Abstract
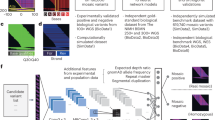
Structural variants (SVs) are a major driver of genetic diversity and disease in the human genome and their discovery is imperative to advances in precision medicine. Existing SV callers rely on hand-engineered features and heuristics to model SVs, which cannot scale to the vast diversity of SVs nor fully harness the information available in sequencing datasets. Here we propose an extensible deep-learning framework, Cue, to call and genotype SVs that can learn complex SV abstractions directly from the data. At a high level, Cue converts alignments to images that encode SV-informative signals and uses a stacked hourglass convolutional neural network to predict the type, genotype and genomic locus of the SVs captured in each image. We show that Cue outperforms the state of the art in the detection of several classes of SVs on synthetic and real short-read data and that it can be easily extended to other sequencing platforms, while achieving competitive performance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The 60× HG002 Illumina WGS short reads, the 28× HG002 PacBio CCS reads and the HG002 v.0.06 truthset are available through the GIAB FTP data repository. In particular, short reads can be downloaded from https://ftp-trace.ncbi.nlm.nih.gov/giab/ftp/data/AshkenazimTrio/HG002_NA24385_son/NIST_HiSeq_HG002_Homogeneity-10953946/NHGRI_Illumina300X_AJtrio_novoalign_bams/HG002.hs37d5.60x.1.bam, the PacBio CCS reads can be downloaded from https://ftp-trace.ncbi.nlm.nih.gov/giab/ftp/data/AshkenazimTrio/HG002_NA24385_son/PacBio_CCS_15kb/alignment/HG002.Sequel.15kb.pbmm2.hs37d5.whatshap.haplotag.RTG.10x.trio.bam and the v.0.06 truthset can be downloaded from https://ftp-trace.ncbi.nlm.nih.gov/giab/ftp/data/AshkenazimTrio/analysis/NIST_SVs_Integration_v0.6/HG002_SVs_Tier1_v0.6.vcf.gz. The CHM1 and CHM13 40× coverage Illumina WGS short reads can be downloaded from the ENA short-read archive (ENA accessions ERR1341794 and ERR1341795, respectively). The CHM1 and CHM13 PacBio long reads can be obtained from the NCBI sequence read archive under accession numbers SRP044331 (CHM1) and SRR11292120 to SRR11292123 (CHM13). The Huddleston et al.20 CHM1 and CHM13 truthsets can be downloaded from http://eichlerlab.gs.washington.edu/publications/Huddleston2016/structural_variants. To obtain a single truthset, we merged the CHM1 and CHM13 VCFs using SURVIVOR and genotyped the calls accordingly (such that records reported in both CHM1 and CHM13 were labeled as homozygous and records only reported in one of the two were labeled as heterozygous). To label duplications, we cross-referenced insertion calls with Supplementary Table 11 from previous work20, which separately reports which published insertion calls are duplications. The synthetic benchmark data, training data, trained models and configurations are available through the associated GitHub repository at https://github.com/PopicLab/cue.
Code availability
The Cue source code and documentation are available on GitHub under the MIT license at https://github.com/PopicLab/cue. The code is also archived in the Code Ocean capsule https://doi.org/10.24433/CO.8949236.v2.
References
Chaisson, M. J. et al. Multi-platform discovery of haplotype-resolved structural variation in human genomes. Nat. Commun. 10, 1–16 (2019).
Mantere, T., Kersten, S. & Hoischen, A. Long-read sequencing emerging in medical genetics. Front. Genet. 10, 426 (2019).
Li, Y. et al. Patterns of somatic structural variation in human cancer genomes. Nature 578, 112–121 (2020).
Chen, X. et al. Manta: rapid detection of structural variants and indels for germline and cancer sequencing applications. Bioinformatics 32, 1220–1222 (2016).
Rausch, T. et al. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 28, i333–i339 (2012).
Layer, R. M., Chiang, C., Quinlan, A. R. & Hall, I. M. LUMPY: a probabilistic framework for structural variant discovery. Genome Biol. 15, R84 (2014).
Wala, J. A. et al. SvABA: genome-wide detection of structural variants and indels by local assembly. Genome Res. https://doi.org/10.1101/gr.221028.117 (2018).
Sedlazeck, F. J. et al. Accurate detection of complex structural variations using single-molecule sequencing. Nat. Methods 15, 461–468 (2018).
Pacific Biosciences. pbsv. https://github.com/PacificBiosciences/pbsv (2018).
Alkan, C., Coe, B. P. & Eichler, E. E. Genome structural variation discovery and genotyping. Nat. Rev. Genet. 12, 363–376 (2011).
Poplin, R. et al. A universal SNP and small-indel variant caller using deep neural networks. Nat. Biotechnol. 36, 983–987 (2018).
Belyeu, J. R. et al. Samplot: a platform for structural variant visual validation and automated filtering. Genome Biol. 22, 1–13 (2021).
Bai, R., Ling, C., Cai, L. & Gao, J. Cnngeno: a high-precision deep-learning-based strategy for the calling of structural variation genotype. Comput. Biol. Chem. 94, 107417 (2021).
Liu, Y., Huang, Y., Wang, G. & Wang, Y. A deep learning approach for filtering structural variants in short-read sequencing data. Brief. Bioinform. 22, bbaa370 (2021).
Cai, L., Wu, Y. & Gao, J. DeepSV: accurate calling of genomic deletions from high-throughput sequencing data using deep convolutional neural network. BMC Bioinform. 20, 1–17 (2019).
Newell, A., Yang, K. & Deng, J. Stacked hourglass networks for human pose estimation. In Proc. European Conference on Computer Vision, 483–499 (Springer, 2016).
Newell, A., Huang, Z. & Deng, J. Associative embedding: end-to-end learning for joint detection and grouping. In Proc. Advances in Neural Information Processing Systems 30: Annual Conference on Neural Information Processing Systems 2017 (Guyon, I. et al.) (Curran Associates, Inc., 2017).
Zook, J. M. et al. A robust benchmark for detection of germline large deletions and insertions. Nat. Biotechnol. 38, 1347–1355 (2020).
Li, H. et al. A synthetic-diploid benchmark for accurate variant-calling evaluation. Nat. Methods 15, 595–597 (2018).
Huddleston, J. et al. Discovery and genotyping of structural variation from long-read haploid genome sequence data. Genome Res. 27, 677–685 (2017).
Li, J., Su, W. & Wang, Z. Simple pose: rethinking and improving a bottom-up approach for multi-person pose estimation. In Proc. AAAI Conference on Artificial Intelligence, Vol. 34, 11354–11361 (AAAI, 2020).
Jeffares, D. C. et al. Transient structural variations have strong effects on quantitative traits and reproductive isolation in fission yeast. Nat. Commun. 8, 1–11 (2017).
English, A. C., Menon, V. K., Gibbs, R., Metcalf, G. A. & Sedlazeck, F. J. Truvari: refined structural variant comparison preserves allelic diversity. Genome Biol. https://doi.org/10.1186/s13059-022-02840-6 (2022).
Tarailo-Graovac, M. & Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr. Protoc. Bioinformatics 25, 4–10 (2009).
Karolchik, D. et al. The UCSC Genome Browser database. Nucleic Acids Res. 31, 51–54 (2003).
Zhao, X. et al. Expectations and blind spots for structural variation detection from long-read assemblies and short-read genome sequencing technologies. Am. J. Hum. Genet. https://doi.org/10.1016/j.ajhg.2021.03.014 (2021).
Collins, R. L. et al. A structural variation reference for medical and population genetics. Nature 581, 444–451 (2020).
Ono, Y., Asai, K. & Hamada, M. PBSIM2: a simulator for long-read sequencers with a novel generative model of quality scores. Bioinformatics 37, 589–595 (2021).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Luo, R., Sedlazeck, F. J., Darby, C. A., Kelly, S. M. & Schatz, M. C. LRSim: a linked-reads simulator generating insights for better genome partitioning. Comput. Struct. Biotechnol. J. 15, 478–484 (2017).
Marks, P. et al. Resolving the full spectrum of human genome variation using linked-reads. Genome Res. 29, 635–645 (2019).
Fang, L. et al. LinkedSV for detection of mosaic structural variants from linked-read exome and genome sequencing data. Nat. Commun. 10, 1–15 (2019).
Ebert, P. et al. Haplotype-resolved diverse human genomes and integrated analysis of structural variation. Science https://doi.org/10.1126/science.abf7117 (2021).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
DWGSIM. Whole genome simulator for next-generation sequencing https://github.com/nh13/DWGSIM (2022).
Heng Li. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at arXiv https://doi.org/10.48550/arXiv.1303.3997 (2013).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. Preprint at arXiv https://doi.org/10.48550/arXiv.1412.6980 (2014).
Acknowledgements
Research reported in this publication was supported by the Broad Institute Schmidt Fellowship and the National Human Genome Research Institute of the National Institutes of Health Award R01HG012467 to V.P. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. I.H. was also supported by the National Institute of General Medical Sciences Maximizing Investigators’ Research Award R35GM138152. We thank H. Brand, M. Talkowski, A. Al’Khafaji and members of their laboratories at the Broad Institute for useful feedback and discussions. We thank the Genomics Platform at the Broad Institute and the SCU at Weill Cornell Medicine for access to GPU computing resources. We also thank A. Kushlak for the data recovery service provided during this project.
Author information
Authors and Affiliations
Contributions
V.P. conceived the study. V.P. implemented the framework, generated training data, trained the models and performed the evaluation across benchmarks. C.R. implemented scripts to annotate and visualize SV callsets and assisted with analysis. F.C. performed runtime benchmarking, interval selection experiments and evaluated SV candidate calls using long reads. V.P. and I.H. selected datasets for the benchmarks. I.H. provided access to GPU resources. D.M. produced call sets of existing tools on several benchmark datasets. K.G. assisted with the interpretation of candidate SV calls. A.M. reviewed the methodology of existing approaches and assisted with analysis. V.P. wrote the paper. All of the authors revised the paper. V.P. supervised the study.
Corresponding author
Ethics declarations
Competing interests
V.P. is a former employee and owns shares of Illumina. Illumina produces sequencing platforms that generate short-read data that were used in this work for SV detection. All other authors have no competing interests.
Peer review
Peer review information
Nature Methods thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Lin Tang, in collaboration with the Nature Methods team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Performance evaluation broken down by SV type on synthetic data at 30x genome coverage.
a. Precision, recall and F1 score for DEL, DUP and INV calling and genotyping. b. Recall-precision curves for each SV type generated using the SV quality thresholds reported in the QUAL VCF field.
Extended Data Fig. 2 Performance evaluation on synthetic data at varying genome coverage.
Precision, recall, and F1 score for DEL, DUP, and INV calling and genotyping computed for chr1 at 10x, 15x, 30x, 45x, and 60x genome coverage. Results are shown for all the SV calls combined (‘ALL’) and broken down by type.
Extended Data Fig. 3 Evaluation of the TP, FN, and FP SV calls in the HG002 benchmark.
a. Histogram showing the number of occurrences of the TP and FN SV calls in gnomAD-SV for each tool. SVs with no match in gnomAD-SV are collected in the zeroth bin. b. TP and FN calls broken down by frequency in gnomAD-SV and genome context. c. Recall-Precision curves generated using the SV quality thresholds reported in the QUAL VCF field. d. The Recall-Precision curve of Cue annotated with a subset of reported SV quality values.
Extended Data Fig. 4 Analysis of a false positive HG002 deletion generated by all short-read callers except Cue.
a. IGV plot showing short-read alignments at the call locus. Discordant read pairs mapped to the same strand (LL and RR mappings) are shown in light and dark blue, RL mappings are shown in green, and read pairs with a discordantly large insert size are shown in red. b. Cue-generated image channels depicting short-read signals that are inconsistent with a valid DEL signature. c. One of the two haplotypes of HG002, reconstructed by de novo assembly of PacBio CCS reads, that explains the main discordant pair mappings in panel a (the other haplotype is identical to the reference). The reconstructed haplotype contains two dispersed DUPs, one inverted dispersed DUP, and no DEL. Colored blocks labeled with letters are distinct short repeats. Gray blocks broken by diagonal lines are long sequences. rc(A) denotes the reverse-complement of A. Haplotypes were reconstructed and compared to the reference as follows. Let W be the sequence of the reference that covers the main patterns of discordant pairs in panel a. We built a joint de Bruijn graph (k=87) on W and on the 190 CCS reads that have some alignment to W, we removed k-mers with frequency one, and we translated W and every read into a walk (which may contain cycles) in the graph.
Extended Data Fig. 5 Schematic of read-pair mapping signatures for a small dispersed DUP and a divergent reference repeat.
Locus ‘A’ is duplicated in the donor genome. Some read pairs map discordantly in the RL orientation (green) or with a large insert size (red). Pairs internal to each copy of the donor map to the single copy of ‘A’ in the reference genome, doubling its coverage. If the reference has a divergent copy of ‘A’ (denoted as ‘a’), a gap in coverage will be observed at ‘a’.
Extended Data Fig. 6 Evaluation of DUP and INV calls in the CHM1 and CHM13 diploid mix benchmark.
a. Upset plot depicting DUP callset overlaps of short-read and long-read callers (only sets larger than 5 events are displayed for conciseness). Overlaps that include Cue are highlighted in orange. b. Breakdown of DUP calls by consensus with long-read and other short-read callers. c. Upset plot depicting INV callset overlaps of short-read and long-read callers. d. Breakdown of INV calls by consensus with long-read and other short-read callers.
Extended Data Fig. 7 Training data generation.
a. High-level overview of the in silico sequencing and image data generation process. b. Annotated training examples (displayed using standard image visualization software using only three Cue channels, including the read-depth channel).
Extended Data Fig. 8 Runtime and memory performance on chr1 of HG002.
a. Sequential runtime. Cue’s runtime is divided into indexing and calling. Lumpy’s runtime is divided into indexing, calling (short block), and genotyping. b. Sequential peak memory. c. Effect of PyTorch parallelism on calling time of Cue. In ‘multi-CPU’ mode we do not limit PyTorch to use a specific number of threads.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2, Supplementary Table 1 and Supplementary Notes 1–5.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Popic, V., Rohlicek, C., Cunial, F. et al. Cue: a deep-learning framework for structural variant discovery and genotyping. Nat Methods 20, 559–568 (2023). https://doi.org/10.1038/s41592-023-01799-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-023-01799-x
This article is cited by
-
De novo and somatic structural variant discovery with SVision-pro
Nature Biotechnology (2024)
-
Facilitating genome structural variation analysis
Nature Methods (2023)
-
A survey of algorithms for the detection of genomic structural variants from long-read sequencing data
Nature Methods (2023)