J. & Sevier, C. S. An unexpected role for the yeast nucleotide exchange factor Sil1 as a reductant acting on the molecular chaperone BiP. eLife 6, e24141 (2017).
Blond-Elguindi, S., Fourie, A. M., Sambrook, J. F. & Gething, M. J. Peptide-dependent stimulation of the ATPase activity of the molecular chaperone BiP is the result of conversion of oligomers to active monomers. J. Biol. Chem. 268, 12730–12735 (1993).
Marcion, G. et al. C-terminal amino acids are essential for human heat shock protein 70 dimerization. Cell Stress Chaperones 20, 61–72 (2015).
Preissler, S. et al. Physiological modulation of BiP activity by trans-protomer engagement of the interdomain linker. eLife 4, e08961 (2015).
Packschies, L. et al. GrpE accelerates nucleotide exchange of the molecular chaperone DnaK with an associative displacement mechanism. Biochemistry 36, 3417–3422 (1997).
Ha, J. H. & McKay, D. B. ATPase kinetics of recombinant bovine 70 kDa heat shock cognate protein and its amino-terminal ATPase domain. Biochemistry 33, 14625–14635 (1994).
Marcinowski, M. et al. Conformational selection in substrate recognition by Hsp70 chaperones. J. Mol. Biol. 425, 466–474 (2013).
Schneider, M. et al. BiPPred: combined sequence- and structure-based prediction of peptide binding to the Hsp70 chaperone BiP. Proteins 84, 1390–1407 (2016).
Hartmann, C., Antes, I. & Lengauer, T. IRECS: a new algorithm for the selection of most probable ensembles of side-chain conformations in protein models. Protein Sci. 16, 1294–1307 (2007).
Antes, I. DynaDock: A new molecular dynamics-based algorithm for protein-peptide docking including receptor flexibility. Proteins 78, 1084–1104 (2010).
Wei, J., Gaut, J. R. & Hendershot, L. M. In vitro dissociation of BiP-peptide complexes requires a conformational change in BiP after ATP binding but does not require ATP hydrolysis. J. Biol. Chem. 270, 26677–26682 (1995).
Kudryavtsev, V. et al. Combining MFD and PIE for accurate single-pair Förster resonance energy transfer measurements. ChemPhysChem 13, 1060–1078 (2012).
Antonik, M., Felekyan, S., Gaiduk, A. & Seidel, C. A. Separating structural heterogeneities from stochastic variations in fluorescence resonance energy transfer distributions via photon distribution analysis. J. Phys. Chem. B 110, 6970–6978 (2006).
Kalinin, S., Felekyan, S., Antonik, M. & Seidel, C. A. Probability distribution analysis of single-molecule fluorescence anisotropy and resonance energy transfer. J. Phys. Chem. B 111, 10253–10262 (2007).
Sánchez-Rico, C., Voith von Voithenberg, L., Warner, L., Lamb, D. C. & Sattler, M. Effects of fluorophore attachment on protein conformation and dynamics studied by spFRET and NMR spectroscopy. Chemistry 23, 14267–14277 (2017).
Hellenkamp, B., Wortmann, P., Kandzia, F., Zacharias, M. & Hugel, T. Multidomain structure and correlated dynamics determined by self-consistent FRET networks. Nat. Methods 14, 174–180 (2017).
Hillger, F. et al. Probing protein-chaperone interactions with single-molecule fluorescence spectroscopy. Angew. Chem. Int. Ed. Engl. 47, 6184–6188 (2008).
Wilbanks, S. M., DeLuca-Flaherty, C. & McKay, D. B. Structural basis of the 70-kilodalton heat shock cognate protein ATP hydrolytic activity. I. Kinetic analyses of active site mutants. J. Biol. Chem. 269, 12893–12898 (1994).
Feige, M. J. et al. An unfolded CH1 domain controls the assembly and secretion of IgG antibodies. Mol. Cell 34, 569–579 (2009).
Haas, I. G. & Wabl, M. Immunoglobulin heavy chain binding protein. Nature 306, 387–389 (1983).
Flynn, G. C., Pohl, J., Flocco, M. T. & Rothman, J. E. Peptide-binding specificity of the molecular chaperone BiP. Nature 353, 726–730 (1991).
Knarr, G., Gething, M. J., Modrow, S. & Buchner, J. BiP binding sequences in antibodies. J. Biol. Chem. 270, 27589–27594 (1995).
Harrison, C. J., Hayer-Hartl, M., Di Liberto, M., Hartl, F. & Kuriyan, J. Crystal structure of the nucleotide exchange factor GrpE bound to the ATPase domain of the molecular chaperone DnaK. Science 276, 431–435 (1997).
Bracher, A. & Verghese, J. The nucleotide exchange factors of Hsp70 molecular chaperones. Front. Mol. Biosci. https://dx.doi.org/10.3389/fmolb.2015.00010 (2015).
Gowda, N.K.C. et al. Nucleotide exchange factors Fes1 and HspBP1 mimic substrate to release misfolded proteins from Hsp70. Nat. Struct. Mol. Biol. https://doi.org/10.1038/s41594-017-0008-2 (in the press).
Butt, T. R., Edavettal, S. C., Hall, J. P. & Mattern, M. R. SUMO fusion technology for difficult-to-express proteins. Protein Expr. Purif. 43, 1–9 (2005).
Nørby, J. G. Coupled assay of Na+,K+-ATPase activity. Methods Enzymol. 156, 116–119 (1988).
Sun, L. et al. The lid domain of Caenorhabditis elegans Hsc70 influences ATP turnover, cofactor binding and protein folding activity. PLoS ONE 7, e33980 (2012).
Schuck, P. Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and lamm equation modeling. Biophys. J. 78, 1606–1619 (2000).
Gaiser, A. M., Kretzschmar, A. & Richter, K. Cdc37-Hsp90 complexes are responsive to nucleotide-induced conformational changes and binding of further cofactors. J. Biol. Chem. 285, 40921–40932 (2010).
Eggeling, C. et al. Data registration and selective single-molecule analysis using multi-parameter fluorescence detection. J. Biotechnol. 86, 163–180 (2001).
Tomov, T. E. et al. Disentangling subpopulations in single-molecule FRET and ALEX experiments with photon distribution analysis. Biophys. J. 102, 1163–1173 (2012).
Nir, E. et al. Shot-noise limited single-molecule FRET histograms: comparison between theory and experiments. J. Phys. Chem. B 110, 22103–22124 (2006).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Zahn, M. et al. Structural studies on the forward and reverse binding modes of peptides to the chaperone DnaK. J. Mol. Biol. 425, 2463–2479 (2013).
Cock, P. J. A. et al. Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 25, 1422–1423 (2009).
Roe, D. R. & Cheatham, T. E. III PTRAJ and CPPTRAJ: Software for processing and analysis of molecular dynamics trajectory data. J. Chem. Theory Comput. 9, 3084–3095 (2013).
Jorgensen, W. L., Maxwell, D. S. & Tirado-Rives, J. Development and testing of the OPLS all-atom force field on conformational energetics and properties of organic liquids. J. Am. Chem. Soc. 118, 11225–11236 (1996).
Acknowledgements
We thank J. Lawatschek and K. Richter for help with experiments, L. Voith von Voithenberg for helpful and critical discussions regarding spFRET experiments as well as data analysis, W. Kügel for contributions to the spFRET analysis software, G. M. Feind for performing the HDX analysis and K. Buhr for help with the structural modeling of the BiP–peptide complexes. We gratefully acknowledge the financial support by the Deutsche Forschungsgemeinschaft to J.B. (DFG), D.C.L. (SFB1035, A11), I.A. (SFB1035, A10) and M.S. (IGSSE) and by the Ludwig-Maximilians-Universität through the Center for NanoScience (CeNS) and the BioImaging Network (BIN).
Author information
Authors and Affiliations
Contributions
M.R., J. Hochmair and C.N. prepared protein and performed biochemical experiments, K.C.B. did analytical ultracentrifugation experiments, C.Z. and J.R. performed nucleotide-exchange experiments and D.K., J. Hendrix and G.A. performed the spFRET experiments. J.B., I.A. and D.C.L. designed experiments. M.S. and I.A. analyzed the HDX data, and D.K., G.A. and A.B. analyzed the single-molecule data. M.R., D.K., C.N., D.C.L., I.A. and J.B. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Bap sequences and completeness of the available crystal structure.
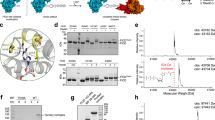
The sequence of human Bap with its signal sequence highlighted in red is depicted. Structural information from yeast Bap (PDB-ID: 3QML) was mapped onto the sequence of human Bap. Sequence regions resolved in the crystal are highlighted in purple. The missing N-terminal amino acids comprise ~25 % of the mature full-length protein.
Supplementary Figure 2 Structural characterization of Bap, Bap-C and Bap-N.
(a) FUV CD spectra were measured for Bap, Bap-C and Bap-N. A comparison between the curves reveals an α-helical secondary structure of Bap and Bap-C and a random coil structure for Bap-N. (b) Thermal transitions of Bap (black) and Bap-C (red) at 220 nm and the corresponding sigmoidal fits showing a one-step thermal denaturation occurring at ~46 °C.
Supplementary Figure 3 Titration measurements to calculate the K d values.
Analytical ultracentrifugation (AUC) measurements were performed in the presence of 1 mM nucleotide to detect the increase in size of the BiP-Bap complex depending on the Bap or Bap-C concentration. The presence of the Bip-Bap complex can be observed by a shift in the sedimentation coefficient with increasing Bap concentrations. (a-c) Titration performed with 1 mM AMP-PNP. (a) AUC sedimentation velocity (AUC-SV) curves for 0.4 μM ATTO 488-labeled BiP-167-638 (final concentration) titrated with different Bap concentrations. (b) AUC-SV curves for 0.4 μM ATTO 488-labeled BiP-167-638 (final concentration) titrated with different concentration of Bap-C. (c) AUC-SV curves for 0.5 μM ATTO 488-labelled BiP-NBD-167 (final concentration) titrated with Bap. The titrations are shown in Figure 2b. (d-g) AUC-SV curves for 0.4 μM ATTO 488-labeled BiP-167-638 (final concentration) titrated with different concentration of Bap (d, e) without nucleotide or (f, g) in presence of 1 mM ADP. (h-j) Interaction of BiP’s isolated NBD with Bap. Sedimentation velocity (SV) was determined by analytical ultracentrifugation (AUC). 0.5 μM Atto 488-labeled BiP NBD-167 was tested without Bap (black line) and in the presence of 5 μM Bap (red line). Measurements were performed (h) in the absence of a nucleotide (apo), (i) with 1 mM ATP or (j) with 1 mM ADP.
Supplementary Figure 4 ATPase stimulation experiments.
(a) Different BiP mutants and DnaK ATPase activity measured in an enzyme coupled assay. Data was normalized to the data of BiP wt. (b) Thin-layer chromatography was employed to separate [α-32P]-ATP from [α-32P]-ADP for the determination of single-turnover ATPase kinetics. 20 μM BiP were mixed with 5 μM ATP containing [α-32P]-ATP. Samples were quenched at defined time points and the band intensities were quantified. Simulation of ATPase in single turnover experiments of BiP and Bap (red) compared to BiP alone (black). The calculated rate constants from the fits are 0.4 min-1 and 1.3 min–1.
Supplementary Figure 5 Prediction of BiP binding sequences in the BAP sequence.
BiPPred prediction scores were determined for all heptamers within the BAP sequence. The score for each heptamer is shown in condensed form as a coloured box below the central residue Z of the heptameric peptide (XXXZXXX)
Supplementary Figure 6 Representative PIE-MFD analysis shown for mutant BiP (167-638) in the presence of 1 mM ADP and 10 μM Bap-N.
(a) A 2D histogram of stoichiometry versus FRET efficiency after filtering for double-labeled molecules. A stoichiometry of ~0.5 indicates a ratio of 1:1 labeling between donor and acceptor fluorophores. (b) A 2D histogram of FRET efficiency versus donor lifetime \({\tau }_{D(A)}\). The relationship between FRET efficiency and donor lifetime for a static population (the static FRET line) is shown in black. No deviation from the static FRET line is observed indicating an absence of conformational dynamics during the ~1 ms long observation time. (c,d) 2D histograms of the burstwise anisotropy of (c) the donor \({r}_{D}\) versus the fluorescence lifetime of the donor \({\tau }_{D(A)}\) and (d) the acceptor \({r}_{A}\) versus the fluorescence lifetime of the acceptor fluorophore \({\tau }_{A}\). The black lines are given by the Perrin equation \(r={r}_{0}/\left(1+\frac{\tau }{\rho }\right)\), where \({r}_{0}\) is the fundamental anisotropy, \(\tau \) is the fluorescence lifetime and \(\rho \) is the rotational correlation time. \({r}_{0}\) is assumed to be 0.4. (e) The time-resolved anisotropy decay for the donor (blue) and acceptor (red) fluorophores are shown. Photons from all double-labelled molecules are pooled together to obtain the cumulative fluorescence decays, from which the anisotropy is determined. Fits to bi-exponential model functions are given by solid lines, accounting for the fast rotation of the fluorophore on the nanosecond timescale and the slow rotation of the protein on the timescale of tens of nanoseconds
Supplementary Figure 7 Origin of the low-FRET conformation in the NBD-lid sensor (BiP-167-519).
(a) SpFRET efficiency distributions of the NBD-lid sensor (BiP-167-519) in the presence of different concentrations of Bap and 1 mM ATP were measured. Sample preparation was performed by incubating 1 μM BiP for 15 min at 37 °C with the final Bap concentrations and then BiP was diluted to spFRET concentrations (~20 pM) while keeping the Bap concentration constant and adding the nucleotide. (b) The spFRET distribution for a small peptide, HTFPVAL (70 μM), bound to BiP in the presence of 1 mM ATP. BiP and BiP with 10 μM Bap in 1 mM ATP are shown for comparison
Supplementary Figure 8 Effect of Bap on the dissociation of ATP from BiP.
SpFRET efficiency distributions of the SBD-lid sensor (BiP-519-638) were determined with different concentrations of ATP (a) in the absence of Bap, (b) in the presence of 10 μM Bap and (c) in the presence of 10 μM Bap-C. (d) The normalized area under the FRET efficiency histograms was summed up until a FRET efficiency value of 0.4 and plotted versus the ATP concentration. The K d of ATP is given by the Boltzmann fit as 4.7 ± 1.3 nM for BiP alone and as 412 ± 7 nM in the presence of Bap. The addition of Bap-C resulted in a K d of 4.1 ± 0.7 nM. Therefore, Bap decreases BiP’s affinity for ATP by a factor of 88 compared to BiP alone or with bound Bap-C. Sample preparation was performed by incubating 1 μM BiP with the final Bap concentration and then keeping the Bap and Bap-C concentrations constant and adding the nucleotide during the dilution of BiP to spFRET concentrations (~20 pM)
Supplementary Figure 9 SpFRET experiments of BiP-Δlid.
(a-c) SpFRET efficiency histograms were measured for the NBD-SBD BiP sensor (gray line), BiP-Δlid (black line), BiP-Δlid in the presence of 10 μM Bap (red line) or BiP-Δlid with 10 μM Bap-C (dark yellow line). Experiments were performed (a) in the absence of nucleotide (apo), (b) with 1 mM ATP or (c) 1 mM ADP included in the buffer. Sample preparation was performed by incubation of BiP (1 μM) with Bap, Bap-C and nucleotide at their final concentrations, which were kept constant during the dilution of BiP to spFRET concentrations (~20 pM). (d-f) SpFRET efficiency histograms for the SBD-lid BiP sensor (BiP 519 - 638), the NBD-SBD BiP sensor (BiP 167 - 519), and BiP- Δlid in 1 mM ADP without peptide or Bap (gray line), with 70 μM HTFPVAL (black line) and with 70 μM HTFPVAL and 10 μM Bap (red line)
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Tables 1–4
Rights and permissions
About this article
Cite this article
Rosam, M., Krader, D., Nickels, C. et al. Bap (Sil1) regulates the molecular chaperone BiP by coupling release of nucleotide and substrate. Nat Struct Mol Biol 25, 90–100 (2018). https://doi.org/10.1038/s41594-017-0012-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-017-0012-6
This article is cited by
-
Identification of ER/SR resident proteins as biomarkers for ER/SR calcium depletion in skeletal muscle cells
Orphanet Journal of Rare Diseases (2022)
-
Multivalent protein–protein interactions are pivotal regulators of eukaryotic Hsp70 complexes
Cell Stress and Chaperones (2022)
-
The Hsp70 chaperone network
Nature Reviews Molecular Cell Biology (2019)
-
Function, evolution, and structure of J-domain proteins
Cell Stress and Chaperones (2019)
-
Nucleotide exchange factors Fes1 and HspBP1 mimic substrate to release misfolded proteins from Hsp70
Nature Structural & Molecular Biology (2018)