Abstract
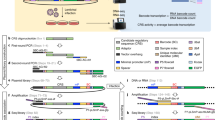
Massively parallel reporter assays (MPRAs) functionally screen thousands of sequences for regulatory activity in parallel. To date, there are limited studies that systematically compare differences in MPRA design. Here, we screen a library of 2,440 candidate liver enhancers and controls for regulatory activity in HepG2 cells using nine different MPRA designs. We identify subtle but significant differences that correlate with epigenetic and sequence-level features, as well as differences in dynamic range and reproducibility. We also validate that enhancer activity is largely independent of orientation, at least for our library and designs. Finally, we assemble and test the same enhancers as 192-mers, 354-mers and 678-mers and observe sizable differences. This work provides a framework for the experimental design of high-throughput reporter assays, suggesting that the extended sequence context of tested elements and to a lesser degree the precise assay, influence MPRA results.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
We developed a fully reproducible MPRA processing pipeline available to process the data into final enhancer activity scores. Raw and processed data have been deposited in the Gene Expression Omnibus at accession number GSE142696.
Code availability
A reproducible processing pipeline for MPRA data is available as a Nextflow-based MPRA processing pipeline named MPRAflow (https://github.com/shendurelab/MPRAflow)44.
References
Banerji, J., Rusconi, S. & Schaffner, W. Expression of a β-globin gene is enhanced by remote SV40 DNA sequences. Cell 27, 299–308 (1981).
Moreau, P. et al. The SV40 72 base repair repeat has a striking effect on gene expression both in SV40 and other chimeric recombinants. Nucleic Acids Res. 9, 6047–6068 (1981).
Banerji, J., Olson, L. & Schaffner, W. A lymphocyte-specific cellular enhancer is located downstream of the joining region in immunoglobulin heavy chain genes. Cell 33, 729–740 (1983).
Neuberger, M. S. Expression and regulation of immunoglobulin heavy chain gene transfected into lymphoid cells. EMBO J. 2, 1373–1378 (1983).
Bernstein, B. E. et al. The NIH roadmap epigenomics mapping consortium. Nat. Biotechnol. 28, 1045–1048 (2010).
Kawaji, H., Kasukawa, T., Forrest, A., Carninci, P. & Hayashizaki, Y. The FANTOM5 collection, a data series underpinning mammalian transcriptome atlases in diverse cell types. Sci. Data 4, 170113 (2017).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
ENCODE Project Consortium. A user’s guide to the encyclopedia of DNA elements (ENCODE). PLoS Biol. 9, e1001046 (2011).
Patwardhan, R. P. et al. High-resolution analysis of DNA regulatory elements by synthetic saturation mutagenesis. Nat. Biotechnol. 27, 1173–1175 (2009).
Patwardhan, R. P. et al. Massively parallel functional dissection of mammalian enhancers in vivo. Nat. Biotechnol. 30, 265–270 (2012).
Melnikov, A. et al. Systematic dissection and optimization of inducible enhancers in human cells using a massively parallel reporter assay. Nat. Biotechnol. 30, 271 (2012).
Vockley, C. M. et al. Massively parallel quantification of the regulatory effects of noncoding genetic variation in a human cohort. Genome Res. 25, 1206–1214 (2015).
Tewhey, R. et al. Direct identification of hundreds of expression-modulating variants using a multiplexed reporter assay. Cell 172, 1132–1134 (2018).
Ulirsch, J. C. et al. Systematic functional dissection of common genetic variation affecting red blood cell traits. Cell 165, 1530–1545 (2016).
Liu, S. et al. Systematic identification of regulatory variants associated with cancer risk. Genome Biol. 18, 194 (2017).
Arnold, C. D. et al. Genome-wide quantitative enhancer activity maps identified by STARR-seq. Science 339, 1074–1077 (2013).
Kwasnieski, J. C., Fiore, C., Chaudhari, H. G. & Cohen, B. A. High-throughput functional testing of ENCODE segmentation predictions. Genome Res. 24, 1595–1602 (2014).
Inoue, F. et al. A systematic comparison reveals substantial differences in chromosomal versus episomal encoding of enhancer activity. Genome Res. 27, 38–52 (2017).
Klein, J. C. et al. Functional testing of thousands of osteoarthritis-associated variants for regulatory activity. Nat. Commun. 10, 2434 (2019).
Arnold, C. D. et al. Quantitative genome-wide enhancer activity maps for five Drosophila species show functional enhancer conservation and turnover during cis-regulatory evolution. Nat. Genet. 46, 685–692 (2014).
Klein, J. C., Keith, A., Agarwal, V., Durham, T. & Shendure, J. Functional characterization of enhancer evolution in the primate lineage. Genome Biol. 19, 99 (2018).
Muerdter, F. et al. Resolving systematic errors in widely used enhancer activity assays in human cells. Nat. Methods 15, 141–149 (2018).
Vanhille, L. et al. High-throughput and quantitative assessment of enhancer activity in mammals by CapStarr-seq. Nat. Commun. 6, 6905 (2015).
Wang, X. et al. High-resolution genome-wide functional dissection of transcriptional regulatory regions and nucleotides in human. Nat. Commun. 9, 5380 (2018).
Klein, J. C. et al. Multiplex pairwise assembly of array-derived DNA oligonucleotides. Nucleic Acids Res. 44, e43 (2016).
Kircher, M. et al. Saturation mutagenesis of disease-associated regulatory elements. Nat. Commun. 10, 3583 (2019).
Hill, A. J. et al. On the design of CRISPR-based single-cell molecular screens. Nat. Methods 15, 271–274 (2018).
Sack, L. M., Davoli, T., Xu, Q., Li, M. Z. & Elledge, S. J. Sources of error in mammalian genetic screens. G3 6, 2781–2790 (2016).
Smith, R. P. et al. Massively parallel decoding of mammalian regulatory sequences supports a flexible organizational model. Nat. Genet. 45, 1021–1028 (2013).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Shiraki, T. et al. Cap analysis gene expression for high-throughput analysis of transcriptional starting point and identification of promoter usage. Proc. Natl Acad. Sci. USA 100, 15776–15781 (2003).
FANTOM Consortium et al. Supplementary figures, tables and texts for FANTOM 5 phase 2. Figshare https://doi.org/10.6084/m9.figshare.1288777 (2015).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Engreitz, J. M. et al. Local regulation of gene expression by lncRNA promoters, transcription and splicing. Nature 539, 452–455 (2016).
van Arensbergen, J. et al. Genome-wide mapping of autonomous promoter activity in human cells. Nat. Biotechnol. https://doi.org/10.1038/nbt.3754 (2016).
Kvon, E. Z., Stampfel, G., Yáñez-Cuna, J. O., Dickson, B. J. & Stark, A. HOT regions function as patterned developmental enhancers and have a distinct cis-regulatory signature. Genes Dev. 26, 908–913 (2012).
Mikhaylichenko, O. et al. The degree of enhancer or promoter activity is reflected by the levels and directionality of eRNA transcription. Genes Dev. 32, 42–57 (2018).
Weingarten-Gabbay, S. et al. Systematic interrogation of human promoters. Genome Res. 29, 171–183 (2019).
Plesa, C., Sidore, A. M., Lubock, N. B., Zhang, D. & Kosuri, S. Multiplexed gene synthesis in emulsions for exploring protein functional landscapes. Science 359, 343–347 (2018).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Klein J. C. et al. A systematic evaluation of the design, orientation, and sequence context dependencies of massively parallel reporter assays. Protoc. Exch. https://doi.org/10.21203/rs.3.pex-1065/v1 (2020).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30, 614–620 (2014).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at arXiv https://arxiv.org/abs/1303.3997 (2013).
Gordon, M. G. et al. lentiMPRA and MPRAflow for high-throughput functional characterization of gene regulatory elements. Nat. Protoc. 15, 2387–2412 (2020).
Karolchik, D. et al. The UCSC Genome Browser database: 2014 update. Nucleic Acids Res. 42, D764–D770 (2014).
Zhou, J. & Troyanskaya, O. G. Predicting effects of noncoding variants with deep learning-based sequence model. Nat. Methods 12, 931–934 (2015).
Alipanahi, B., Delong, A., Weirauch, M. T. & Frey, B. J. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nat. Biotechnol. 33, 831–838 (2015).
Roadmap Epigenomics Consortium. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Acknowledgements
We thank S. Kim and other members of the Shendure and Ahituv laboratories for general advice and critical feedback on the manuscript. This work was supported by the National Human Genome Research Institute grants 1UM1HG009408 (N.A. and J.S.), 5R01HG009136 (J.S.), 1R21HG010065 (N.A.), 1R21HG010683 (N.A.) and 5F30HG009479 (J.K.); National Institute of Mental Health grants 1R01MH109907 (N.A.) and 1U01MH116438 (N.A.); NRSA National Institutes of Health fellowship 5T32HL007093 (V.A.); and the Uehara Memorial Foundation (F.I.). J.S. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
J.K. and A.K. performed all cloning and sequencing for the nine assays and all experimental work for orientation and length sections. J.K. and J.S. conceived the HMPA protocol, and J.K. and A.K. developed and optimized it. A.K. produced schematic figures. M.K. developed the initial MPRA analysis pipeline. V.A. performed the computational analyses and generated all remaining figures and tables. F.I. performed the transfections and lentiviral transductions for the nine assays, carried out luciferase reporter experiments and wrote the associated methods sections. B.M. designed cloning steps and guided the development and testing of the MPRA assays. J.K., V.A., N.A. and J.S. wrote the remainder of the paper. N.A. and J.S. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
V.A. is an employee of Calico Life Sciences LLC.
Additional information
Peer review information Lei Tang was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Notes 1–3 and Figs. 1–17.
Supplementary Table 1
Genomic coordinates (human genome build hg19) and sequences for all designed elements, both naturally occurring as well as synthetic positive and negative controls, in the experiments testing the nine assays, element orientation and element size.
Supplementary Table 2
Activity scores computed for each element for each of the nine MPRA assays tested as well as HSS_full, HSS_b2, ORI_full and ORI_b2. Provided are averaged activity scores across replicates as well as individual scores for each replicate alongside normalized DNA counts, normalized RNA counts and the number of barcodes per element.
Supplementary Table 3
Summary of 915 features considered in a model trained to predict enhancer activity, with an overview of features considered, feature type (computationally predicted or experimentally derived), data source and number of features in the category.
Supplementary Table 4
Definition of each feature considered in the lasso regression models, with detailed metadata corresponding to the data source of origin, species of origin, sample accession IDs and additional factor-specific information. Also provided are pre-computed tables of the features used during training for the nine MPRA assays as well as the assay testing different size classes.
Supplementary Table 5
Coefficients fit for the full lasso regression models for each of eight MPRA assays shown in Supplementary Fig. 6, differential comparisons shown in Supplementary Figs. 7 and 17, the assay testing different size classes shown in Supplementary Fig. 14 and the corresponding differential pairwise comparisons shown in Supplementary Fig. 15.
Supplementary Table 6
Activity scores computed for each element in the forward (‘F’) and reverse (‘R’) orientations in the orientation assay. Provided are averaged activity scores across replicates as well as individual scores for each replicate alongside normalized DNA counts, normalized RNA counts and the number of barcodes per element.
Supplementary Table 7
Activity scores computed for each element in the short, medium and long elements in the assay testing for different size classes. Provided are averaged activity scores across replicates as well as individual scores for each replicate alongside normalized DNA counts, normalized RNA counts and the number of barcodes per element.
Supplementary Table 8
All primer, adaptor and oligonucleotide sequences utilized throughout the manuscript (excluding HMPA). When applicable, this includes the assay and step for which the primer was used.
Supplementary Table 9
All sequence indexes used for each experiment.
Supplementary Table 10
All primer and adaptor sequences used for HMPA.
Rights and permissions
About this article
Cite this article
Klein, J.C., Agarwal, V., Inoue, F. et al. A systematic evaluation of the design and context dependencies of massively parallel reporter assays. Nat Methods 17, 1083–1091 (2020). https://doi.org/10.1038/s41592-020-0965-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0965-y
This article is cited by
-
Proformer: a hybrid macaron transformer model predicts expression values from promoter sequences
BMC Bioinformatics (2024)
-
Massively parallel screen uncovers many rare 3′ UTR variants regulating mRNA abundance of cancer driver genes
Nature Communications (2024)
-
Integrative functional genomic analyses identify genetic variants influencing skin pigmentation in Africans
Nature Genetics (2024)
-
Single-cell spatial multi-omics and deep learning dissect enhancer-driven gene regulatory networks in liver zonation
Nature Cell Biology (2024)
-
Functional dissection of human cardiac enhancers and noncoding de novo variants in congenital heart disease
Nature Genetics (2024)