Abstract
Single-cell RNA sequencing offers snapshots of whole transcriptomes but obscures the temporal RNA dynamics. Here we present single-cell metabolically labeled new RNA tagging sequencing (scNT-seq), a method for massively parallel analysis of newly transcribed and pre-existing mRNAs from the same cell. This droplet microfluidics-based method enables high-throughput chemical conversion on barcoded beads, efficiently marking newly transcribed mRNAs with T-to-C substitutions. Using scNT-seq, we jointly profiled new and old transcriptomes in ~55,000 single cells. These data revealed time-resolved transcription factor activities and cell-state trajectories at the single-cell level in response to neuronal activation. We further determined rates of RNA biogenesis and decay to uncover RNA regulatory strategies during stepwise conversion between pluripotent and rare totipotent two-cell embryo (2C)-like stem cell states. Finally, integrating scNT-seq with genetic perturbation identifies DNA methylcytosine dioxygenase as an epigenetic barrier into the 2C-like cell state. Time-resolved single-cell transcriptomic analysis thus opens new lines of inquiry regarding cell-type-specific RNA regulatory mechanisms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All sequencing data associated with this study have been deposited in the Gene Expression Omnibus (GEO) database under accession code GSE141851. Source data are provided with this paper.
Code availability
The analysis source code underlying the final version of this paper are available on the GitHub repository (https://github.com/wulabupenn/scNT-seq/).
References
Rabani, M. et al. Metabolic labeling of RNA uncovers principles of RNA production and degradation dynamics in mammalian cells. Nat. Biotechnol. 29, 436–442 (2011).
Rabani, M. et al. High-resolution sequencing and modeling identifies distinct dynamic RNA regulatory strategies. Cell 159, 1698–1710 (2014).
Tanay, A. & Regev, A. Scaling single-cell genomics from phenomenology to mechanism. Nature 541, 331–338 (2017).
Herzog, V. A. et al. Thiol-linked alkylation of RNA to assess expression dynamics. Nat. Methods 14, 1198–1204 (2017).
Schofield, J. A., Duffy, E. E., Kiefer, L., Sullivan, M. C. & Simon, M. D. TimeLapse-seq: adding a temporal dimension to RNA sequencing through nucleoside recoding. Nat. Methods 15, 221–225 (2018).
Riml, C. et al. Osmium-mediated transformation of 4-thiouridine to cytidine as key to study RNA dynamics by sequencing. Angew. Chem. Int. Ed. Engl. 56, 13479–13483 (2017).
Erhard, F. et al. scSLAM-seq reveals core features of transcription dynamics in single cells. Nature 571, 419–423 (2019).
Hendriks, G. J. et al. NASC-seq monitors RNA synthesis in single cells. Nat. Commun. 10, 3138 (2019).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Hu, P. et al. Dissecting cell-type composition and activity-dependent transcriptional state in mammalian brains by massively parallel single-nucleus RNA-seq. Mol. Cell 68, 1006–1015 (2017).
Hu, P. et al. Single-nucleus transcriptomic survey of cell diversity and functional maturation in postnatal mammalian hearts. Genes Dev. 32, 1344–1357 (2018).
Lake, B. B. et al. Integrative single-cell analysis of transcriptional and epigenetic states in the human adult brain. Nat. Biotechnol. 36, 70–80 (2018).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Qiu, Q. et al. Massively parallel and time-resolved RNA sequencing in single cells with scNT-seq. Protocol Exchange https://doi.org/10.21203/rs.3.pex-1019/v1 (2020).
Yap, E. L. & Greenberg, M. E. Activity-regulated transcription: bridging the gap between neural activity and behavior. Neuron 100, 330–348 (2018).
Tyssowski, K. M. et al. Different neuronal activity patterns induce different gene expression programs. Neuron 98, 530–546 (2018).
Duffy, E. E., Schofield, J. A. & Simon, M. D. Gaining insight into transcriptome-wide RNA population dynamics through the chemistry of 4-thiouridine. Wiley Interdiscip. Rev. RNA 10, e1513 (2019).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Kannan, M. B., Solovieva, V. & Blank, V. The small MAF transcription factors MAFF, MAFG and MAFK: current knowledge and perspectives. Biochim. Biophys. Acta 1823, 1841–1846 (2012).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Schwalb, B. et al. TT-seq maps the human transient transcriptome. Science 352, 1225–1228 (2016).
Qiu, X. et al. Mapping vector field of single cells. Preprint at bioRxiv https://doi.org/10.1101/696724 (2019).
Kolodziejczyk, A. A. et al. Single cell RNA-sequencing of pluripotent states unlocks modular transcriptional variation. Cell Stem Cell 17, 471–485 (2015).
Macfarlan, T. S. et al. Embryonic stem cell potency fluctuates with endogenous retrovirus activity. Nature 487, 57–63 (2012).
Eckersley-Maslin, M. A. et al. MERVL/Zscan4 network activation results in transient genome-wide DNA demethylation of mESCs. Cell Rep. 17, 179–192 (2016).
Fu, X., Wu, X., Djekidel, M. N. & Zhang, Y. Myc and Dnmt1 impede the pluripotent to totipotent state transition in embryonic stem cells. Nat. Cell Biol. 21, 835–844 (2019).
Lu, F., Liu, Y., Jiang, L., Yamaguchi, S. & Zhang, Y. Role of Tet proteins in enhancer activity and telomere elongation. Genes Dev. 28, 2103–2119 (2014).
Russo, J., Heck, A. M., Wilusz, J. & Wilusz, C. J. Metabolic labeling and recovery of nascent RNA to accurately quantify mRNA stability. Methods 120, 39–48 (2017).
Battich, N. et al. Sequencing metabolically labeled transcripts in single cells reveals mRNA turnover strategies. Science 367, 1151–1156 (2020).
Wu, H. & Zhang, Y. Reversing DNA methylation: mechanisms, genomics and biological functions. Cell 156, 45–68 (2014).
Wu, H. et al. Dual functions of Tet1 in transcriptional regulation in mouse embryonic stem cells. Nature 473, 389–393 (2011).
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR–Cas-mediated genome engineering. Cell 153, 910–918 (2013).
Hughes, T. K. et al. Highly efficient, massively-parallel single-cell RNA-seq reveals cellular states and molecular features of human skin pathology. Preprint at bioRxiv https://doi.org/10.1101/689273 (2019).
Cao, J., Zhou, W., Steemers, F., Trapnell, C. & Shendure, J. Sci-fate characterizes the dynamics of gene expression in single cells. Nat. Biotechnol. https://doi.org/10.1038/s41587-020-0480-9 (2020).
Gupta, I. et al. Single-cell isoform RNA sequencing characterizes isoforms in thousands of cerebellar cells. Nat. Biotechnol. https://doi.org/10.1038/nbt.4259 (2018).
Kiefer, L., Schofield, J. A. & Simon, M. D. Expanding the nucleoside recoding toolkit: revealing RNA population dynamics with 6-thioguanosine. J. Am. Chem. Soc. 140, 14567–14570 (2018).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Schutsky, E. K. et al. Nondestructive, base-resolution sequencing of 5-hydroxymethylcytosine using a DNA deaminase. Nat. Biotechnol. 36, 1083–1090 (2018).
Alles, J. et al. Cell fixation and preservation for droplet-based single-cell transcriptomics. BMC Biol. 15, 44 (2017).
Chen, J. et al. PBMC fixation and processing for Chromium single-cell RNA sequencing. J. Transl. Med. 16, 198 (2018).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies and species. Nat. Biotechnol. 36, 411–420 (2018).
McInnes, L., Healy, J. & Melville, J. UMAP: uniform manifold approximation and projection for dimension reduction. Preprint at https://arxiv.org/abs/1802.03426 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Jurges, C., Dolken, L. & Erhard, F. Dissecting newly transcribed and old RNA using GRAND-SLAM. Bioinformatics 34, i218–i226 (2018).
Petukhov, V. et al. DropEst: pipeline for accurate estimation of molecular counts in droplet-based single-cell RNA-seq experiments. Genome Biol. 19, 78 (2018).
Acknowledgements
We are grateful to all members of the Wu laboratory for helpful discussion. This work was supported by the Penn Epigenetics Institute, the National Human Genome Research Institute (NHGRI; grants R00-HG007982 and R01-HG010646), National Heart Lung and Blood Institute (NHLBI grant DP2-HL142044) and National Cancer Institute (NCI; grant U2C-CA233285 to H.W.). The work of P.G.C. is partially supported by Stand Up To Cancer Convergence 2.0.
Author information
Authors and Affiliations
Contributions
H.W., Q.Q. and P.H. conceived and developed the scNT-seq method. Q.Q. conducted most of the experiments. P.H. performed most of the computational analysis and generated the Tet-TKO mESC line. X.Q. contributed to RNA velocity analysis. K.W.G. and P.G.C. contributed to the statistical modeling. Q.Q., P.H. and H.W. analyzed the results and wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Lei Tang was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Performance and quality control analyses of scNT-seq.
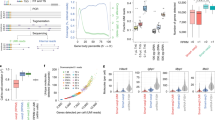
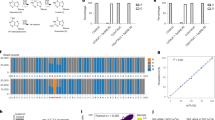
a. Scatterplots showing the number of detected gene per cell (y-axis, upper panel) or UMI per cell (y-axis, lower panel) as a function of aligned reads per cell (x-axis) between 4sU (red, 462 cells), TFEA (blue, 211 cells), and 4sU/TFEA (green, 578 cells) experiments. 4sU, cells labeled with 4sU (100 μM, 4 hours (h)). TFEA, beads treated with TFEA/NaIO4 chemical reaction. 4sU/TFEA, cells labeled with 4sU and beads treated with TFEA/NaIO4 chemical reaction. The fitted lines of different experiments were shown. The estimated numbers of gene or UMI detected per cell at matching sequencing depth (50,000 aligned read per cell) are shown on the right. b. Shown are all transcripts (with unique UMIs) for the ACTG1 gene from one untreated control K562 cell (upper panel) and one TFEA/NaIO4-treated cell (lower panel). Grey circles denote uridine sites without T-to-C substitution, and “X”s denote sites with T-to-C substitutions. The read coverage for each T-to-C substitution is color-scaled. All 9 sequencing reads of the 2nd UMI (in red box) from the TFEA/NaIO4-treated cell are highlighted below. c. Bar plot showing nucleotide substitution rates in mESCs with different labeling parameters (100 μM 4sU for 4 h or 200 μM 4sU for 24 h) and sample processing methods (freshly isolated versus methanol fixation followed by cyro-preservation and rehydration with two different rehydration buffers: PBS-based versus SSC-based). A sample (100 μM 4sU for 4 h) that was not treated with TFEA/NaIO4 served as the control. d. Scatterplots showing Pearson’s correlation between two biologically independent replicates of mESCs (rep1: 427 cells and rep2: 733 cells). The expression levels of new (n = 10,925 genes) and old (n = 14,496 genes) transcripts were quantified as natural log transformation of (TP10K + 1).
Extended Data Fig. 2 Cell-type clustering and analysis of activity-dependent gene expression programs in mouse cortical neuronal cultures.
a. Experimental scheme of characterizing neuronal activation in primary mouse cortical cultures with scNT-seq. Cells were treated with KCl from 15 min to 120 min. Cells from all treatment conditions were labeled with 4sU for 2 h before harvest for scNT-seq. b. Left, UMAP plot for 20,547 cells from mouse cortical cultures (the same UMAP plot in Fig. 2a). The cells are colored by different time points of neuronal activation. Right, violin plot showing the distribution of total RNA levels for representative cell-type specific marker genes. c. Clustered heat map showing new RNA levels (z-scaled natural log transformation of (TP10K + 1)) of neuronal activity induced genes across different cell-types. d. Clustered heat map showing new RNA levels (z-scaled natural log transformation of (TP10K + 1)) of early- and late-response genes in excitatory neurons with different durations of KCl stimulation. 97 significantly induced genes were clustered into two groups (early- and late-response). The expression levels of early- and late-response genes are in Supplementary Table 2. e. Venn diagram showing a significant overlap between Maff and Fosb regulon targets (243 genes, P-value = 1.64 x 10-164, Two-sided Fisher’s exact test).
Extended Data Fig. 3 UMI-based statistical correction of newly-transcribed RNA fraction.
a. Density plot showing the distribution of number of covered uridine sites per read (60 bp) or per UMI (UMI-linked transcript) in excitatory neurons with 60 min of KCl stimulation. b. Bar plot of the number of T-to-C substitutions per read (60 bp) or UMI. Shown is the analysis of excitatory neurons with 60 min of KCl stimulation. c. Shown are all unique transcripts (with unique UMIs) of the Fos (an activity-induced gene) and Mapt (a slow turnover housekeeping gene) from a single excitatory neuron with 60 min of KCl stimulation. Grey circles represent uridines without T-to-C conversion, while crosses (“X”s) denote uridines with T-to-C substitution in at least one read. The read coverage for each T-to-C substitution is color-scaled. d. Comparison of uncorrected and statistically corrected new RNA levels of each detected gene (n=9,082 genes) in excitatory neurons (with 60 min of KCl stimulation). Four representative activity-induced genes (Fos, Jun, Egr1, and Npas4) and two housekeeping genes (Mapt and Actb) are highlighted with red circles. e. Scatter plot showing the new transcript fraction (over total RNAs; y-axis) of excitatory neurons with 60 min of KCl stimulation as a function of differential gene expression (between 60 min and 0 min of KCl stimulation; x-axis). Two-sided Wilcoxon rank sum test was used to assess significance of the difference, and the P-value was adjusted by Bonferroni correction. Genes were color-coded by statistical significance of differential gene expression. The fraction of new transcripts, expression fold-change, and adjusted P-value of each gene are in Source Data Extended Data Fig. 3.
Extended Data Fig. 4 scNT-seq enables metabolic labeling-based time-resolved RNA velocity in excitatory neurons.
a. UMAP visualization of excitatory neurons (13,511 cells, with >500 genes detected per cell) that were characterized by standard splicing kinetics-based (left) or metabolic labeling based RNA velocity (right) analyses. Cells are color-coded by time points. The streamlines indicate the integration paths that connect local projections from the observed state to extrapolated future state. The thickness of streamline indicates the magnitude of velocity. UMAP plots in lower panels (same as upper panels) show randomized velocity controls for splicing (left) or metabolic labeling (right) based RNA velocity. Permutation of velocity flows was performed by shuffling velocity for genes in each cell and then randomly flipping the sign of shuffled velocity values. b. UMAP (same as right of a) visualization of Ex neurons colored by the average new RNA expression level (natural log transformation of (TP10K + 1)) of 24 early- (left) or 73 late-response (right) genes. c. UMAP (same as right of a) showing Ex neurons colored by the regulon activity of three representative TFs (Jun, Mef2d, and Maff).
Extended Data Fig. 5 Comparison of splicing-based and metabolic labeling-based RNA velocity analysis methods.
The excitatory neurons (n=3,066 cells, with >2,000 genes detected per cell) were analyzed by either splicing kinetics-based a, or metabolic labeling-based b, RNA velocity. Shown are the phase portraits (left), UMAP plots colored by smoothed spliced (in a) or total (in b) RNA level based on local averaging (middle), and RNA velocity values (right) of three representative activity-induced genes (Egr1, Fos and Homer1).
Extended Data Fig. 6 Quality control for metabolic labeling based RNA velocity analysis.
a. UMAP (as in right panels of Fig. 3a) visualization of high-quality Ex neurons (3,066 cells, >2,000 genes detected per cell) colored by time points (left), number of gene detected (middle), and number of UMI detected per cell (right). b. UMAP (as in right panels of Fig. 3a) visualization of high-quality Ex neurons colored by the new RNA levels (natural log transformation of (TP10K + 1)) of six representative genes, including three early-response genes (Egr1, Fos, Jun) and three late-response genes (Homer1, Gadd45g, Nr4a2). c. UMAP (as in right panels of Extended Data Fig. 4a) visualization of all Ex neurons (13,511 cells, >500 genes detected per cell) colored by time points (left), number of gene detected (middle), and number of UMI detected per cell (right). d. UMAP (as in right panels of Extended Data Fig. 4a) visualization of all Ex neurons colored by the new RNA levels (natural log transformation of (TP10K + 1)) of six representative genes (same as in b).
Extended Data Fig. 7 scNT-seq reveals different stem cell states in mESC cultures.
a. UMAP visualization of 4,633 WT cells (from two biological replicates) colored by different cell-types or cell-states. Feeders are contaminating mouse embryonic fibroblasts. b. UMAP visualization of two biological replicates in (a). c. Violin plots showing total RNA levels (natural log transformation of (TP10K + 1)) of representative marker genes for feeders or specific stem cell states. d. UMAP (same as in (a)) visualization of cells colored by total RNA levels (natural log transformation of (TP10K + 1)) of four representative marker genes. e. Violin plots showing both new and old RNA levels (natural log transformation of (TP10K + 1)) of selected genes across three stem cell states.
Extended Data Fig. 8 Pulse-chase scNT-seq reveals state-specific mRNA half-life.
a. Violin plots showing levels of labeled and total transcripts of two representative genes (Sox2 and Top2a) during pulse-chase assay. The expression level is measured in natural log transformation of (TP10K + 1). b. Enrichment analysis of GO terms within stable (top 10% genes with longest half-lives) and unstable genes (top 10% genes with shortest half-life) in pluripotent state mESCs. Enrichment analysis was performed via a one-sided hypergeometric test. P-value was then corrected by FDR. The P-values of GO terms are in Source Data Extended Data Fig. 8. c. Clustered heat map showing the mRNA half-life of 2,616 genes across three stem cell states. These genes are clustered to six groups based on the scaled RNA half-lives in three cell states. The state-specific half-lives are in Supplementary Table 4. d. Shown are mRNA decay curves of representative genes from each group. The fraction of labeled transcripts was calculated for each time point and normalized to chase (0 h), then fit to a single-exponential decay model to derive RNA half-lives (t1/2).
Extended Data Fig. 9 scNT-seq analysis of the pluripotent-to-2C transition in mESCs.
a. Scatter MA-plot showing differential expression of new, old, and total RNAs between pluripotent and 2C-like states. Dashed line denotes 1.5-fold change between states. b. Heat map showing enriched GO terms for state-specific genes. Significance of enrichment (FDR) is scaled by colors. Enrichment analysis was performed using a one-sided hypergeometric test. P-value was then corrected by FDR. The exact P-values of GO terms are in Source Data Extended Data Fig. 9. c. Normalized new and old RNA levels (natural log transformation of (TP10K + 1)) of major DNA methylation regulators across three stem cell states. d. Validation of genotypes of the Tet1 (-11bp/+1bp) and Tet2 (-7bp/-1bp) genes in Tet-TKO cells by aligning scNT-seq reads to the CRISPR-Cas9 genome editing sites. e. UMAP visualization (same as in Fig. 5d) of mESCs colored by cell-cycle states (left) or the new RNA level (natural log transformation of (TP10K + 1)) of Zscan4a (right). f. Venn diagrams showing significant overlap between Tet1 and Myc regulon target genes (upper) (P-value = 2.42 x 10-25, two-sided Fisher’s exact test), and between Tet1 and Max regulon target genes (lower) (P-value = 1.96 x 10-62, two-sided Fisher’s exact test).
Extended Data Fig. 10 Benchmarking the 2nd SS scNT-seq protocol in human K562 cells.
a. Bar plot showing nucleotide substitution rates in K562 cells analyzed with different experimental protocols. 4sU, metabolic labeling with 4sU (100 μM, 4 h); TFEA, on-bead TFEA/NaIO4 chemical reaction; 2nd SS, second strand synthesis. b. PCA plots showing K562 cells colored by the total RNA level of the TOP2A gene (natural log transformation of (TP10K + 1)) in three experimental protocols (same as in Fig. 6d). c. Violin plots showing the new-to-total RNA ratios of 8 representative cell-cycle genes in datasets generated by 2nd SS (4sU/TFEA/2nd SS, n =795 cells) or standard (4sU/TFEA, n = 533 cell) scNT-seq protocols. See ‘Data visualization’ in the Methods for definitions of box-plot elements. d. Same as in c but showing new and old RNA levels (natural log transformation of (TP10K + 1)) in three cell-cycle states (G1/S/G2M).
Supplementary information
Supplementary Information
Supplementary protocol for scNT-seq
Supplementary Table
Supplementary Table 1: Summary of samples and sequencing experiments. Supplementary Table 2: Early- and late-response genes in excitatory neurons. Supplementary Table 3: Cell-type-specific regulon activity of 79 TFs in response to distinct activity durations. Supplementary Table 4: State-specific mRNA half-life of 2,616 genes in mESCs. Supplementary Table 5: Synthesis/degradation rates and observed total RNA levels of 445 state-specific genes in three stem cell states of mESCs. Supplementary Table 6: List of genes differentially expressed between WT and Tet-TKO mESCs. Supplementary Table 7: Comparison of different single-cell metabolic labeling-based RNA-sequencing methods.
Source data
Source Data Fig. 2
Statistical source data for Fig. 2e.
Source Data Fig. 3
Statistical source data for Fig. 3c.
Source Data Fig. 5
Statistical source data for Fig. 5g.
Source Data Extended Data Fig. 3
Statistical source data for Extended Data Fig. 3e.
Source Data Extended Data Fig. 8
Statistical source data for Extended Data Fig. 8b.
Source Data Extended Data Fig. 9
Statistical source data for Extended Data Fig. 9b.
Rights and permissions
About this article
Cite this article
Qiu, Q., Hu, P., Qiu, X. et al. Massively parallel and time-resolved RNA sequencing in single cells with scNT-seq. Nat Methods 17, 991–1001 (2020). https://doi.org/10.1038/s41592-020-0935-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0935-4
This article is cited by
-
Spatiotemporal multi-omics: exploring molecular landscapes in aging and regenerative medicine
Military Medical Research (2024)
-
DeepKINET: a deep generative model for estimating single-cell RNA splicing and degradation rates
Genome Biology (2024)
-
Powerful and accurate detection of temporal gene expression patterns from multi-sample multi-stage single-cell transcriptomics data with TDEseq
Genome Biology (2024)
-
Cell-type-specific mRNA transcription and degradation kinetics in zebrafish embryogenesis from metabolically labeled single-cell RNA-seq
Nature Communications (2024)
-
Dissecting key regulators of transcriptome kinetics through scalable single-cell RNA profiling of pooled CRISPR screens
Nature Biotechnology (2024)