Abstract
Mutation of SMARCA4 (BRG1), the ATPase of BAF (mSWI/SNF) and PBAF complexes, contributes to a range of malignancies and neurologic disorders. Unfortunately, the effects of SMARCA4 missense mutations have remained uncertain. Here we show that SMARCA4 cancer missense mutations target conserved ATPase surfaces and disrupt the mechanochemical cycle of remodeling. We find that heterozygous expression of mutants alters the open chromatin landscape at thousands of sites across the genome. Loss of DNA accessibility does not directly overlap with Polycomb accumulation, but is enriched in ‘A compartments’ at active enhancers, which lose H3K27ac but not H3K4me1. Affected positions include hundreds of sites identified as superenhancers in many tissues. Dominant-negative mutation induces pro-oncogenic expression changes, including increased expression of Myc and its target genes. Together, our data suggest that disruption of enhancer accessibility represents a key source of altered function in disorders with SMARCA4 mutations in a wide variety of tissues.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Hodges, C., Kirkland, J. G. & Crabtree, G. R. The many roles of BAF (mSWI/SNF) and PBAF complexes in cancer. Cold Spring Harb. Perspect. Med. 6, a026930 (2016).
Son, E. Y. & Crabtree, G. R. The role of BAF (mSWI/SNF) complexes in mammalian neural development. Am. J. Med. Genet. C. Semin. Med. Genet. 166, 333–349 (2014).
Kadoch, C. et al. Proteomic and bioinformatic analysis of mammalian SWI/SNF complexes identifies extensive roles in human malignancy. Nat. Genet. 45, 592–601 (2013).
Shain, A. H. & Pollack, J. R. The spectrum of SWI/SNF mutations, ubiquitous in human cancers. PLoS One 8, e55119 (2013).
Versteege, I. et al. Truncating mutations of hSNF5/INI1 in aggressive paediatric cancer. Nature 394, 203–206 (1998).
Biegel, J. A. et al. Germ-line and acquired mutations of INI1 in atypical teratoid and rhabdoid tumors. Cancer Res. 59, 74–79 (1999).
Kadoch, C. & Crabtree, G. R. Reversible disruption of mSWI/SNF (BAF) complexes by the SS18-SSX oncogenic fusion in synovial sarcoma. Cell 153, 71–85 (2013).
Davoli, T. et al. Cumulative haploinsufficiency and triplosensitivity drive aneuploidy patterns and shape the cancer genome. Cell 155, 948–962 (2013).
Wilson, B. G. & Roberts, C. W. SWI/SNF nucleosome remodellers and cancer. Nat. Rev. Cancer 11, 481–492 (2011).
Wei, D. et al. SNF5/INI1 deficiency redefines chromatin remodeling complex composition during tumor development. Mol. Cancer Res. 12, 1574–1585 (2014).
Wang, X. et al. SMARCB1-mediated SWI/SNF complex function is essential for enhancer regulation. Nat. Genet. 49, 289–295 (2017).
Oruetxebarria, I. et al. P16INK4a is required for hSNF5 chromatin remodeler-induced cellular senescence in malignant rhabdoid tumor cells. J. Biol. Chem. 279, 3807–3816 (2004).
Kia, S. K., Gorski, M. M., Giannakopoulos, S. & Verrijzer, C. P. SWI/SNF mediates polycomb eviction and epigenetic reprogramming of the INK4b-ARF-INK4a locus. Mol. Cell. Biol. 28, 3457–3464 (2008).
Wilson, B. G. et al. Epigenetic antagonism between polycomb and SWI/SNF complexes during oncogenic transformation. Cancer Cell 18, 316–328 (2010).
Lee, R. S. et al. A remarkably simple genome underlies highly malignant pediatric rhabdoid cancers. J. Clin. Invest. 122, 2983–2988 (2012).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
de Bruijn, D. R. et al. The synovial-sarcoma-associated SS18-SSX2 fusion protein induces epigenetic gene (de)regulation. Cancer Res. 66, 9474–9482 (2006).
Lubieniecka, J. M. et al. Histone deacetylase inhibitors reverse SS18-SSX-mediated polycomb silencing of the tumor suppressor early growth response 1 in synovial sarcoma. Cancer Res. 68, 4303–4310 (2008).
Zhang, Z. K. et al. Cell cycle arrest and repression of cyclin D1 transcription by INI1/hSNF5. Mol. Cell. Biol. 22, 5975–5988 (2002).
Wang, X. et al. Oncogenesis caused by loss of the SNF5 tumor suppressor is dependent on activity of BRG1, the ATPase of the SWI/SNF chromatin remodeling complex. Cancer Res. 69, 8094–8101 (2009).
Mathur, R. et al. ARID1A loss impairs enhancer-mediated gene regulation and drives colon cancer in mice. Nat. Genet. 49, 296–302 (2017).
Alver, B. H. et al. The SWI/SNF chromatin remodelling complex is required for maintenance of lineage specific enhancers. Nat. Commun. 8, 14648 (2017).
Miller, E. L. et al. TOP2 synergizes with BAF chromatin remodeling for both resolution and formation of facultative heterochromatin. Nat. Struct. Mol. Biol. 24, 344–352 (2017).
Fillmore, C. M. et al. EZH2 inhibition sensitizes BRG1 and EGFR mutant lung tumours to TopoII inhibitors. Nature 520, 239–242 (2015).
Imielinski, M. et al. Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell 150, 1107–1120 (2012).
The Cancer Genome Atlas Research Network. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014).
Witkowski, L. et al. Germline and somatic SMARCA4 mutations characterize small cell carcinoma of the ovary, hypercalcemic type. Nat. Genet. 46, 438–443 (2014).
Jelinic, P. et al. Recurrent SMARCA4 mutations in small cell carcinoma of the ovary. Nat. Genet. 46, 424–426 (2014).
Karnezis, A. N. et al. Dual loss of the SWI/SNF complex ATPases SMARCA4/BRG1 and SMARCA2/BRM is highly sensitive and specific for small cell carcinoma of the ovary, hypercalcaemic type. J. Pathol. 238, 389–400 (2016).
Hodis, E. et al. A landscape of driver mutations in melanoma. Cell 150, 251–263 (2012).
Le Loarer, F. et al. SMARCA4 inactivation defines a group of undifferentiated thoracic malignancies transcriptionally related to BAF-deficient sarcomas. Nat. Genet. 47, 1200–1205 (2015).
Love, C. et al. The genetic landscape of mutations in Burkitt lymphoma. Nat. Genet. 44, 1321–1325 (2012).
Lunning, M. A. & Green, M. R. Mutation of chromatin modifiers; an emerging hallmark of germinal center B-cell lymphomas. Blood Cancer J 5, e361 (2015).
Stanton, B. Z. et al. Smarca4 ATPase mutations disrupt direct eviction of PRC1 from chromatin. Nat. Genet. 49, 282–288 (2017).
The Cancer Genome Atlas Research Network et al. The cancer genome atlas pan-cancer analysis project. Nat. Genet. 45, 1113–1120 (2013).
Zehir, A. et al. Mutational landscape of metastatic cancer revealed from prospective clinical sequencing of 10,000 patients. Nat. Med. 23, 703–713 (2017).
Liu, X., Li, M., Xia, X., Li, X. & Chen, Z. Mechanism of chromatin remodelling revealed by the Snf2-nucleosome structure. Nature 544, 440–445 (2017).
Xia, X., Liu, X., Li, T., Fang, X. & Chen, Z. Structure of chromatin remodeler Swi2/Snf2 in the resting state. Nat. Struct. Mol. Biol. 23, 722–729 (2016).
Hauk, G., McKnight, J. N., Nodelman, I. M. & Bowman, G. D. The chromodomains of the Chd1 chromatin remodeler regulate DNA access to the ATPase motor. Mol. Cell 39, 711–723 (2010).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Laurent, B. C., Treich, I. & Carlson, M. The yeast SNF2/SWI2 protein has DNA-stimulated ATPase activity required for transcriptional activation. Genes Dev. 7, 583–591 (1993).
Johnson, T. A., Elbi, C., Parekh, B. S., Hager, G. L. & John, S. Chromatin remodeling complexes interact dynamically with a glucocorticoid receptor-regulated promoter. Mol. Biol. Cell 19, 3308–3322 (2008).
Gurskaya, N. G. et al. Engineering of a monomeric green-to-red photoactivatable fluorescent protein induced by blue light. Nat. Biotechnol. 24, 461–465 (2006).
Sif, S., Stukenberg, P. T., Kirschner, M. W. & Kingston, R. E. Mitotic inactivation of a human SWI/SNF chromatin remodeling complex. Genes Dev. 12, 2842–2851 (1998).
Ho, L. et al. esBAF facilitates pluripotency by conditioning the genome for LIF/STAT3 signalling and by regulating polycomb function. Nat. Cell Biol. 13, 903–913 (2011).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Whyte, W. A. et al. Enhancer decommissioning by LSD1 during embryonic stem cell differentiation. Nature 482, 221–225 (2012).
Francis, N. J., Kingston, R. E. & Woodcock, C. L. Chromatin compaction by a polycomb group protein complex. Science 306, 1574–1577 (2004).
Endoh, M. et al. Histone H2A mono-ubiquitination is a crucial step to mediate PRC1-dependent repression of developmental genes to maintain ES cell identity. PLoS Genet. 8, e1002774 (2012).
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Ha, M. & Hong, S. Gene-regulatory interactions in embryonic stem cells represent cell-type specific gene regulatory programs. Nucleic Acids Res. 45, 10428–10435 (2017).
Khan, A. & Zhang, X. dbSUPER: a database of super-enhancers in mouse and human genome. Nucleic Acids Res. 44 D1, D164–D171 (2016).
Bultman, S. et al. A Brg1 null mutation in the mouse reveals functional differences among mammalian SWI/SNF complexes. Mol. Cell 6, 1287–1295 (2000).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Dekker, J., Marti-Renom, M. A. & Mirny, L. A. Exploring the three-dimensional organization of genomes: interpreting chromatin interaction data. Nat. Rev. Genet. 14, 390–403 (2013).
Brookes, E. et al. Polycomb associates genome-wide with a specific RNA polymerase II variant, and regulates metabolic genes in ESCs. Cell Stem Cell 10, 157–170 (2012).
Araya, C. L. et al. Identification of significantly mutated regions across cancer types highlights a rich landscape of functional molecular alterations. Nat. Genet. 48, 117–125 (2016).
Soufi, A., Donahue, G. & Zaret, K. S. Facilitators and impediments of the pluripotency reprogramming factors’ initial engagement with the genome. Cell 151, 994–1004 (2012).
Rahl, P. B. et al. c-Myc regulates transcriptional pause release. Cell 141, 432–445 (2010).
Lin, C. Y. et al. Transcriptional amplification in tumor cells with elevated c-Myc. Cell 151, 56–67 (2012).
Ho, L. et al. An embryonic stem cell chromatin remodeling complex, esBAF, is essential for embryonic stem cell self-renewal and pluripotency. Proc. Natl. Acad. Sci. USA 106, 5181–5186 (2009).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Pohl, A. & Beato, M. bwtool: a tool for bigWig files. Bioinformatics 30, 1618–1619 (2014).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Jin, W. et al. Genome-wide detection of DNase I hypersensitive sites in single cells and FFPE tissue samples. Nature 528, 142–146 (2015).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Kidder, B. L. & Zhao, K. Efficient library preparation for next-generation sequencing analysis of genome-wide epigenetic and transcriptional landscapes in embryonic stem cells. Methods Mol. Biol. 1150, 3–20 (2014).
Hahne, F. & Ivanek, R. Visualizing genomic data using Gviz and Bioconductor. Methods Mol. Biol. 1418, 335–351 (2016).
Anders, S., Pyl, P. T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Tibshirani, R. Regression shrinkage and selection via the lasso. J. R. Stat. Soc. B 58, 267–288 (1996).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Krieger, E. & Vriend, G. YASARA View - molecular graphics for all devices - from smartphones to workstations. Bioinformatics 30, 2981–2982 (2014).
Hooft, R. W., Vriend, G., Sander, C. & Abola, E. E. Errors in protein structures. Nature 381, 272 (1996).
Hooft, R. W., Sander, C., Scharf, M. & Vriend, G. The PDBFINDER database: a summary of PDB, DSSP and HSSP information with added value. Comput. Appl. Biosci. 12, 525–529 (1996).
Qiu, J. & Elber, R. SSALN: an alignment algorithm using structure-dependent substitution matrices and gap penalties learned from structurally aligned protein pairs. Proteins 62, 881–891 (2006).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Acknowledgements
We thank C. Weber, E. Chory, K. Cui, G. Hu, and E. Son for helpful discussions and dedicate this manuscript to the lasting memory of Joseph P. Calarco. We gratefully acknowledge the assistance of the DNA Sequencing and Genomics Core facility of NHLBI, the Stanford Cell Sciences Imaging Facility (S10OD01227601), and the Stanford BioX3 cluster (S10RR02664701). Ring1a −/− ;Ring1b fl/fl mESCs were a generous gift from H. Koseki (RIKEN, Japan). This work was supported by the Simons Foundation Autism Research Initiative (G.R.C.), NIH grants R37NS046789 (G.R.C.), R01CA163915 (G.R.C.), T32CA009151 (J.G.K.), R00CA187565 (H.C.H.), the Division of Intramural Research of the NHLBI/NIH (K.Z.)., the Czech Science Foundation grant 16-06357S (V.V.), the Cancer Prevention & Research Institute of Texas grant RR170036 (H.C.H.), and the Howard Hughes Medical Institute (G.R.C.).
Author information
Authors and Affiliations
Contributions
H.C.H. and B.Z.S. conceived of and performed experiments and wrote the paper. K.C., H.C.H., and V.V. developed the homology model. H.C.H, C.-Y.C., and E.L.M. performed analyses. B.Z.S., H.C.H., C.-Y.C., J.G.K., and W.L.K. prepared materials. K.Z. and G.R.C. designed experiments and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 The SMARCA4 ATP cleft and DNA groove are frequently mutated in cancer.
(a) Surface rendering of the SMARCA4 homology model viewed from the DNA-binding surface. Details of homology model are described in the Methods section. Surface residues are colored based on mutation frequency of deleterious missense mutations at the time of analysis in cBioPortal, for residue positions 758–1335. DNA is shown in yellow. (b) Rotation of the view in (a) 90° around the x-axis to show the ATP-binding cleft. ATP is shown in blue. (c) Rotation of the view in (a) 180° around the y-axis. (d) Rotation of the view in (c) –90° around the x-axis reveals few other surface mutation hotspots.
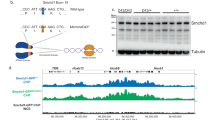
Supplementary Figure 2 Construction of a Smarca4-Dendra2 fluorescent knockin mouse.
(a) A 15.5-kbp homology template was used to target the last exon of SMARCA4 to create a SMARCA4-Dendra2 fusion. Thymidine kinase from herpes simplex virus (HSV-tk) was used for negative selection with gancyclovir, and positive selection from neo was achieved by selecting for G418-resistant colonies. (b) Southern blot of resulting heterozygous knockin mESC colony using the probe shown in panel (a). (c) Western blotting of the heterozygous knockin mESC colony shows a new band with higher molecular weight that also stains positive using anti-Dendra2 antibodies (Evrogen). (d) Fluorescent knockin SMARCA4 constructs assemble with endogenous BAF and PBAF subunits to form natural complexes. (e) Confocal image of resulting fluorescent mESC colony. (f) Homozygous mice obtained through breeding are phenotypically normal. Strain has been deposited for cryopreservation at Jackson Lab (stock #27901). PCR primers for genotyping are provided in the Methods section.
Supplementary Figure 3 SMARCA4-Dendra2 is highly stable and undergoes dynamic changes in interaction with chromatin throughout the cell cycle.
(a) Following photoconversion with 405-nm light using a custom-made LED lamp, SMARCA4-Dendra2 irreversibly photoswitches from green to red. (b) Photoswitching is readily detectable using flow cytometry. (c) The persistence of red fluorescence provides a measure of the stability of the protein fusion. Red fluorescence persists with a half-life of 11.8 ± 1.2 h, comparable to the canonical 12-hour half-life of mESCs. This result indicates that fluorescent fusions of SMARCA4 are stable. (d) Live-cell imaging of mitosis. Mitotic cells are readily distinguishable from interphasic cells due to the altered distribution of fluorescence intensity. (e) Fluorescence recovery after photobleaching (FRAP) of an individual mESC colony. (f) FRAP recovery times of SMARCA4-Dendra2 fusions are faster in mitosis, when the complex is excluded from condensed mitotic chromatin (g) Statistical comparison of FRAP recovery times of SMARCA4-Dendra2 fluorescent fusions. Mitotic cells show significantly increased FRAP recovery times (p<0.002, KS test). (h) Custom LED array for bulk photoconversion of cells using 405-nm LEDs (DigiKey 492-1349-ND).
Supplementary Figure 4 ATPase-dead SMARCA4 shows altered interaction with chromatin during interphase.
(a) Fluorescence deconvolution image of fixed mESCs expressing lentivirally transduced SMARCA4-GFP. During mitosis, SMARCA4 is excluded from chromatin. (b) Live-cell imaging of SMARCA4-GFP and DNA (imaged using DRAQ5) shows comparable results. (c) ATPase-dead K785R mutant (K798R in some citations), a cancer mutation found in the ATP-binding cleft) shows a dramatic dynamic defect, consistent with failure to release immobile chromatin. (d) Wild-type SMARCA4-GFP and SMARCA4-Dendra2 have indistinguishable dynamics in FRAP assays, while K785R mutants show a significant increase in the FRAP recovery times during interphase (p<2.2e-16). (e) During mitosis, mutant and wild-type complexes show similar dynamics, consistent with a model whereby ATP-dependent dynamics predominate during interphase when the complex engages chromatin (Figure 2g).
Supplementary Figure 5 Genome-wide alterations of ATAC-seq following acute Smarca4 knockout and relationship with Ring1b ChIP-seq.
(a) Heat maps and plot showing sites that lose accessibility following acute deletion of SMARCA4. (b) Heat maps and plot showing sites that do not change accessibility following acute deletion of SMARCA4. (c) Heat maps and plot showing sites that gain accessibility following acute deletion of SMARCA4. The criteria for classification is provided in the Methods section. (d) Genome-wide signal of ATAC-seq and Ring1b ChIP-seq, in relation to other chromatin features from the Mouse ENCODE Project. ATAC-seq is highly correlated with permissive marks, while Ring1b shows a high degree of correlation with H3K27me3, the mark of Polycomb Repressive Complex 2. (e) Enrichment of enhancers, transcription start sites (TSSs), transcription termination sites (TTSs), and intergenic regions in ATAC-seq datasets by class. ATAC-seq peaks are enriched at enhancers and TSSs. Small deviations between the knockout and WT/G784E expression reflect variability in genome-wide peak calling. (f) Enrichment of enhancers, transcription start sites (TSSs), transcription termination sites (TTSs), and intergenic regions in Ring1b ChIP-seq datasets by class. Ring1b peaks are enriched at TSSs.
Supplementary Figure 6 Analysis of altered chromatin signals and consistency of results across independent cell-culture replicates.
All fold changes below refer to changes that arise in G784E/WT mESCs compared to WT/WT mESCs. (a) MA plot of fold change of ATAC read density at individual sites across the genome. (b) Reproducibility of ATAC changes across independent cell-culture replicates. (c) MA plot of fold change of H3K4me1 read density at individual sites across the genome. (d) Reproducibility of H3K4me1 changes across independent cell-culture replicates. (e) MA plot of fold change of H3K27ac read density at individual sites across the genome. (f) Reproducibility of H3K27ac changes across independent cell-culture replicates. (g) MA plot of fold change of RNAP2 read density at individual sites across the genome. (h) Reproducibility of RNAP2 changes across independent cell-culture replicates.
Supplementary Figure 7 Examples of enhancer changes across the genome.
(a) Example of enhancer accessibility loss coinciding with H3K27ac loss but not H3K4me1 loss on chromosome 8. (b) Example of enhancer accessibility loss coinciding with H3K27ac loss but not H3K4me1 loss on chromosome 10. (c) Example of enhancer accessibility loss coinciding with H3K27ac loss but not H3K4me1 loss on chromosome 2. These changes are representative of the dominant-negative effects of SMARCA4 mutations on the enhancer landscape.
Supplementary Figure 8 Summary of ChIP-seq data at sites with increased and decreased accessibility measured by ATAC-seq.
(a) Summary of individual H3K4me1 ChIP-seq tracks at ATAC-seq sites identified as increased or decreased. (b) Summary of individual RNAP2 ChIP-seq tracks at ATAC-seq sites identified as increased or decreased. (c) Summary of individual H3K27ac ChIP-seq tracks at ATAC-seq sites identified as increased or decreased. (d) Mean RNA polymerase 2 (RNAP2) ChIP-seq read density at TSSs and enhancers in wild-type and heterozygous mutant cells.
Supplementary Figure 9 SMARCA2 (BRM) is repressed in wild-type and mutant SMARCA4 mESCs.
Expression of SMARCA2 compared to other genes in mutant and wild-type mESCs. Little to no SMARCA2 is expressed in mESCs, either in wild-type or mutant cells. Counting of transcripts reveals that SMARCA2 is expressed at less than 1% compared to SMARCA4, hence confounding effects due to SMARCA2 expression can be excluded. Plotted values are mean normalized expression counts (a.u.), error bars are 95% confidence intervals from independent cell-culture replicates (n=2).
Source data
Rights and permissions
About this article
Cite this article
Hodges, H.C., Stanton, B.Z., Cermakova, K. et al. Dominant-negative SMARCA4 mutants alter the accessibility landscape of tissue-unrestricted enhancers. Nat Struct Mol Biol 25, 61–72 (2018). https://doi.org/10.1038/s41594-017-0007-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-017-0007-3
This article is cited by
-
Polycomb repressive complex 2 and its core component EZH2: potential targeted therapeutic strategies for head and neck squamous cell carcinoma
Clinical Epigenetics (2024)
-
MYC overexpression and SMARCA4 loss cooperate to drive medulloblastoma formation in mice
Acta Neuropathologica Communications (2023)
-
The esBAF and ISWI nucleosome remodeling complexes influence occupancy of overlapping dinucleosomes and fragile nucleosomes in murine embryonic stem cells
BMC Genomics (2023)
-
SWI/SNF complexes in hematological malignancies: biological implications and therapeutic opportunities
Molecular Cancer (2023)
-
Mitotic bookmarking by SWI/SNF subunits
Nature (2023)