Abstract
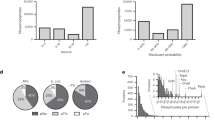
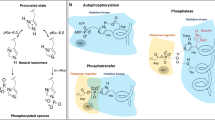
It has been suggested that in mammalian cells histidine residues in proteins may become as frequently phosphorylated as serine, threonine and tyrosine, and may play a key role in mammalian signaling. Here we applied a robust workflow that earlier allowed us to detect histidine phosphorylation in bacteria unambiguously, to probe for histidine phosphorylation in four human cell lines. Initially, seemingly hundreds of protein histidine phosphorylations were picked up in all studied human cell lines. However, careful examination of the data, and several control experiments, led us to the conclusion that >99% of these initially assigned pHis sites were not genuine, and should be site localized to neighboring Ser/Thr residues. Nevertheless, our methods are selective enough to detect just a handful of genuine pHis sites in mammalian cells, representing well-known enzymatic intermediates. Consequently, we do not find any evidence in our data supporting that protein histidine phosphorylation plays a role in mammalian signaling.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Data availability
All raw data that support the findings of this study have been deposited in ProteomeXchange with the accession number PXD028360.
References
Fuhs, S. R. & Hunter, T. pHisphorylation: the emergence of histidine phosphorylation as a reversible regulatory modification. Curr. Opin. Cell Biol. 45, 8–16 (2017).
Adam, K. & Hunter, T. Histidine kinases and the missing phosphoproteome from prokaryotes to eukaryotes. Lab. Invest. 98, 233–247 (2018).
Adam, K., Ning, J., Reina, J. & Hunter, T. NME/NM23/NDPK and histidine phosphorylation. Int. J. Mol. Sci. 21, 5848 (2020).
Hindupur, S. K. et al. The protein histidine phosphatase LHPP is a tumour suppressor. Nature 555, 678–682 (2018).
Kee, J. M., Oslund, R. C., Perlman, D. H. & Muir, T. W. A pan-specific antibody for direct detection of protein histidine phosphorylation. Nat. Chem. Biol. 9, 416–421 (2013).
Fuhs, S. R. et al. Monoclonal 1- and 3-phosphohistidine antibodies: new tools to study histidine phosphorylation. Cell 162, 198–210 (2015).
Hardman, G. et al. Strong anion exchange‐mediated phosphoproteomics reveals extensive human non‐canonical phosphorylation. EMBO J. 38, e100847 (2019).
Potel, C. M., Lin, M. H., Heck, A. J. R. & Lemeer, S. Widespread bacterial protein histidine phosphorylation revealed by mass spectrometry-based proteomics. Nat. Methods 15, 187–190 (2018).
Potel, C. M. et al. Gaining confidence in the elusive histidine phosphoproteome. Anal. Chem. 91, 5542–5547 (2019).
Hardman, G. & Eyers, C. E. High-throughput characterization of histidine phosphorylation sites using UPAX and tandem mass. Methods Mol Biol. 2077, 225–235 (2020).
Xue, L. et al. Sensitive kinase assay linked with phosphoproteomics for identifying direct kinase substrates. Proc. Natl Acad. Sci. USA 109, 5615–5620 (2012).
Potel, C. M., Fasci, D. & Heck, A. J. R. Mix and match of the tumor metastasis suppressor Nm23 protein isoforms in vitro and in vivo. FEBS J. 285, 2856–2868 (2018).
Acknowledgements
We thank B. van Breukelen for assistance with preparing the figures. This research received funding through the Netherlands Organization for Scientific Research through a VIDI grant (project no. 723.013.008) to S.L. and the Spinoza Award SPI.2017.028 to A.J.R.H. This project received additional funding from the European Union’s Horizon 2020 research and innovation program under grant agreement no. 686547 (EPIC-XS).
Author information
Authors and Affiliations
Contributions
N.M.L., A.J.R.H. and S.L. designed the experiments. N.M.L. performed the experiments. N.M.L. and S.L. performed data analysis. N.M.L., A.J.R.H. and S.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks the anonymous reviewers for their contribution to the peer review of this work. Arunima Singh was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Notes 1–6 and Supplementary Figs. 1–7
Rights and permissions
About this article
Cite this article
Leijten, N.M., Heck, A.J.R. & Lemeer, S. Histidine phosphorylation in human cells; a needle or phantom in the haystack?. Nat Methods 19, 827–828 (2022). https://doi.org/10.1038/s41592-022-01524-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-022-01524-0