Abstract
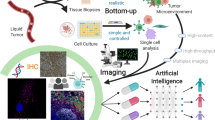
Understanding intratumoral heterogeneity—the molecular variation among cells within a tumor—promises to address outstanding questions in cancer biology and improve the diagnosis and treatment of specific cancer subtypes. Single-cell analyses, especially RNA sequencing and other genomics modalities, have been transformative in revealing novel biomarkers and molecular regulators associated with tumor growth, metastasis and drug resistance. However, these approaches fail to provide a complete picture of tumor biology, as information on cellular location within the tumor microenvironment is lost. New technologies leveraging multiplexed fluorescence, DNA, RNA and isotope labeling enable the detection of tens to thousands of cancer subclones or molecular biomarkers within their native spatial context. The expeditious growth in these techniques, along with methods for multiomics data integration, promises to yield a more comprehensive understanding of cell-to-cell variation within and between individual tumors. Here we provide the current state and future perspectives on the spatial technologies expected to drive the next generation of research and diagnostic and therapeutic strategies for cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Hooke, R. Micrographia: Or Some Physiological Descriptions of Minute Bodies Made by Magnifying Glasses, with Observations and Inquiries Thereupon (Courier, 2003).
Hajdu, S. I. A note from history: landmarks in history of cancer, part 3. Cancer 118, 1155–1168 (2012).
Coons, A. H., Creech, H. J. & Jones, R. N. Immunological properties of an antibody containing a fluorescent group. Proc. Soc. Exp. Biol. Med. 47, 200–202 (1941).
Cobb, M. Who discovered messenger RNA? Curr. Biol. 25, R526–R532 (2015).
Berger, M. F. & Mardis, E. R. The emerging clinical relevance of genomics in cancer medicine. Nat. Rev. Clin. Oncol. 15, 353–365 (2018).
Stuart, T. & Satija, R. Integrative single-cell analysis. Nat. Rev. Genet. 20, 257–272 (2019).
Kashima, Y. et al. Single-cell sequencing techniques from individual to multiomics analyses. Exp. Mol. Med. 52, 1419–1427 (2020).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Klein, C. A. Parallel progression of primary tumours and metastases. Nat. Rev. Cancer 9, 302–312 (2009).
Marusyk, A., Janiszewska, M. & Polyak, K. Intratumor heterogeneity: the Rosetta Stone of therapy resistance. Cancer Cell 37, 471–484 (2020).
DeGregori, J. Adaptive Oncogenesis: A New Understanding of How Cancer Evolves Inside Us (Harvard University Press, 2018).
Turajlic, S., Sottoriva, A., Graham, T. & Swanton, C. Resolving genetic heterogeneity in cancer. Nat. Rev. Genet. 20, 404–416 (2019).
Marine, J. C., Dawson, S. J. & Dawson, M. A. Non-genetic mechanisms of therapeutic resistance in cancer. Nat. Rev. Cancer 20, 743–756 (2020).
Sottoriva, A. et al. A Big Bang model of human colorectal tumor growth. Nat. Genet. 47, 209–216 (2015).
Casasent, A. K., Edgerton, M. & Navin, N. E. Genome evolution in ductal carcinoma in situ: invasion of the clones. J. Pathol. 241, 208–218 (2017).
Lan, X. et al. Fate mapping of human glioblastoma reveals an invariant stem cell hierarchy. Nature 549, 227–232 (2017).
Nguyen, L. V. et al. DNA barcoding reveals diverse growth kinetics of human breast tumour subclones in serially passaged xenografts. Nat. Commun. 5, 5871 (2014).
Merino, D. et al. Barcoding reveals complex clonal behavior in patient-derived xenografts of metastatic triple negative breast cancer. Nat. Commun. 10, 766 (2019).
Echeverria, G. V. et al. High-resolution clonal mapping of multi-organ metastasis in triple negative breast cancer. Nat. Commun. 9, 5079 (2018).
Livet, J. et al. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature 450, 56–62 (2007).
Snippert, H. J. et al. Intestinal crypt homeostasis results from neutral competition between symmetrically dividing Lgr5 stem cells. Cell 143, 134–144 (2010). The authors developed a multicolor reporter mouse termed R26R-Confetti based on the Brainbow-2.1 cassette. After Cre-mediated recombination, one of four fluorescent marker proteins is stochastically expressed, allowing clonal lineage tracing of stem cells.
Weber, K., Bartsch, U., Stocking, C. & Fehse, B. A multicolor panel of novel lentiviral ‘gene ontology’ (LeGO) vectors for functional gene analysis. Mol. Ther. 16, 698–706 (2008). A panel of LeGO vectors provide a flexible tool for investigating clonality and gene networks using simultaneous ectopic expression of fluorescent proteins, transgenes and shRNAs.
Weissman, T. A. & Pan, Y. A. Brainbow: new resources and emerging biological applications for multicolor genetic labeling and analysis. Genetics 199, 293–306 (2014).
El-Nachef, D. et al. A rainbow reporter tracks single cells and reveals heterogeneous cellular dynamics among pluripotent stem cells and their differentiated derivatives. Stem Cell Rep. 15, 226–241 (2020).
Boone, P. G. et al. A cancer rainbow mouse for visualizing the functional genomics of oncogenic clonal expansion. Nat. Commun. 10, 5490 (2019).
Kim, G. B. et al. Rapid generation of somatic mouse mosaics with locus-specific, stably integrated transgenic elements. Cell 179, 251–267 (2019).
Schepers, A. G. et al. Lineage tracing reveals Lgr5+ stem cell activity in mouse intestinal adenomas. Science 337, 730–735 (2012).
Fumagalli, A. et al. Plasticity of Lgr5-negative cancer cells drives metastasis in colorectal cancer. Cell Stem Cell 26, 569–578 (2020).
Reeves, M. Q., Kandyba, E., Harris, S., Del Rosario, R. & Balmain, A. Multicolour lineage tracing reveals clonal dynamics of squamous carcinoma evolution from initiation to metastasis. Nat. Cell Biol. 20, 699–709 (2018).
Zomer, A. et al. Intravital imaging of cancer stem cell plasticity in mammary tumors. Stem Cells 31, 602–606 (2013).
Rios, A. C. et al. Intraclonal plasticity in mammary tumors revealed through large-scale single-cell resolution 3D imaging. Cancer Cell 35, 618–632 (2019).
Tiede, S. et al. Multi-color clonal tracking reveals intra-stage proliferative heterogeneity during mammary tumor progression. Oncogene 40, 12–27 (2021).
Weber, K. et al. RGB marking facilitates multicolor clonal cell tracking. Nat. Med. 17, 504–509 (2011).
Roh, V. et al. Cellular barcoding identifies clonal substitution as a hallmark of local recurrence in a surgical model of head and neck squamous cell carcinoma. Cell Rep. 25, 2208–2222 (2018).
Lamprecht, S. et al. Multicolor lineage tracing reveals clonal architecture and dynamics in colon cancer. Nat. Commun. 8, 1406 (2017).
Mohme, M. et al. Optical barcoding for single-clone tracking to study tumor heterogeneity. Mol. Ther. 25, 621–633 (2017).
Lambert, T. J. FPbase: a community-editable fluorescent protein database. Nat. Methods 16, 277–278 (2019).
Anzalone, A. V., Jimenez, M. & Cornish, V. W. FRAME-tags: genetically encoded fluorescent markers for multiplexed barcoding and time-resolved tracking of live cells. Preprint at bioRxiv https://doi.org/10.1101/2021.04.09.436507 (2021).
Weber, K., Thomaschewski, M., Benten, D. & Fehse, B. RGB marking with lentiviral vectors for multicolor clonal cell tracking. Nat. Protoc. 7, 839–849 (2012).
Medaglia, C. et al. Spatial reconstruction of immune niches by combining photoactivatable reporters and scRNA-seq. Science 358, 1622–1626 (2017).
Yamashita, R. et al. Deep learning model for the prediction of microsatellite instability in colorectal cancer: a diagnostic study. Lancet Oncol. 22, 132–141 (2021).
Naik, N. et al. Deep learning-enabled breast cancer hormonal receptor status determination from base-level H&E stains. Nat. Commun. 11, 5727 (2020).
Tóth, Z. E. & Mezey, É. Simultaneous visualization of multiple antigens with tyramide signal amplification using antibodies from the same species. J. Histochem. Cytochem. 55, 545–554 (2007). Microwave-mediated antibody stripping in combination with tyramide signal amplification enables use of multiple antibodies from the same host, and both rare and abundant antigens can be detected.
Banik, G. et al. High-dimensional multiplexed immunohistochemical characterization of immune contexture in human cancers. Methods Enzymol. 635, 1–20 (2020).
Tsujikawa, T. et al. Quantitative multiplex immunohistochemistry reveals myeloid-inflamed tumor-immune complexity associated with poor prognosis. Cell Rep. 19, 203–217 (2017).
Stack, E. C., Wang, C., Roman, K. A. & Hoyt, C. C. Multiplexed immunohistochemistry, imaging, and quantitation: a review, with an assessment of tyramide signal amplification, multispectral imaging and multiplex analysis. Methods 70, 46–58 (2014).
Lin, J. R., Fallahi-Sichani, M. & Sorger, P. K. Highly multiplexed imaging of single cells using a high-throughput cyclic immunofluorescence method. Nat. Commun. 6, 8390 (2015).
Gerdes, M. J. et al. Highly multiplexed single-cell analysis of formalin-fixed, paraffin-embedded cancer tissue. Proc. Natl Acad. Sci. USA 110, 11982–11987 (2013).
Herbel, C. et al. Abstract 4694: Evaluation of tumor-associated antigen expression with the MACSima high-content imaging platform. Cancer Res. 79, 4694 (2019).
Goltsev, Y. et al. Deep profiling of mouse splenic architecture with CODEX multiplexed imaging. Cell 174, 968–981 (2018). An indexable tagging system whereby antibodies are labeled with uniquely designed oligonucleotide duplexes. Pairs of antibodies are visualized in a multicycle protocol to allow antigen multiplexing.
Schürch, C. M. et al. Coordinated cellular neighborhoods orchestrate antitumoral immunity at the colorectal cancer invasive front. Cell 182, 1341–1359 (2020).
Saka, S. K. et al. Immuno-SABER enables highly multiplexed and amplified protein imaging in tissues. Nat. Biotechnol. 37, 1080–1090 (2019).
Bendall, S. C. et al. Single-cell mass cytometry of differential immune and drug responses across a human hematopoietic continuum. Science 332, 687–696 (2011).
Giesen, C. et al. Highly multiplexed imaging of tumor tissues with subcellular resolution by mass cytometry. Nat. Methods 11, 417–422 (2014). IMC combines CyTOF mass cytometry and a high-resolution laser ablation system with immunocytochemistry and IHC techniques to image 32 proteins simultaneously. Here IMC is used to study microenvironmental heterogeneity in human breast cancer, while in ref. 55 it is used for multimodal RNA and protein imaging.
Schulz, D. et al. Simultaneous multiplexed imaging of mrna and proteins with subcellular resolution in breast cancer tissue samples by mass cytometry. Cell Syst. 6, 25–36 (2018).
Angelo, M. et al. Multiplexed ion beam imaging (MIBI) of human breast tumors. Nat. Med. 20, 436–442 (2014). MIBI uses secondary ion mass spectrometry to image antibodies tagged with isotopically pure elemental metal reporters. Here MIBI is used to study pathogenesis in human breast cancer, and it has also been used in several cancer studies, including in ref. 57.
Keren, L. et al. A structured tumor-immune microenvironment in triple negative breast cancer revealed by multiplexed ion beam imaging. Cell 174, 1373–1387 (2018).
Keren, L. et al. MIBI-TOF: a multiplexed imaging platform relates cellular phenotypes and tissue structure. Sci. Adv. 5, eaax5851 (2019).
Beechem, J. M. High-plex spatially resolved RNA and protein detection using digital spatial profiling: a technology designed for immuno-oncology biomarker discovery and translational research. Methods Mol. Biol. 2055, 563–583 (2020).
Brady, L. et al. Inter- and intra-tumor heterogeneity of metastatic prostate cancer determined by digital spatial gene expression profiling. Nat. Commun. 12, 1426 (2021).
Gall, J. G. & Pardue, M. L. Formation and detection of RNA–DNA hybrid molecules in cytological preparations. Proc. Natl Acad. Sci. USA 63, 378–383 (1969).
Pardue, M. L. & Gall, J. G. Molecular hybridization of radioactive DNA to the DNA of cytological preparations. Proc. Natl Acad. Sci. USA 64, 600–604 (1969).
Rudkin, G. T. & Stollar, B. D. High resolution detection of DNA–RNA hybrids in situ by indirect immunofluorescence. Nature 265, 472–473 (1977).
Bauman, J. G. J., Wiegant, J., Borst, P. & van Duijn, P. A new method for fluorescence microscopical localization of specific DNA sequences by in situ hybridization of fluorochrome-labelled RNA. Exp. Cell. Res. 128, 485–490 (1980).
Takei, Y. et al. Integrated spatial genomics reveals global architecture of single nuclei. Nature 590, 344–350 (2021).
Su, J. H., Zheng, P., Kinrot, S. S., Bintu, B. & Zhuang, X. Genome-scale imaging of the 3D organization and transcriptional activity of chromatin. Cell 182, 1641–1659 (2020).
Payne, A. C. et al. In situ genome sequencing resolves DNA sequence and structure in intact biological samples. Science 371, eaay3446 (2021).
Nguyen, H. Q. et al. 3D mapping and accelerated super-resolution imaging of the human genome using in situ sequencing. Nat. Methods 17, 822–832 (2020).
Femino, A. M., Fay, F. S., Fogarty, K. & Singer, R. H. Visualization of single RNA transcripts in situ. Science 280, 585–590 (1998).
Raj, A., Van Den Bogaard, P., Rifkin, S. A., Van Oudenaarden, A. & Tyagi, S. Imaging individual mRNA molecules using multiple singly labeled probes. Nat. Methods 5, 877–879 (2008). A sensitive smFISH method for detecting individual mRNA molecules in fixed cells via hybridization of multiple short probes. This paved the way for future multiplexed smFISH methods.
Shaffer, S. M. et al. Rare cell variability and drug-induced reprogramming as a mode of cancer drug resistance. Nature 546, 431–435 (2017).
Lubeck, E. & Cai, L. Single-cell systems biology by super-resolution imaging and combinatorial labeling. Nat. Methods 9, 743–748 (2012).
Codeluppi, S. et al. Spatial organization of the somatosensory cortex revealed by osmFISH. Nat. Methods 15, 932–935 (2018).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015). MERFISH is a temporal barcoded smFISH method that measures 100–1,001 genes with high spatial resolution and detection efficiency. Xia et al.76 build on this to enable detection of 10,000+ mRNAs in situ, leading to the identification of approximately 1,600 cell cycle-dependent genes.
Alon, S. et al. Expansion sequencing: spatially precise in situ transcriptomics in intact biological systems. Science 371, eaax2656 (2021).
Xia, C., Fan, J., Emanuel, G., Hao, J. & Zhuang, X. Spatial transcriptome profiling by MERFISH reveals subcellular RNA compartmentalization and cell cycle-dependent gene expression. Proc. Natl Acad. Sci. USA 116, 19490–19499 (2019).
Goh, J. J. L. et al. Highly specific multiplexed RNA imaging in tissues with split-FISH. Nat. Methods 17, 689–693 (2020).
Lubeck, E., Coskun, A. F., Zhiyentayev, T., Ahmad, M. & Cai, L. Single-cell in situ RNA profiling by sequential hybridization. Nat. Methods 11, 360–361 (2014). seqFISH uses temporal barcoding to measure 12 genes. Eng et al.80 expand this to achieve multiplexing of 10,000 genes in single cells using pseudocolors to circumvent the problem of optical crowding.
Shah, S., Lubeck, E., Zhou, W. & Cai, L. In situ transcription profiling of single cells reveals spatial organization of cells in the mouse hippocampus. Neuron 92, 342–357 (2016).
Eng, C.-H. L. et al. Transcriptome-scale super-resolved imaging in tissues by RNA seqFISH+. Nature 568, 235–239 (2019).
Frieda, K. L. et al. Synthetic recording and in situ readout of lineage information in single cells. Nature 541, 107–111 (2017).
Askary, A. et al. In situ readout of DNA barcodes and single base edits facilitated by in vitro transcription. Nat. Biotechnol. 38, 66–75 (2020).
Chow, K.-H. K. et al. Imaging cell lineage with a synthetic digital recording system. Science 372, eabb3099 (2021).
Wang, F. et al. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J. Mol. Diagn. 14, 22–29 (2012).
Nichterwitz, S. et al. Laser capture microscopy coupled with Smart-seq2 for precise spatial transcriptomic profiling. Nat. Commun. 7, 12139 (2016).
Chen, J. et al. Spatial transcriptomic analysis of cryosectioned tissue samples with Geo-seq. Nat. Protoc. 12, 566–580 (2017).
Casasent, A. K. et al. Multiclonal invasion in breast tumors identified by topographic single cell sequencing. Cell 172, 205–217 (2018).
Pennycuick, A. et al. Immune surveillance in clinical regression of preinvasive squamous cell lung cancer. Cancer Discov. 10, 1489–1499 (2020).
Lovatt, D. et al. Transcriptome in vivo analysis (TIVA) of spatially defined single cells in live tissue. Nat. Methods 11, 190–196 (2014).
Larsson, L., Frisén, J. & Lundeberg, J. Spatially resolved transcriptomics adds a new dimension to genomics. Nat. Methods 18, 15–18 (2021).
Ståhl, P. L. et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353, 78–82 (2016). An oligonucleotide array with positional barcodes to capture mRNA from histological tissue sections, demonstrating that individual experiments can recover thousands of transcript-coupled spatial barcodes at 10-μm resolution. This study paved the way for higher-resolution spatial transcriptomics to single-cell resolution.
Berglund, E. et al. Spatial maps of prostate cancer transcriptomes reveal an unexplored landscape of heterogeneity. Nat. Commun. 9, 2419 (2018).
Thrane, K., Eriksson, H., Maaskola, J., Hansson, J. & Lundeberg, J. Spatially resolved transcriptomics enables dissection of genetic heterogeneity in stage III cutaneous malignant melanoma. Cancer Res. 78, 5970–5979 (2018).
Rodriques, S. G. et al. Slide-seq: a scalable technology for measuring genome-wide expression at high spatial resolution. Science 363, 1463–1467 (2019).
Vickovic, S. et al. High-definition spatial transcriptomics for in situ tissue profiling. Nat. Methods 16, 987–990 (2019).
Liu, Y. et al. High-spatial-resolution multi-omics sequencing via deterministic barcoding in tissue. Cell 183, 1665–1681 (2020).
Chen, A. et al. Large field of view-spatially resolved transcriptomics at nanoscale resolution. Preprint at bioRxiv https://doi.org/10.1101/2021.01.17.427004 (2021).
Ke, R. et al. In situ sequencing for RNA analysis in preserved tissue and cells. Nat. Methods 10, 857–860 (2013).
Larsson, C., Grundberg, I., Söderberg, O. & Nilsson, M. In situ detection and genotyping of individual mRNA molecules. Nat. Methods 7, 395–397 (2010).
Gyllborg, D. et al. Hybridization-based in situ sequencing (HybISS) for spatially resolved transcriptomics in human and mouse brain tissue. Nucleic Acids Res. 48, e112 (2020).
Lee, J. H. et al. Fluorescent in situ sequencing (FISSEQ) of RNA for gene expression profiling in intact cells and tissues. Nat. Protoc. 10, 442–458 (2015).
Lee, J. H. et al. Highly multiplexed subcellular RNA sequencing in situ. Science 343, 1360–1363 (2014). A protocol for unbiased gene expression profiling in situ in fixed histological samples. RNA is converted into cross-linked cDNA amplicons and sequenced manually on a confocal microscope.
Massingham, T. & Goldman, N. Error-correcting properties of the SOLiD exact call chemistry. BMC Bioinformatics 13, 145 (2012).
Strell, C. et al. Placing RNA in context and space—methods for spatially resolved transcriptomics. FEBS J. 286, 1468–1481 (2019).
Lee, J. H. Quantitative approaches for investigating the spatial context of gene expression. Wiley Interdiscip. Rev. Syst. Biol. Med. 9, e1369 (2017).
Baharlou, H., Canete, N. P., Cunningham, A. L., Harman, A. N. & Patrick, E. Mass cytometry imaging for the study of human diseases—applications and data analysis strategies. Front. Immunol. 10, 2657 (2019).
Efremova, M. & Teichmann, S. A. Computational methods for single-cell omics across modalities. Nat. Methods 17, 14–17 (2020).
Van Valen, D. A. et al. Deep learning automates the quantitative analysis of individual cells in live-cell imaging experiments. PLoS Comput. Biol. 12, e1005177 (2016).
Caicedo, J. C. et al. Nucleus segmentation across imaging experiments: the 2018 Data Science Bowl. Nat. Methods 16, 1247–1253 (2019).
Berg, S. et al. ilastik: interactive machine learning for (bio)image analysis. Nat. Methods 16, 1226–1232 (2019).
Zhu, Q., Shah, S., Dries, R., Cai, L. & Yuan, G. C. Identification of spatially associated subpopulations by combining scRNAseq and sequential fluorescence in situ hybridization data. Nat. Biotechnol. 36, 1183–1190 (2018).
Hollandi, R. et al. nucleAIzer: a parameter-free deep learning framework for nucleus segmentation using image style transfer. Cell Syst. 10, 453–458 (2020).
Schmidt, U., Weigert, M., Broaddus, C. & Myers, G. Cell detection with star-convex polygons. in International Conference on Medical Image Computing and Computer-Assisted Intervention 265–273 (Springer, 2018).
Weigert, M., Schmidt, U., Haase, R., Sugawara, K. & Myers, G. Star-convex polyhedra for 3D object detection and segmentation in microscopy. in Proceedings of the IEEE/CVF Winter Conference on Applications of Computer Vision 3666–3673 (IEEE, 2020).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Palla, G. et al. Squidpy: a scalable framework for spatial single cell 2 analysis. Preprint at bioRxiv https://doi.org/10.1101/2021.02.19.431994 (2021).
Pham, D. et al. stLearn: integrating spatial location, tissue morphology and gene expression to find cell types, cell–cell interactions and spatial trajectories within undissociated tissues. Preprint at bioRxiv https://doi.org/10.1101/2020.05.31.125658 (2020).
Righelli, D. et al. SpatialExperiment: infrastructure for spatially resolved transcriptomics data in R using Bioconductor. Preprint at bioRxiv https://doi.org/10.1101/2021.01.27.428431 (2021).
Dries, R. et al. Giotto: a toolbox for integrative analysis and visualization of spatial expression data. Genome Biol. 22, 78 (2021).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Bergenstråhle, J., Larsson, L. & Lundeberg, J. Seamless integration of image and molecular analysis for spatial transcriptomics workflows. BMC Genomics 21, 482 (2020).
Lun, A. T. L., Bach, K. & Marioni, J. C. Pooling across cells to normalize single-cell RNA sequencing data with many zero counts. Genome Biol. 17, 75 (2016).
Senabouth, A. et al. Ascend: R package for analysis of single-cell RNA-seq data. Gigascience 8, giz087 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Perkel, J. M. Starfish enterprise: finding RNA patterns in single cells. Nature 572, 549–551 (2019).
Alquicira-Hernandez, J., Sathe, A., Ji, H. P., Nguyen, Q. & Powell, J. E. ScPred: accurate supervised method for cell-type classification from single-cell RNA-seq data. Genome Biol. 20, 264 (2019).
Elosua-Bayes, M., Nieto, P., Mereu, E., Gut, I. & Heyn, H. SPOTlight: seeded NMF regression to deconvolute spatial transcriptomics spots with single-cell transcriptomes. Nucleic Acids Res. 49, e50 (2021).
Kleshchevnikov, V. et al. Comprehensive mapping of tissue cell architecture via integrated single cell and spatial transcriptomics. Preprint at bioRxiv https://doi.org/10.1101/2020.11.15.378125 (2020).
Cable, D. M. et al. Robust decomposition of cell type mixtures in spatial transcriptomics. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-00830-w (2021). RCTD uses cross-platform integration of single-cell sequencing data and spatial transcriptomics to predict cell type composition and location. Technical variation is accounted for between different technologies.
Moncada, R. et al. Integrating microarray-based spatial transcriptomics and single-cell RNA-seq reveals tissue architecture in pancreatic ductal adenocarcinomas. Nat. Biotechnol. 38, 333–342 (2020). Among the first to use scRNA-seq data as a reference to resolve cell type information in each microarray-based spatial spot, this approach was used to find a spatially restricted distribution of macrophages and fibroblasts in pancreatic ductal adenocarcinoma tissue.
Ji, A. L. et al. Multimodal analysis of composition and spatial architecture in human squamous cell carcinoma. Cell 182, 497–514 (2020). scRNA-seq, spatial transcriptomics and spatial proteomic technologies are combined to study the cellular and molecular profiles of ten human skin squamous cell carcinoma and matched normal tissue samples. This multimodal approach and integrative analysis were used to demonstrate the heterogeneity of tumor cells at tumor edges, as well as their spatial relationships and communication networks with immune and stromal cells.
Welch, J. D. et al. Single-cell multi-omic integration compares and contrasts features of brain cell identity. Cell 177, 1873–1887 (2019).
Lopez, R. et al. A joint model of unpaired data from scRNA-seq and spatial transcriptomics for imputing missing gene expression measurements. Preprint at https://arxiv.org/abs/1905.02269 (2019).
Edsgärd, D., Johnsson, P. & Sandberg, R. Identification of spatial expression trends in single-cell gene expression data. Nat. Methods 15, 339–342 (2018).
Sun, S., Zhu, J. & Zhou, X. Statistical analysis of spatial expression patterns for spatially resolved transcriptomic studies. Nat. Methods 17, 193–200 (2020).
Bergenstråhle, J., Bergenstråhle, L. & Lundeberg, J. SpatialCPie: an R/Bioconductor package for spatial transcriptomics cluster evaluation. BMC Bioinformatics 21, 161 (2020).
Zhao, E. et al. Spatial transcriptomics at subspot resolution with BayesSpace. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-00935-2 (2021).
Biancalani, T. et al. Deep learning and alignment of spatially-resolved whole transcriptomes of single cells in the mouse brain with Tangram. Preprint at bioRxiv https://doi.org/10.1101/2020.08.29.272831 (2020).
Hu, J. et al. Integrating gene expression, spatial location and histology to identify spatial domains and spatially variable genes by graph convolutional network. Preprint at bioRxiv https://doi.org/10.1101/2020.11.30.405118 (2020).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 19, 477 (2018).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Kueckelhaus, J. et al. Inferring spatially transient gene expression pattern from spatial transcriptomic studies. Preprint at bioRxiv https://doi.org/10.1101/2020.10.20.346544 (2020).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Tran, M. et al. Spatial analysis of ligand–receptor interactions in skin cancer at genome-wide and single-cell resolution. Preprint at bioRxiv https://doi.org/10.1101/2020.09.10.290833 (2020).
Schapiro, D. et al. HistoCAT: analysis of cell phenotypes and interactions in multiplex image cytometry data. Nat. Methods 14, 873–876 (2017).
Elyanow, R., Zeira, R., Land, M. & Raphael, B. J. STARCH: copy number and clone inference from spatial transcriptomics data. Phys. Biol. 18, 035001 (2021).
Kiemen, A. et al. In situ characterization of the 3D microanatomy of the pancreas and pancreatic cancer at single cell resolution. Preprint at bioRxiv https://doi.org/10.1101/2020.12.08.416909 (2020).
Su, A. et al. A deep learning model for molecular label transfer that enables cancer cell identification from histopathology images. Preprint at bioRxiv https://doi.org/10.1101/2021.03.18.436004 (2021).
Tan, X., Su, A., Tran, M. & Nguyen, Q. SpaCell: integrating tissue morphology and spatial gene expression to predict disease cells. Bioinformatics 36, 2293–2294 (2020).
Stoltzfus, C. R. et al. CytoMAP: a spatial analysis toolbox reveals features of myeloid cell organization in lymphoid tissues. Cell Rep. 31, 107523 (2020).
Browaeys, R., Saelens, W. & Saeys, Y. NicheNet: modeling intercellular communication by linking ligands to target genes. Nat. Methods 17, 159–162 (2020).
He, B. et al. Integrating spatial gene expression and breast tumour morphology via deep learning. Nat. Biomed. Eng. 4, 827–834 (2020). stNet demonstrates successful application of deep learning to predict gene expression using H&E images, creating the potential for predicting breast cancer gene markers using histological images.
Schmauch, B. et al. A deep learning model to predict RNA-seq expression of tumours from whole slide images. Nat. Commun. 11, 3877 (2020).
Bergenstråhle, L. et al. Super-resolved spatial transcriptomics by deep data fusion. Preprint at bioRxiv https://doi.org/10.1101/2020.02.28.963413 (2020).
Hu, K. H. et al. ZipSeq: barcoding for real-time mapping of single cell transcriptomes. Nat. Methods 17, 833–843 (2020).
Wang, X. et al. Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science 361, eaat5691 (2018).
Acknowledgements
For Fig. 2, we thank L. Cai for contributing seqFISH data; S. Lakhani and P. Kalita de Croft for clinical annotation of breast cancer tissue; and Ionpath for assisting with tissue staining and generation of MIBI data. We thank D. Brown and T. Weber for critical feedback. M.-L.A.-L. is supported by funding from the Viertel Foundation Senior Medical Research Fellowship, NHMRC grant GNT1182155 and the Harry Secomb Foundation, managed by Perpetual. Q.N. is supported by Australian Research Council DECRA fellowship DE190100116 and NHMRC grant GNT2001514. D.M. is supported by Susan G. Komen and Cancer Australia, CCR19606878, a grant from the National Breast Cancer Foundation, Australia, IIRS-19-082 and the Grant-in-Aid Scheme administered by Cancer Council Victoria. The Olivia Newton-John Cancer Research Institute gratefully acknowledges the generous support of the Love Your Sister Foundation. S.H.N. is supported by NHMRC grants GNT1062820, GNT1100033, GNT1101378, GNT1124812 and GNT1145184. We also acknowledge support from the ACRF Centre for Imaging the Tumour Environment at the Olivia Newton-John Cancer Research Institute, the ACRF Program for Resolving Cancer Complexity and Therapeutic Resistance at WEHI and the Operational Infrastructure Support Program provided by the Victorian government and the NHMRC Independent Research Institutes Infrastructure Support Scheme (IRIISS) grant. The contents of the published material are solely the responsibility of the individual authors and do not reflect the views of Cancer Australia, NHMRC or other funding agencies.
Author information
Authors and Affiliations
Contributions
All authors contributed to the writing of the manuscript, with all figures generated by S.M.L. along with X.T. for Fig. 6b. Primary data for Fig. 2 were contributed by V.C.W., J.B., K.L.R. and D.M. (Fig. 2a), M.-L.A.-L. (Fig. 2b), and X.T. and Q.N. (Fig. 2d).
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Rita Strack was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Table 1.
Rights and permissions
About this article
Cite this article
Lewis, S.M., Asselin-Labat, ML., Nguyen, Q. et al. Spatial omics and multiplexed imaging to explore cancer biology. Nat Methods 18, 997–1012 (2021). https://doi.org/10.1038/s41592-021-01203-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-021-01203-6
This article is cited by
-
Mapping cancer biology in space: applications and perspectives on spatial omics for oncology
Molecular Cancer (2024)
-
Thermal-plex: fluidic-free, rapid sequential multiplexed imaging with DNA-encoded thermal channels
Nature Methods (2024)
-
STEM enables mapping of single-cell and spatial transcriptomics data with transfer learning
Communications Biology (2024)
-
Microenvironmental reorganization in brain tumors following radiotherapy and recurrence revealed by hyperplexed immunofluorescence imaging
Nature Communications (2024)
-
Decoder-seq enhances mRNA capture efficiency in spatial RNA sequencing
Nature Biotechnology (2024)