Abstract
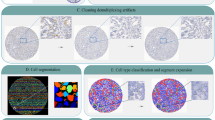
Advances in multiplexed in situ imaging are revealing important insights in spatial biology. However, cell type identification remains a major challenge in imaging analysis, with most existing methods involving substantial manual assessment and subjective decisions for thousands of cells. We developed an unsupervised machine learning algorithm, CELESTA, which identifies the cell type of each cell, individually, using the cell’s marker expression profile and, when needed, its spatial information. We demonstrate the performance of CELESTA on multiplexed immunofluorescence images of colorectal cancer and head and neck squamous cell carcinoma (HNSCC). Using the cell types identified by CELESTA, we identify tissue architecture associated with lymph node metastasis in HNSCC, and validate our findings in an independent cohort. By coupling our spatial analysis with single-cell RNA-sequencing data on proximal sections of the same specimens, we identify cell–cell crosstalk associated with lymph node metastasis, demonstrating the power of CELESTA to facilitate identification of clinically relevant interactions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The scRNA-seq data are deposited at GEO: GSE140042. HNSCC imaging data are hosted at Synapse.org SageBionetworks at https://doi.org/10.7303/syn26242593. The benchmark public imaging data can be found at https://doi.org/10.7937/tcia.2020.fqn0-0326. Source data are provided with this paper.
Code availability
All codes related to CELESTA can be found at https://github.com/plevritis/CELESTA. The source codes are also hosted at Code Ocean at https://doi.org/10.24433/CO.0677810.v1 (ref. 49).
References
Stack, E. C., Wang, C., Roman, K. A. & Hoyt, C. C. Multiplexed immunohistochemistry, imaging, and quantitation: a review, with an assessment of tyramide signal amplification, multispectral imaging and multiplex analysis. Methods 70, 46–58 (2014).
Angelo, M. et al. Multiplexed ion beam imaging (MIBI) of human breast tumors. Nat. Med. 20, 436–442 (2014).
Wang, Y. J. et al. Multiplexed in situ imaging mass cytometry analysis of the human endocrine pancreas and immune system in type 1 diabetes. Cell Metab. 29, 769–783 (2019).
Ptacek, J. et al. Multiplexed ion beam imaging (MIBI) for characterization of the tumor microenvironment across tumor types. Lab. Invest. 100, 1111–1123 (2020).
Parra, E. R., Francisco-Cruz, A. & Wistuba, I. I. State-of-the-art of profiling immune contexture in the era of multiplexed staining and digital analysis to study paraffin tumor tissues. Cancers (Basel) 11, 247 (2019).
Schürch, C. M. et al. Coordinated cellular neighborhoods orchestrate antitumoral immunity at the colorectal cancer invasive front. Cell 182, 1341–1359 (2020).
Gillies, R. J., Verduzco, D. & Gatenby, R. A. Evolutionary dynamics of carcinogenesis and why targeted therapy does not work. Nat. Rev. Cancer 12, 487–493 (2012).
Heindl, A., Nawaz, S. & Yuan, Y. Mapping spatial heterogeneity in the tumor microenvironment: a new era for digital pathology. Lab. Invest. 95, 377–384 (2015).
Alfarouk, K. O., Ibrahim, M. E., Gatenby, R. A. & Brown, J. S. Riparian ecosystems in human cancers. Evol. Appl. 6, 46–53 (2013).
Little, S. E. et al. Receptor tyrosine kinase genes amplified in glioblastoma exhibit a mutual exclusivity in variable proportions reflective of individual tumor heterogeneity. Cancer Res. 72, 1614–1620 (2012).
Herzenberg, L. A., Tung, J., Moore, W. A., Herzenberg, L. A. & Parks, D. R. Interpreting flow cytometry data: a guide for the perplexed. Nat. Immunol. 7, 681–685 (2006).
Aghaeepour, N. et al. Critical assessment of automated flow cytometry data analysis techniques. Nat. Methods 10, 228–238 (2013).
Shekhar, K., Brodin, P., Davis, M. M. & Chakraborty, A. K. Automatic classification of cellular expression by nonlinear stochastic embedding (ACCENSE). Proc. Natl Acad. Sci. USA 111, 202–207 (2014).
Goltsev, Y. et al. Deep profiling of mouse splenic architecture with CODEX multiplexed imaging. Cell 174, 968–981 (2018).
Black, S. et al. CODEX multiplexed tissue imaging with DNA-conjugated antibodies. Nat. Protoc. 16, 3802–3835 (2021).
Ren, X. et al. Reconstruction of cell spatial organization from single-cell RNA sequencing data based on ligand-receptor mediated self-assembly. Cell Res. 30, 763–778 (2020).
Lee, H. C., Kosoy, R., Becker, C. E., Dudley, J. T. & Kidd, B. A. Automated cell type discovery and classification through knowledge transfer. Bioinformatics 33, 1689–1695 (2017).
Wu, F. Y. The Potts model. Rev. Mod. Phys. 54, 235–268 (1982).
Storath, M., Weinmann, A., Frikel, J. & Unser, M. Joint image reconstruction and segmentation using the Potts model. Inverse Probl. 31, 025003 (2015).
Celeux, G., Forbes, F. & Peyrard, N. EM-based image segmentation using Potts models with external field. [Research Report] RR-4456 INRIA (2002). https://hal.inria.fr/inria-00072132
Pettit, J. B. et al. Identifying cell types from spatially referenced single-cell expression datasets. PLoS Comput. Biol. 10, e1003824 (2014).
Li, Q., Yi, F., Wang, T., Xiao, G. & Liang, F. Lung cancer pathological image analysis using a hidden Potts model. Cancer Inform. 16, 1176935117711910 (2017).
Celeux, G., Forbes, F. & Peyrard, N. EM procedures using mean field-like approximations for Markov model-based image segmentation. Pattern Recogn. 36, 131–144 (2003).
Samusik, N., Good, Z., Spitzer, M. H., Davis, K. L. & Nolan, G. P. Automated mapping of phenotype space with single-cell data. Nat. Methods 13, 493–496 (2016).
Van Gassen, S. et al. FlowSOM: using self-organizing maps for visualization and interpretation of cytometry data. Cytometry A 87, 636–645 (2015).
Aghaeepour, N., Nikolic, R., Hoos, H. H. & Brinkman, R. R. Rapid cell population identification in flow cytometry data. Cytometry A 79, 6–13 (2011).
Liu, X. et al. A comparison framework and guideline of clustering methods for mass cytometry data. Genome Biol. 20, 297 (2019).
Denisenko, E. et al. Systematic assessment of tissue dissociation and storage biases in single-cell and single-nucleus RNA-seq workflows. Genome Biol. 21, 130 (2020).
Leslie, T. F. & Kronenfeld, B. J. The colocation quotient: a new measure of spatial association between categorical subsets of points. Geogr. Anal. 43, 306–326 (2011).
Arnol, D., Schapiro, D., Bodenmiller, B., Saez-Rodriguez, J. & Stegle, O. Modeling cell–cell interactions from spatial molecular data with spatial variance component analysis. Cell Rep. 29, 202–211 (2019).
Ramilowski, J. A. et al. A draft network of ligand-receptor-mediated multicellular signalling in human. Nat. Commun. 6, 7866 (2015).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Puram, S. V. et al. Single-cell transcriptomic analysis of primary and metastatic tumor ecosystems in head and neck cancer. Cell 171, 1611–1624 (2017).
Reticker-Flynn, N. et al. Lymph node colonization induces tumor-immune tolerance to promote distant metastasis. Cell (2022). https://doi.org/10.1016/j.cell.2022.04.019
Zhu, G. et al. CXCR3 as a molecular target in breast cancer metastasis: inhibition of tumor cell migration and promotion of host anti-tumor immunity. Oncotarget 6, 43408–43419 (2015).
Cambien, B. et al. Organ-specific inhibition of metastatic colon carcinoma by CXCR3 antagonism. Br. J. Cancer 100, 1755–1764 (2009).
Walser, T. C. et al. Antagonism of CXCR3 inhibits lung metastasis in a murine model of metastatic breast cancer. Cancer Res. 66, 7701–7707 (2006).
Kim, D., Curthoys, N. M., Parent, M. T. & Hess, S. T. Bleed-through correction for rendering and correlation analysis in multi-colour localization microscopy. J. Opt. 15, 094011 (2013).
Rich, R. M. et al. Elimination of autofluorescence background from fluorescence tissue images by use of time-gated detection and the AzaDiOxaTriAngulenium (ADOTA) fluorophore. Anal. Bioanal. Chem. 405, 2065–2075 (2013).
Groom, J. R. & Luster, A. D. CXCR3 in T cell function. Exp. Cell Res. 317, 620–631 (2011).
Wightman, S. C. et al. Oncogenic CXCL10 signalling drives metastasis development and poor clinical outcome. Br. J. Cancer 113, 327–335 (2015).
Ranasinghe, R. & Eri, R. Modulation of the CCR6-CCl20 axis: a potential therapeutic target in inflammation and cancer. Medicina 54, 88 (2018).
Rubie, C. et al. CCL20/CCR6 expression profile in pancreatic cancer. J. Transl. Med. 8, 45 (2010).
Osuala, K. O. & Sloane, B. F. Many roles of CCL20: emphasis on breast cancer. Postdoc J. 2, 7–16 (2014).
Kindermann, R. & Snell, J. L. Markov Random Fields and their Applications (American Mathematical Society, 1980).
Tusher, V. G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl Acad. Sci. USA 98, 5116–5121 (2001).
Haribhai, D. et al. Regulatory T cells dynamically control the primary immune response to foreign antigen. J. Immunol. 178, 2961–2972 (2007).
Zhang, W., Lim, T., Li, I. & Plevritis, S. CELESTA (automate machine learning cell type identification for multiplexed in situ imaging data) [Source Code]. Code Ocean https://doi.org/10.24433/CO.0677810.v1 (2022).
Acknowledgements
This work was supported by the National Institute of Health, National Cancer Institute U54 CA209971. In addition, Q.-T.L. was supported by T31IP1598 from the TRDRP (Tobacco-Related Disease Research program). Q.-T.L., R.L. and E.G.E. were additionally supported by NIH grant R01 DE029672-01A1. Z.G. was supported by funding from Parker Institute for Cancer Immunotherapy at San Francisco, CA, USA and Stanford Cancer Institute, CA, USA. The authors are grateful to the Nolan Lab of Stanford University for their input on CODEX imaging data.
Author information
Authors and Affiliations
Contributions
W.Z., I.L. and S.K.P. conceived and designed the algorithm. W.Z. and I.L. performed computational analyses as well as manual assessment for cell type assignment using clustering and gating methods. N.E.R.-F. performed the mouse functional studies. Z.G., W.Z. and X.Z. performed the scRNA-seq analysis. S.C. performed tissue sample collection of the HNSCC cohort. S.C. and S.S. performed cell dissociation and sequencing library preparation for scRNA-seq. A.J.G., S.K.P. and J.B.S. supervised sequencing data generation. N.S. generated CODEX imaging data. Y.L. performed manual assessment on cell type assignments for the HNSCC cohort samples. R.L. and Q.-T.L. curated the clinical information for the tissue microarray cohort. C.S.K. built the HNSCC tissue microarray data and provided pathological inputs. S.K.P., E.G.E. and G.P.N. supervised the study. All authors contributed to and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
Z.G. is an inventor on two patent applications and an investor in Boom Capital Ventures, and a consultant for Mubadala Ventures, GLG and Atheneum Partners. Q.-T.L. has received grants from Varian and serves as a consultant for Nanobiotix, Coherus, Merck and Roche. All other authors have no competing interests.
Peer review
Peer review information
Nature Methods thanks Darren Tyson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Nina Vogt was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Neighborhood enrichment analysis, expression distributions of protein markers and illustration of final updated prior knowledge cell type signature matrix.
(a) Cell neighborhood enrichment analysis using Schurch et al.6 cell type annotations. Red versus blue indicates that cells of a given cell type (columns) are significantly enriched versus are not enriched, respectively, in the 5-nearest neighborhood of a cell type of interest (rows). Cells of the same or similar cell type are enriched in each other’s neighborhoods. Statistical significance is determined with p-value right tail < 0.05 and Benjamini–Hochberg adjusted p-value < 0.05. Legend for cell count indicates the number of cells below 2,000, (2,000–4,000), (4,000–6,000), (6,000–8,000) and over 8,000 for each cell type across the 70 samples. (b) Histograms of protein expressions in a representative sample. Red curves illustrate fitted bimodal Gaussian mixture model. The protein expression levels were ArcSinh transformed. (c) Illustration of final updated prior knowledge cell type signature matrix on a representative sample from Schurch et al. data. The initial user-defined cell type signature matrix is shown in Supplementary Table 1. There were no NK cells identified in this sample, and thus information on NK cells is not updated in the cell type signature matrix. White to red color indicates values from 0 to 1. Gray color indicates NA values.
Extended Data Fig. 2 Comparison between CELESTA and Schurch et al.6 annotations on the colorectal cancer dataset.
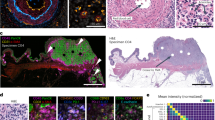
(a) Confusion matrix for each cell type identified by CELESTA (rows) versus Schurch et al.6 (column) for 70 samples. White to red color indicates values from low to high. (b) Nuclei staining for sample core 032. (c) Cytokeratin staining for sample core 032. (d) Tumor cells identified by Schurch et al. (yellow crosses) overlaid on cytokeratin staining for sample core 032. (e) Tumor cells identified by CELESTA overlaid on cytokeratin staining for sample core 032. (f) Average canonical cell type marker expressions across all the 70 samples on cells identified to be tumor cells by (i) both CELESTA and Schurch et al. (black), (ii) only CELESTA (orange), and (iii) only Schurch et al. (blue). (g) Similar to (f) but with error bars indicating 95% confidence interval based on sampling the same number of cells from each category across n = 70 samples and center values indicate mean values.
Extended Data Fig. 3 Testing leave-one-out marker and cell type resolution strategy and sensitivity analysis of user-defined parameters (hyperparameters) in CELESTA using the Schurch et al.6 dataset.
(a) Assigned cell type proportions for testing of different cell type signature matrices with each time leaving one cell type marker and corresponding cell type out. (b) Comparison of CELESTA’s performance with (yellow) and without (purple) cell type resolution strategy. (c) Average numbers of neighboring cells as a function of the bandwidth parameter across n = 70 samples. Error bar indicates standard deviation, and center value indicates mean values. (d) F1 score as a function of the number of nearest neighbors. Left panel: major cell populations. Right panel: cell types with smaller populations. (e) Effect of different values for the threshold of high marker probability expression. Left panel: Number of cells assigned to unknown cell types as a function of the threshold for high marker probability expression. Middle panel: F1 scores as a function of the threshold for high marker probability expression, for major cell types. Right panel: F1 scores as a function of the threshold for high marker probability expression, for cell types with smaller populations.
Extended Data Fig. 4 Comparison of expression probabilities versus original staining across a representative sample.
Expression probability for a given marker for each cell CELESTA (left) compared to marker staining on the original image (right). For the CELESTA result, the marker expression probability is shown at the XY coordinates of the cell, where the XY coordinates represents the cell’s center; marker expression probabilities are color-coded for values over 0.5 in light blue to over 0.9 in dark blue. Markers illustrated are: (a) aSMA, a mesenchymal marker, (b) cytokeratin, a tumor marker and (c) CD31, an endothelial marker.
Extended Data Fig. 5 Analysis of two different clustering-based methods (namely, flowMeans and FlowSOM) used to assign cell types on the Schurch et al.6 dataset.
(a) Heatmaps of cluster marker expressions on different numbers of clusters (n = 20, 30, 50) with two independent annotators (Anno1 and Anno2) to assign cluster cell types based on manual assessment of cluster protein marker expressions; light green indicates matched annotations and dark green indicates mismatched annotations. (b) Percentage of matched cluster annotations between the two annotators as a function of the number of clusters, for two different clustering methods. (c) Number of cell types identified by the two annotators as a function of the number of clusters, for the two different clustering methods. (d) The percentage of cells assigned to unknown cell types with CELESTA and the two different clustering methods, as a function of the number of clusters and the annotator. (e) F1 scores per cell type, comparing CELESTA and cell type assignments from the two annotators using the two different clustering methods, where annotations from Schurch et al. are used as ground truth. Abbreviations: Anno1 for Annotator 1; Anno2 for Annotator 2.
Extended Data Fig. 6 Visual assessment of CELESTA’s performance for a representative HNSCC sample.
(a)-(f) Identified cells are shown as yellow crosses using the x and y coordinates overlaid on canonical marker staining (white) CODEX images. For each cell type, nuclei staining and three example markers (positive and negative) important for the cell type are shown. Cell types shown (a)-(f): malignant cells, endothelial cells, fibroblast cells, B cells, NK cells, plasmacytoid dendritic cells.
Extended Data Fig. 7 Additional visual assessment of CELESTA’s performance for a representative HNSCC sample.
(a)-(f) Identified cells are shown as yellow crosses using the x and y coordinates overlaid on canonical marker staining (white) CODEX images. For each cell type, nuclei staining and three example markers (positive and negative) important for the cell type are shown. Cell types shown (a)-(f): T cells, conventional dendritic cells, neutrophils, CD8 + T cells, CD4 + T cells, Treg cells.
Extended Data Fig. 8 Gating strategies on the head and neck squamous cell carcinoma (HNSCC) samples.
Gating strategies used to identify key cell types relevant to the HNSCC study including malignant cells, endothelial cells and subtypes of T cells.
Extended Data Fig. 9 Additional scRNA-seq analysis of primary HNSCC samples and scRNA-seq analysis using public domain data from Puram et al. (2017)34.
(a) UMAP plot of identified cell clusters with node status on the study HNSCC cohort. (b)-(c) UMAP plots highlighting expression of FOXP3, IL2RA, CXCR3, CD4 and CD8A. (d) CXCR3 expression in different T cell clusters showing that CXCR3 is differentially expressed in N0 (n = 2) versus N + (n = 2) samples only in the Treg cells. (e) Violin plot of STAT1 expression in the Treg cluster between N + (n = 2) and N0 (n = 2) samples. STAT1 is a CXCR3 inducer. (f) Violin plot of CXCL9 and CXCL11 in the malignant cell cluster between N + (n = 2) and N0 (n = 2) samples. CXCL9 and CXCL11 are both ligands of CXCR3, but they are not differentially expressed in our data. (g) Heatmap shows expressions of CD274 (PD-L1), MUC1, EMT markers (CDH1 and VIM) and stemness markers (CD44 and CD24). (h) UMAP of identified cell clusters using the Puram et al. dataset. (i) UMAP of identified cell type clusters with node status color-coded. (j) UMAP plots of CD4, CD8A, and FOXP3. (k) UMAP plot of CXCR3. (l) Violin plots of CXCR3 in the T cell clusters between N + (n = 12) and N0 (n = 6) samples. (m) Violin plot of CXCL10 in malignant cell cluster 0 between N + (n = 12) and N0 (n = 6) samples. Differentially expressed genes were identified using SAMR and false discovery rate was used to adjust p-values. Center line of box plot defines data median, top value indicates largest value within 1.5 times interquartile range above 75th percentile, bottom value indicates smallest value within 1.5 times interquartile range below 25th percentile, and upper and lower bounds of the box plot indicate 75th and 25th percentile respectively. *: adjusted p-value < 0.05, **: adjusted p-value < 0.01, ***: adjusted p-value < 0.005, ****: adjusted p-value < 0.001.
Extended Data Fig. 10 Gating strategies used for mouse model studies.
Gating strategies used to study CXCL10–CXCR3 crosstalk between malignant and Treg cells in the functional studies.
Supplementary information
Supplementary Information
Supplementary Tables 1–6 and Supplementary Notes 1–3
Source data
Source Data Fig. 3
Source data for cell type annotations related to Fig.3
Source Data Fig. 5
Source data for density correlations related to Fig.5g
Source Data Fig. 6
Source data for functional mouse studies in Fig.6
Source Data Extended Data Fig. 1
Source data for cell neighborhood enrichment analysis related to Extended Data Fig.1a
Source Data Extended Data Fig. 2
Source data used to generate the confusion matrix in Extended Data Fig.2
Source Data Extended Data Fig. 5
Source data for the cluster annotations used in Extended Data Fig.5
Rights and permissions
About this article
Cite this article
Zhang, W., Li, I., Reticker-Flynn, N.E. et al. Identification of cell types in multiplexed in situ images by combining protein expression and spatial information using CELESTA. Nat Methods 19, 759–769 (2022). https://doi.org/10.1038/s41592-022-01498-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-022-01498-z
This article is cited by
-
MAPS: pathologist-level cell type annotation from tissue images through machine learning
Nature Communications (2024)
-
Precise immunofluorescence canceling for highly multiplexed imaging to capture specific cell states
Nature Communications (2024)
-
Pianno: a probabilistic framework automating semantic annotation for spatial transcriptomics
Nature Communications (2024)
-
Spatial relationships in the urothelial and head and neck tumor microenvironment predict response to combination immune checkpoint inhibitors
Nature Communications (2024)
-
Mapping the single cell spatial immune landscapes of the melanoma microenvironment
Clinical & Experimental Metastasis (2024)