Abstract
An exciting frontier in circuit neuroscience lies at the intersection between neural network mapping and single-cell genomics. Monosynaptic rabies viruses provide a promising platform for the merger of circuit mapping methods with -omics approaches. However, three key limitations have hindered the extraction of physiologically meaningful gene expression profiles from rabies-mapped circuits: inherent viral cytotoxicity, high viral immunogenicity and virus-induced alteration of cellular transcriptional regulation. These factors alter the transcriptional and translational profiles of infected neurons and their neighboring cells. To overcome these limitations we applied a self-inactivating genomic modification to the less immunogenic rabies strain, CVS-N2c, to generate a self-inactivating CVS-N2c rabies virus (SiR-N2c). SiR-N2c not only eliminates undesired cytotoxic effects but also substantially reduces gene expression alterations in infected neurons and dampens the recruitment of innate and acquired immune responses, thus enabling open-ended interventions on neural networks and their genetic characterization using single-cell genomic approaches.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All materials described in this paper can be obtained for non-commercial purposes after signing a material transfer agreement (MTA) with the UKRI. Data generated during this study are included in the manuscript and supporting files. The DNA constructs to generate the viral vectors used in this study are available from Addgene (99610, 194352–194356). The raw NGS dataset has been deposited into NCBI’s Sequence Read Archive (SRA) and is accessible through accession number PRJNA901288. Further requests for information, resources and reagents should be directed to the Lead Contact, Marco Tripodi.
References
McCulloch, W. S. & Pitts, W. A logical calculus of the ideas immanent in nervous activity. 1943. Bull. Math. Biol. 52, 99–115 (1990). discussion 73–97.
Sherrington, C. S. The Integrative Action of the Nervous System (Cambridge University Press, 1947).
Li, P. H. et al. Automated reconstruction of a serial-section EM Drosophila brain with flood-filling networks and local realignment. Microsc. Microanal. 25 (Suppl. 2), 1364–1365 (2019).
White, J. G., Southgate, E., Thomson, J. N. & Brenner, S. The structure of the nervous system of the nematode Caenorhabditis elegans. Philos. Trans. R. Soc. Lond. B Biol. Sci. 314, 1–340 (1986).
Luo, L., Callaway, E. M. & Svoboda, K. Genetic dissection of neural circuits: a decade of progress. Neuron 98, 865 (2018).
Iourov, I. Y., Vorsanova, S. G. & Yurov, Y. B. Single cell genomics of the brain: focus on neuronal diversity and neuropsychiatric diseases. Curr. Genomics 13, 477–488 (2012).
Rosenberg, A. B. et al. Single-cell profiling of the developing mouse brain and spinal cord with split-pool barcoding. Science 360, 176–182 (2018).
Wickersham, I. R. et al. Monosynaptic restriction of transsynaptic tracing from single, genetically targeted neurons. Neuron 53, 639–647 (2007).
Wickersham, I. R., Finke, S., Conzelmann, K. K. & Callaway, E. M. Retrograde neuronal tracing with a deletion-mutant rabies virus. Nat. Methods 4, 47–49 (2007).
Stepien, A. E., Tripodi, M. & Arber, S. Monosynaptic rabies virus reveals premotor network organization and synaptic specificity of cholinergic partition cells. Neuron 68, 456–472 (2010).
Tripodi, M., Stepien, A. E. & Arber, S. Motor antagonism exposed by spatial segregation and timing of neurogenesis. Nature 479, 61–66 (2011).
Reardon, T. R. et al. Rabies virus CVS-N2c(DeltaG) strain enhances retrograde synaptic transfer and neuronal viability. Neuron 89, 711–724 (2016).
Huang, K. W. & Sabatini, B. L. Single-cell analysis of neuroinflammatory responses following intracranial injection of G-deleted rabies viruses. Front Cell Neurosci. 14, 65 (2020).
Zhao, P. et al. Analysis of expression profiles of long noncoding RNAs and mRNAs in brains of mice infected by rabies virus by RNA sequencing. Sci. Rep. 8, 11858 (2018).
Prosniak, M., Hooper, D. C., Dietzschold, B. & Koprowski, H. Effect of rabies virus infection on gene expression in mouse brain. Proc. Natl Acad. Sci. USA 98, 2758–2763 (2001).
Ciabatti, E., Gonzalez-Rueda, A., Mariotti, L., Morgese, F. & Tripodi, M. Life-long genetic and functional access to neural circuits using self-inactivating rabies virus. Cell 170, 382–392 (2017).
Chatterjee, S. et al. Nontoxic, double-deletion-mutant rabies viral vectors for retrograde targeting of projection neurons. Nat. Neurosci. 21, 638–646 (2018).
Zhang, D. et al. Genome-wide transcriptional profiling reveals two distinct outcomes in central nervous system infections of rabies virus. Front. Microbiol. 7, 751 (2016).
Miao, F. M. et al. Comparison of immune responses to attenuated rabies virus and street virus in mouse brain. Arch. Virol. 162, 247–257 (2017).
Wang, Z. W. et al. Attenuated rabies virus activates, while pathogenic rabies virus evades, the host innate immune responses in the central nervous system. J. Virol. 79, 12554–12565 (2005).
Dietzschold, B., Li, J., Faber, M. & Schnell, M. Concepts in the pathogenesis of rabies. Future Virol. 3, 481–490 (2008).
Conzelmann, K. K., Cox, J. H., Schneider, L. G. & Thiel, H. J. Molecular cloning and complete nucleotide sequence of the attenuated rabies virus SAD B19. Virology 175, 485–499 (1990).
Ciabatti, E. et al. Genomic stability of self-inactivating rabies. Preprint at bioRxiv https://doi.org/10.1101/2020.09.19.304683 (2020).
Menegas, W., Akiti, K., Amo, R., Uchida, N. & Watabe-Uchida, M. Dopamine neurons projecting to the posterior striatum reinforce avoidance of threatening stimuli. Nat. Neurosci. 21, 1421–1430 (2018).
Jin, L. et al. “Self-inactivating” rabies viruses are susceptible to loss of their intended attenuating modification. Preprint at bioRxiv https://doi.org/10.1101/550640 (2022).
Takeuchi, O. & Akira, S. MDA5/RIG-I and virus recognition. Curr. Opin. Immunol. 20, 17–22 (2008).
Hooper, D. C. et al. Collaboration of antibody and inflammation in clearance of rabies virus from the central nervous system. J. Virol. 72, 3711–3719 (1998).
Eddleston, M. & Mucke, L. Molecular profile of reactive astrocytes: implications for their role in neurologic disease. Neuroscience 54, 15–36 (1993).
Sasaki, Y., Ohsawa, K., Kanazawa, H., Kohsaka, S. & Imai, Y. Iba1 is an actin-cross-linking protein in macrophages/microglia. Biochem. Biophys. Res. Commun. 286, 292–297 (2001).
Morimoto, K. et al. Rabies virus quasispecies: implications for pathogenesis. Proc. Natl Acad. Sci. USA 95, 3152–3156 (1998).
Kim, E. J., Jacobs, M. W., Ito-Cole, T. & Callaway, E. M. Improved monosynaptic neural circuit tracing using engineered rabies virus glycoproteins. Cell Rep. 15, 692–699 (2016).
Russo, S. J. & Nestler, E. J. The brain reward circuitry in mood disorders. Nat. Rev. Neurosci. 14, 609–625 (2013).
Tasic, B. et al. Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat. Neurosci. 19, 335–346 (2016).
Zeisel, A. et al. Brain structure. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Kohl, J. et al. Functional circuit architecture underlying parental behaviour. Nature 556, 326–331 (2018).
Kim, E. J. et al. Extraction of distinct neuronal cell types from within a genetically continuous population. Neuron 107, 274–282 (2020).
Alkaslasi, M. R. et al. Single nucleus RNA-sequencing defines unexpected diversity of cholinergic neuron types in the adult mouse spinal cord. Nat. Commun. 12, 2471 (2021).
Blum, J. A. et al. Single-cell transcriptomic analysis of the adult mouse spinal cord reveals molecular diversity of autonomic and skeletal motor neurons. Nat. Neurosci. 24, 572–583 (2021).
Sathyamurthy, A. et al. Massively parallel single nucleus transcriptional profiling defines spinal cord neurons and their activity during behavior. Cell Rep. 22, 2216–2225 (2018).
Mo, A. et al. Epigenomic signatures of neuronal diversity in the mammalian brain. Neuron 86, 1369–1384 (2015).
Russ, D. E. et al. A harmonized atlas of mouse spinal cord cell types and their spatial organization. Nat. Commun. 12, 5722 (2021).
Levine, A. J. et al. Identification of a cellular node for motor control pathways. Nat. Neurosci. 17, 586–593 (2014).
Arber, S. Motor circuits in action: specification, connectivity, and function. Neuron 74, 975–989 (2012).
Lake, B. B. et al. Neuronal subtypes and diversity revealed by single-nucleus RNA sequencing of the human brain. Science 352, 1586–1590 (2016).
Tiklova, K. et al. Single-cell RNA sequencing reveals midbrain dopamine neuron diversity emerging during mouse brain development. Nat. Commun. 10, 581 (2019).
Gradinaru, V. et al. Targeting and readout strategies for fast optical neural control in vitro and in vivo. J. Neurosci. 27, 14231–14238 (2007).
Courtine, G. & Sofroniew, M. V. Spinal cord repair: advances in biology and technology. Nat. Med. 25, 898–908 (2019).
Takeoka, A., Vollenweider, I., Courtine, G. & Arber, S. Muscle spindle feedback directs locomotor recovery and circuit reorganization after spinal cord injury. Cell 159, 1626–1639 (2014).
Roselli, F. & Caroni, P. From intrinsic firing properties to selective neuronal vulnerability in neurodegenerative diseases. Neuron 85, 901–910 (2015).
Osakada, F. & Callaway, E. M. Design and generation of recombinant rabies virus vectors. Nat. Protoc. 8, 1583–1601 (2013).
Wickersham, I. R., Sullivan, H. A. & Seung, H. S. Production of glycoprotein-deleted rabies viruses for monosynaptic tracing and high-level gene expression in neurons. Nat. Protoc. 5, 595–606 (2010).
Mi, H. et al. Protocol update for large-scale genome and gene function analysis with the PANTHER classification system (v.14.0). Nat. Protoc. 14, 703–721 (2019).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics 10, 48 (2009).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Garrison, E. & Marth, G. Haplotype-based variant detection from short-read sequencing. Preprint at arXiv https://doi.org/10.48550/arXiv.1207.3907 (2012).
Nagy, C. et al. Single-nucleus transcriptomics of the prefrontal cortex in major depressive disorder implicates oligodendrocyte precursor cells and excitatory neurons. Nat. Neurosci. 23, 771–781 (2020).
Raudvere, U. et al. g:Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 47, W191–W198 (2019).
Acknowledgements
The authors thank the creators of the SeqMonk program (Babraham Institute) and the SeqSeek atlas41, which were used in the analysis pipelines. The authors also thank the members of the LMB Biological Service Group for their support with the in vivo work. This study was supported by the Medical Research Council (MC_UP_1201/2) to M.T., the European Research Council (STG 677029) to M.T., the Rosetrees Trust with an MBPhD fellowship to H.L. (M598) and by the Cambridge Philosophical Society and St Edmund’s College (University of Cambridge) with the Henslow Research Fellowship to A.G.-R.
Author information
Authors and Affiliations
Contributions
M.T., H.L. and E.C. conceived the project and analyzed the data; H.L. and E.C. conducted the experiments and wrote the manuscript with input from M.T.; E.C. and H.L. co-developed the SiR-N2c technology; E.C. produced SiR viruses with the help of H.L. and F.M.; H.L. led the in vivo characterization of the virus and performed the immunology and bulk RNA-seq experiments; H.L. performed the long-term survival experiments and peripheral neurotropism experiments with the help of E.C.; E.C. performed the pseudotyped rabies tracing experiment; E.C., E.W. and F.N. performed the single-cell genomic experiments; H.L. and S.M. analyzed the bulk and single-cell genomic data; A.G.-R. performed the electrophysiological recordings.
Corresponding authors
Ethics declarations
Competing interests
E.C. and M.T. are inventors of a patent related to the SiR technology (WO2018203049A1). All other authors have no competing interests.
Peer review
Peer review information
Nature Methods thanks Naoshige Uchida and the other, anonymous, reviewers for their contribution to the peer review of this work. Primary Handling Editor: Nina Vogt, in collaboration with the Nature Methods team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Sequencing of SiR viral productions confirms viral productions maintain the self- inactivating TEVs-PEST sequence.
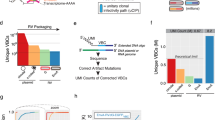
a) Scheme of the procedure to sequence individual viral particles from independent SiR preparations: ssRNA genomes from a viral production are extracted and treated with DNAse I. The RNA is retrotranscribed and the entire N-TEVs-PEST sequence and portion of the P gene is PCR amplified. The amplicon is then inserted into a pBluescript KS(+) sequencing vector to create genomic libraries. (b) List of detected mutations divided by batch (47 individual clones per batch). The position of the mutations is calculated referring to +1 as the first base of the nucleoprotein N coding sequence. (c) The CA1 of mice were injected with SiR-N2c virus or PBS control. 1 week p.i. the hippocampi were dissected for total RNA-seq sample preparation. Sequenced reads were aligned to the SiR-N2c viral genome using Burrows-Wheeler Aligner (BWA)54. Aligned reads were then screened for mutations using freebayes variant detector55. (d) Visualization of reads from SiR- N2c-injected samples aligned to the SiR-N2c genome using Seqmonk software. Samples from PBS-injected samples show only few chance alignments in comparison. (e) Zoom in to TEVs-PEST region of the genome (f) Visualization of mutagenic events identified across SiR-N2c genome flanking the TEVs-PEST coding region (yellow) from a total of 3 injected samples.
Extended Data Fig. 2 Rabies viral infection induces innate immune responses in vitro.
a) Immunostaining of neuro2A cells infected with ΔG−Rabies-B19-GFP shows induction of NF-kB (left) and p38 MAPK (right) activation 24 h post-infection. Treatment with PBS or 100 ng TNFα were used as negative and positive controls, respectively, for induction of innate interferon responses. (b) RT–qPCR fold changes of immune response genes IL6 and RIG-I in neuro2A following ΔG-Rabies-B19, ΔG-Rabies-N2c, SiR-B19, SiR-N2c infection. Fold changes are normalized to SiR-N2c induced transcript levels. (mean ± SEM, n = 3, one-way ANOVA with Tukey correction for multiple comparisons; top-down, left to right: P = ****5.47e-5; **1.2e-3; *1.3e-2; *2.7e-2. Scale bars 100 μm for NF-kB images, 50 μm for MAPK images.
Extended Data Fig. 3 ΔG-Rabies-N2c infection induces an extensive immune response and perturbs synaptic function in the mouse hippocampus.
(a) Scatter plot depicting differentially regulated transcripts in ΔG-Rabies-N2c-injected vs. PBS-injected control mice (blue). Gene ontology (GO) terms enriched in up- and downregulated transcripts following ΔG-Rabies-N2c injection compared to PBS-injected controls organized by biological process (b), reactome pathway (c) and molecular function (d) (Fisher’s exact test with Benjamini and Hochberg multiple testing correction (n = 3 per condition).
Extended Data Fig. 4 Hierarchy of GO terms of ‘immune system process’ enriched in upregulated RNA transcripts from ΔG-Rabies-N2c virus-infected hippocampi.
Enrichment analysis using GOrilla web-based tool (exact mHG p-value computation with p-value threshold 10−3) (ref. 53).
Extended Data Fig. 5 SiR-N2c displays increased peripheral neurotropism and retains transsynaptic spreading capabilities.
(a) CRE-reporter pups (P3-P5) were injected with either SiR-N2c-CRE or oG-coated SiR-B19-CRE in the quadriceps muscle area to target motor neuron terminals. (b) Quantification of total motor neurons infected post-injection, normalized by relative titer of the two viruses. (mean ± SEM, n = 3, two-tailed, unpaired Student’s T-test; P = ***0.0002) (c) Scheme of the strategy for transsynaptic labeling of premotor neurons. tdTomato reporter pups (P3-P5) were injected with a mixture of SiR-N2c-CRE and AAV-G-nucFLAG in the quadriceps muscle area to target motor neuron terminals. (d) Representative images of the spinal cord of mice injected with SiR-N2c-CRE in absence of AAV-G-nucFLAG displaying Chat+ motor neurons (cyan) labeled with tdTomato and FLAG (n = 3) (e) Zoomed images of tdTomato+ motor neurons. (f) Representative images of tdTomato+ SiR-N2c-CRE infected neurons in the spinal cord of injected mice at 1 week p.i. stained for Chat (cyan) and FLAG (gray) (n = 3). (g) Zoomed images of tdTomato +/FLAG+ starter motor neurons. Scale bars: 200 µm for entire spinal cord, 50 µm for zoomed images.
Extended Data Fig. 6 SiR-N2c infection induces lower glial immune response in the spinal cord compared to ΔG-Rabies-N2c.
(a) tdTomato CRE-reporter pups were injected in the quadriceps muscle area with a mixture of either ΔG-Rabies- N2c-CRE or SiR-N2c-CRE and AAV-G-nucFLAG and killed up to 2 weeks p.i. for immunofluorescence studies (b–e). Comparison of GFAP+ staining for activated astrocytes in spinal cords from pups injected with ΔG-Rabies- N2c (B) or SiR-N2c (C), with close up panel showing regions with tdTomato+ neurons. Comparison of Iba1+ staining for activated microglia in spinal cords from pups injected with ΔG-Rabies-N2c (D) or SiR-N2c (E), with close up panel showing regions with tdTomato+ neurons. Scale bars 200 μm for entire spinal cord images and 50 μm for zoomed images.
Extended Data Fig. 7 SiR-N2c EnvA-pseudotyped specificity.
(a) CRE-reporter mice were either not injected or injected in the NAc with AAV-TVA or AAV-TVA-G. 3 weeks later NAc was injected with SiR-N2c-CRE (EnvA). (b-d) Representative images of whole brains of Rosa-LoxPSTOPLoxP-tdTomato mice. (e–g) Representative images of the NAc of injected (nuclear GFP in F and TVA-GFP fusion in G). (h-j) Zoomed images of starter neurons in the NAc. Scale bars: 1 mm for whole brains (B-D), 100 μm for NAc images (E–G), 15 μm for zoomed panels (H-J).
Extended Data Fig. 8 SiR-N2c traced GFP+ neurons show enrichment for specific neuronal subtypes within the spinal premotor circuit.
A stacked tile plot where the horizontal length of rectangles on the x axis represents the relative frequency of neuron subtypes in SiR-N2c traced GFP+ neurons versus non-virally traced GFP- control spinal cord neurons. An oval represents 0% frequency. Each rectangle and circle is colored to represent the Pearson residual from a two-sided chi-squared test. X is placed adjacent to neuronal subtypes that were not identified in ΔG-Rabies-N2c-infected GFP+ neurons.
Extended Data Fig. 9 SiR-N2c-infected neurons show less disruption of transcriptional regulation compared to ΔG-Rabies-N2c-infected neurons.
(a) Enrichment of Biological Process Gene Ontology terms related to neuronal structure, function and RNA synthesis in differentially downregulated genes (gene set enrichment analysis, cumulative hypergeometric test with g:Set Counts and Sizes correction method)57, in ΔG-Rabies-N2c-infected neurons compared to non-infected neurons (blue) and in SiR-N2c-infected neurons compared to non-infected neurons (orange, p > 0.05). (b) Enrichment of Cell Component Gene Ontology terms related to neuronal structures in differentially downregulated genes in ΔG-Rabies-N2c-infected neurons compared to non-infected neurons (blue)and in SiR-N2c-infected neurons compared to non-infected neurons (orange). Gene set enrichment analysis using cumulative hypergeometric test with g:Set Counts and Sizes correction method57. (c) Gene ontology terms enriched in genes upregulated in SiR-N2c-infected neurons compared to ΔG-Rabies-N2c-infected neurons (blue) or upregulated in ΔG-Rabies-N2c-infected neurons compared to SiR-N2c-infected neurons (red). Enrichment analysis using GOrilla web-based tool, exact mHG p-value computation with p-value threshold 10−3 53. (d-g) Gene expression levels of canonical neuronal genes (Kirrel3, Grik3, Igf1r) and vesicle membrane protein Sec61a1 in uninfected neurons (green), SiR-N2c-infected neurons (pink), and ΔG-Rabies-N2c-infected neurons (blue). (mean ± SEM, differential expression analyses, two-sided chi-squared test of differential proportion genes; n = 224 for GFP-, 628 for SiR GFP+, and 580 for WT GFP+. Each experimental condition consists of neurons from 3 biologically independent samples). From left to right: P = ****3.2e-11, ****1.6e-06, ****4.1e-07, ****3.7e-09.
Supplementary information
Supplementary Information
Supplementary Tables 1–3.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lee, H., Ciabatti, E., González-Rueda, A. et al. Combining long-term circuit mapping and network transcriptomics with SiR-N2c. Nat Methods 20, 580–589 (2023). https://doi.org/10.1038/s41592-023-01787-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-023-01787-1