Abstract
Photobleaching limits extended imaging of fluorescent biological samples. We developed DNA-based ‘FluoroCubes’ that are similar in size to the green fluorescent protein, have single-point attachment to proteins, have a ~54-fold higher photobleaching lifetime and emit ~43-fold more photons than single organic dyes. We demonstrate that DNA FluoroCubes provide outstanding tools for single-molecule imaging, allowing the tracking of single motor proteins for >800 steps with nanometer precision.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
Example raw datasets of DNA FluoroCubes with six dyes, single dye cubes (FluoroCubes with a single dye), one-dye dsDNA constructs, single, biotinylated dyes and compact cubes used to determine the photophysical properties are hosted on Zenodo: https://doi.org/10.5281/zenodo.3561024 (ref. 36). All other data files are available from the authors upon request.
Code availability
μManager acquisition and analysis software is available partly under the Berkeley Software Distribution license, partly under the GNU Lesser General Public License and development is hosted on GitHub at https://github.com/nicost/micro-manager. The latest version for MacOS and Windows can be downloaded here: https://micro-manager.org/wiki/Download%20Micro-Manager_Latest%20Release (v.2.0 gamma). The custom written Python code used to determine the total number of photons, the average number of photons per frame, and the half-life (photostability) for all fluorescent samples used in this study is hosted on Zenodo at https://doi.org/10.5281/zenodo.3629746 (ref. 37). All other code is available from the authors upon request.
References
Joo, C., Balci, H., Ishitsuka, Y., Buranachai, C. & Ha, T. Advances in single-molecule fluorescence methods for molecular biology. Annu. Rev. Biochem. 77, 51–76 (2008).
Resch-Genger, U., Grabolle, M., Cavaliere-Jaricot, S., Nitschke, R. & Nann, T. Quantum dots versus organic dyes as fluorescent labels. Nat. Methods 5, 763–775 (2008).
Zheng, Q. et al. Ultra-stable organic fluorophores for single-molecule research. Chem. Soc. Rev. 43, 1044–1056 (2014).
Shaner, N. C., Steinbach, P. A. & Tsien, R. Y. A guide to choosing fluorescent proteins. Nat. Methods 2, 905–909 (2005).
Wichner, S. M. et al. Covalent protein labeling and improved single-molecule optical properties of aqueous CdSe/CdS quantum dots. ACS Nano 11, 6773–6781 (2017).
Wolfbeis, O. S. An overview of nanoparticles commonly used in fluorescent bioimaging. Chem. Soc. Rev. 44, 4743–4768 (2015).
Schröder, T., Scheible, M. B., Steiner, F., Vogelsang, J. & Tinnefeld, P. Interchromophoric interactions determine the maximum brightness density in DNA origami structures. Nano Lett. 19, 1275–1281 (2019).
Woehrstein, J. B. et al. Sub–100-nm metafluorophores with digitally tunable optical properties self-assembled from DNA. Sci. Adv. 3, e1602128 (2017).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Douglas, S. M. et al. Rapid prototyping of 3D DNA-origami shapes with caDNAno. Nucleic Acids Res. 37, 5001–5006 (2009).
Ke, Y., Ong, L. L., Shih, W. M. & Yin, P. Three-dimensional structures self-assembled from DNA bricks. Science 338, 1177–1183 (2012).
Wei, B., Dai, M. & Yin, P. Complex shapes self-assembled from single-stranded DNA tiles. Nature 485, 623–626 (2012).
Scheible, M. B. et al. A compact DNA cube with side length 10 nm. Small 11, 5200–5205 (2015).
Zheng, Q. et al. Electronic tuning of self-healing fluorophores for live-cell and single-molecule imaging. Chem. Sci. 8, 755–762 (2017).
Grimm, J. B. et al. A general method to improve fluorophores for live-cell and single-molecule microscopy. Nat. Methods 12, 244–250 (2015).
Aitken, C. E., Marshall, R. A. & Puglisi, J. D. An oxygen scavenging system for improvement of dye stability in single-molecule fluorescence experiments. Biophys. J. 94, 1826–1835 (2008).
Rasnik, I., McKinney, S. A. & Ha, T. Nonblinking and long-lasting single-molecule fluorescence imaging. Nat. Methods 3, 891–893 (2006).
Nicoli, F. et al. Proximity-induced H-aggregation of cyanine dyes on DNA-duplexes. J. Phys. Chem. A 120, 9941–9947 (2016).
Tomishige, M., Klopfenstein, D. R. & Vale, R. D. Conversion of Unc104/KIF1A kinesin into a processive motor after dimerization. Science 297, 2263–2267 (2002).
Okada, Y., Higuchi, H. & Hirokawa, N. Processivity of the single-headed kinesin KIF1A through biased binding to tubulin. Nature 424, 574–577 (2003).
Yildiz, A., Tomishige, M., Vale, R. D. & Selvin, P. R. Kinesin walks hand-over-hand. Science 303, 676–678 (2004).
Stepp, W. L., Merck, G., Mueller-Planitz, F. & Ökten, Z. Kinesin-2 motors adapt their stepping behavior for processive transport on axonemes and microtubules. EMBO Rep. 18, 1947–1956 (2017).
Reddy, B. J. N. et al. Heterogeneity in kinesin function. Traffic 18, 658–671 (2017).
Ozhalici-Unal, H. & Armitage, B. A. Fluorescent DNA nanotags based on a self-assembled DNA tetrahedron. ACS Nano 3, 425–433 (2009).
Kretschy, N., Sack, M. & Somoza, M. M. Sequence-dependent fluorescence of Cy3- and Cy5-labeled double-stranded DNA. Bioconjug. Chem. 27, 840–848 (2016).
Femino, A. M., Fay, F. S., Fogarty, K. & Singer, R. H. Visualization of single RNA transcripts in situ. Science 280, 585–590 (1998).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. RNA imaging. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Jungmann, R. et al. Multiplexed 3D cellular super-resolution imaging with DNA-PAINT and Exchange-PAINT. Nat. Methods 11, 313–318 (2014).
Niekamp, S. et al. Nanometer-accuracy distance measurements between fluorophores at the single-molecule level. Proc. Natl Acad. Sci. USA 116, 4275–4284 (2019).
Los, G. V. et al. HaloTag: a novel protein labeling technology for cell imaging and protein analysis. ACS Chem. Biol. 3, 373–382 (2008).
Niekamp, S., Stuurman, N. & Ronald, D. Folding of FluoroCubes v.1 (protocols.io.8k2huye, 2019); https://doi.org/10.17504/protocols.io.8k2huye
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Edelstein, A., Amodaj, N., Hoover, K., Vale, R. & Stuurman, N. Computer control of microscopes using µManager. Curr. Protoc. Mol. Biol. Ch.14, Unit14.20 (2010).
Qiu, W. et al. Dynein achieves processive motion using both stochastic and coordinated stepping. Nat. Struct. Mol. Biol. 19, 193–200 (2012).
Niekamp, S., Stuurman, N. & Vale, R. D. A 6-nm Ultra-Photostable DNA FluoroCube for Fluorescence Imaging (Zenodo, 2019); https://doi.org/10.5281/ZENODO.3561024
Niekamp, S., Stuurman, N. & Vale, R. D. Python code for a 6-nm Ultra-Photostable DNA FluoroCube for Fluorescence Imaging (Zenodo, 2020); https://doi.org/10.5281/zenodo.3629746
Acknowledgements
We are grateful to J. Sung (UCSF) for critical discussions of the manuscript. We thank D. Mullins (UCSF) for teaching us how to use the ISS K2 multifrequency fluorometer. We are thankful to Y.-W. Jun and Y. Zhao (UCSF) for teaching us how to use the Malvern Zetasizer ZS90. We thank L. Lavis (Janelia Research Campus) for the suggestion of comparing sulfonated versus nonsulfonated dyes and for providing the JF549 and JF646 dyes. We are thankful to S. Douglas (UCSF) and A.G. York (Calico Laboratories) for feedback on the manuscript after preprinting. A. Carter (MRC Laboratory of Molecular Biology) and E. Villa (UCSD) supplied the MATLAB script for step detection of kinesin. We acknowledge funding from the National Institutes of Health grant nos. R01GM097312 and 1R35GM118106 (to R.D.V. and S.N.), and the Howard Hughes Medical Institute (to R.D.V., S.N. and N.S.).
Author information
Authors and Affiliations
Contributions
S.N., N.S. and R.D.V. designed the research. S.N. prepared samples, collected TIRF microscopy data and electron microscopy data and analyzed it. S.N., N.S. and R.D.V. wrote the article. All authors read and commented on the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Rita Strack was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–21, Tables 1–13 and Supplementary Protocol.
Supplementary Video 1
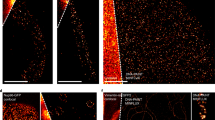
Single-molecule imaging of ATTO 488 six-dye FluoroCubes and ATTO 488 one-dye dsDNA. The movie shows surface-immobilized single molecules of ATTO 488 six-dye FluoroCubes and ATTO 488 one-dye dsDNA over ~50 min under continuous laser illumination. The graph shows the photostability of six-dye FluoroCubes and one-dye dsDNA. The survival rate was quantified by counting the percentage of probes in the ‘on’ state at any given time from 0 to 3,000 s. Time is in minutes. Each experiment was repeated five or four times with freshly assembled six-dye FluoroCubes or one-dye dsDNA, respectively, and on freshly prepared microscope slides. All acquired movies looked very similar and grave consistent results (see Fig. 1 l–n).
Supplementary Video 2
Single-molecule imaging of ATTO 565 six-dye FluoroCubes and ATTO 565 one-dye dsDNA. The movie shows surface-immobilized single molecules of ATTO 565 six-dye FluoroCubes and ATTO 565 one-dye dsDNA over ~50 min under continuous laser illumination. The graph shows the photostability of six-dye FluoroCubes and one-dye dsDNA. The survival rate was quantified by counting the percentage of probes in the ‘on’ state at any given time from 0 to 3,000 s. Time is in minutes. Each experiment was repeated five or four times with freshly assembled six-dye FluoroCubes or one-dye dsDNA, respectively, and on freshly prepared microscope slides. All acquired movies looked very similar and grave consistent results (see Fig. 1 l–n).
Supplementary Video 3
Single-molecule imaging of ATTO 647N six-dye FluoroCubes and ATTO 647N one-dye dsDNA The movie shows surface-immobilized single molecules of ATTO 647N six-dye FluorocCubes and ATTO 647N one-dye dsDNA over ~50 min under continuous laser illumination. The graph shows the photostability of six-dye FluoroCubes and one-dye dsDNA. The survival rate was quantified by counting the percentage of probes in the ‘on’ state at any given time from 0 to 3,000 s. Time is in minutes. Each experiment was repeated five or four times with freshly assembled six-dye FluoroCubes or one-dye dsDNA, respectively, and on freshly prepared microscope slides. All acquired movies looked very similar and grave consistent results (see Fig. 1l–n)
Supplementary Video 4
Single-molecule imaging of Cy3 six-dye FluoroCubes and Cy3 one-dye dsDNA; The movie shows surface-immobilized single molecules of Cy3 six-dye FluoroCubes and Cy3 one-dye dsDNA over ~50 min under continuous laser illumination. The graph shows the photostability of six-dye FluoroCubes and one-dye dsDNA. The survival rate was quantified by counting the percentage of probes in the ‘on’ state at any given time from 0 to 3,000 s. Time is in minutes. Each experiment was repeated five or four times with freshly ssembled six-dye FluoroCubes or one-dye dsDNA, respectively, and on freshly prepared microscope slides. All acquired movies looked very similar and grave consistent results (see Fig. 1l–n).
Supplementary Video 5
Single-molecule imaging of Cy5 six-dye FluoroCubes and Cy5 one-dye dsDNA The movie shows surface-immobilized single molecules of Cy5 six-dye FluoroCubes and Cy5 one-dye dsDNA over ~50 min under continuous laser illumination. The graph shows the photostability of six-dye FluoroCubes and one-dye dsDNA. The survival rate was quantified by counting the percentage of probes in the ‘on’ state at any given time from 0 to 3,000 s. Time is in minutes. Each experiment was repeated five or four times with freshly assembled six-dye FluoroCubes or one-dye dsDNA, respectively, and on freshly prepared microscope slides. All acquired movies looked very similar and grave consistent results (see Fig. 1l–n)
Supplementary Video 6
Single-molecule imaging of Cy3N six-dye FluoroCubes and Cy3N one-dye dsDNA The movie shows surface-immobilized single molecules of Cy3N six-dye FluoroCubes and Cy3N one-dye dsDNA over ~50 min under continuous laser illumination. The graph shows the photostability of six-dye FluoroCubes and one-dye dsDNA. The survival rate was quantified by counting the percentage of probes in the ‘on’ state at any given time from 0 to 3,000 s. Time is in minutes. Each experiment was repeated five or four times with freshly assembled six-dye FluoroCubes or one-dye dsDNA, respectively, and on freshly prepared microscope slides. All acquired movies looked very similar and grave consistent results (see Fig. 1l–n).
Supplementary Video 7
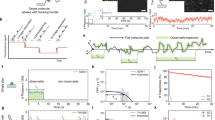
Movement of Kif1A along an axoneme The movie shows the movement of Kif1A along an axoneme. This particular molecule was tracked for the stepping trace shown in Fig. 2. Note that the axoneme is not visible. The movie plays with 30× real time. Time is in minutes. Scale bar is 1 μm. The imaging of kinesin stepping was repeated multiple times with very similar outcomes (see Supplementary Fig. 20). However, all traces were collected with motors from the same preparation but with freshly prepared microscope slides.
Rights and permissions
About this article
Cite this article
Niekamp, S., Stuurman, N. & Vale, R.D. A 6-nm ultra-photostable DNA FluoroCube for fluorescence imaging. Nat Methods 17, 437–441 (2020). https://doi.org/10.1038/s41592-020-0782-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0782-3
This article is cited by
-
Tracking single particles for hours via continuous DNA-mediated fluorophore exchange
Nature Communications (2021)
-
Advanced imaging and labelling methods to decipher brain cell organization and function
Nature Reviews Neuroscience (2021)
-
Turn-key mapping of cell receptor force orientation and magnitude using a commercial structured illumination microscope
Nature Communications (2021)