Abstract
Virtual realities are powerful tools to analyze and manipulate interactions between animals and their environment and to enable measurements of neuronal activity during behavior. In many species, however, optical access to the brain and/or the behavioral repertoire are limited. We developed a high-resolution virtual reality for head-restrained adult zebrafish, which exhibit cognitive behaviors not shown by larvae. We noninvasively measured activity throughout the dorsal telencephalon by multiphoton calcium imaging. Fish in the virtual reality showed regular swimming patterns and were attracted to animations of conspecifics. Manipulations of visuo-motor feedback revealed neurons that responded selectively to the mismatch between the expected and the actual visual consequences of motor output. Such error signals were prominent in multiple telencephalic areas, consistent with models of predictive processing. A virtual reality system for adult zebrafish therefore provides opportunities to analyze neuronal processing mechanisms underlying higher brain functions including decision making, associative learning, and social interactions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Original image series and VR presentations that support the findings of this study are too large to be included in the publication. These data are available from the corresponding author upon reasonable request.
Code availability
Software for controlling VR and coordinating VR with two-photon imaging can be downloaded at https://github.com/HUANGKUOHUA/Zebrafish-in-virtual-reality.git.
References
Holscher, C. Rats are able to navigate in virtual environments. J. Exp. Biol. 208, 561–569 (2005).
Harvey, C. D., Collman, F., Dombeck, D. A. & Tank, D. W. Intracellular dynamics of hippocampal place cells during virtual navigation. Nature 461, 941–946 (2009).
Minderer, M., Harvey, C. D., Donato, F. & Moser, E. I. Virtual reality explored. Nature 533, 324–325 (2016).
Reiser, M. B. & Dickinson, M. H. A modular display system for insect behavioral neuroscience. J. Neurosci. Methods 167, 127–139 (2008).
Kim, S. S., Rouault, H., Druckmann, S. & Jayaraman, V. Ring attractor dynamics in the Drosophila central brain. Science 356, 849–853 (2017).
Dombeck, D. A. & Reiser, M. B. Real neuroscience in virtual worlds. Curr. Opin. Neurobiol. 22, 3–10 (2012).
Larsch, J. & Baier, H. Biological motion as an innate perceptual mechanism driving social affiliation. Curr. Biol. 28, 3523–3532.e4 (2018).
Stowers, J. R. et al. Virtual reality for freely moving animals. Nat. Methods 14, 995–1002 (2017).
Keller, G. B., Bonhoeffer, T. & Hübener, M. Sensorimotor mismatch signals in primary visual cortex of the behaving mouse. Neuron 74, 809–815 (2012).
Dombeck, D. A., Khabbaz, A. N., Collman, F., Adelman, T. L. & Tank, D. W. Imaging large-scale neural activity with cellular resolution in awake, mobile mice. Neuron 56, 43–57 (2007).
Maimon, G., Straw, A. D. & Dickinson, M. H. Active flight increases the gain of visual motion processing in Drosophila. Nat. Neurosci. 13, 393–399 (2010).
Seelig, J. D. et al. Two-photon calcium imaging from head-fixed Drosophila during optomotor walking behavior. Nat. Methods 7, 535–540 (2010).
Ahrens, M. B. et al. Brain-wide neuronal dynamics during motor adaptation in zebrafish. Nature 485, 471–477 (2012).
Picardo, M. A. et al. Population-level representation of a temporal sequence underlying song production in the Zebra Finch. Neuron 90, 866–876 (2016).
Kerr, J. N. D. & Denk, W. Imaging in vivo: watching the brain in action. Nat. Rev. Neurosci. 9, 195–205 (2008).
Ahrens, M. B., Orger, M. B., Robson, D. N., Li, J. M. & Keller, P. J. Whole-brain functional imaging at cellular resolution using light-sheet microscopy. Nat. Methods 10, 413–420 (2013).
Panier, T. et al. Fast functional imaging of multiple brain regions in intact zebrafish larvae using selective plane illumination microscopy. Front. Neural Circuits 7, 65 (2013).
Schulze, L. et al. Transparent Danionella translucida as a genetically tractable vertebrate brain model. Nat. Methods 15, 977–983 (2018).
Portugues, R., Severi, K. E., Wyart, C. & Ahrens, M. B. Optogenetics in a transparent animal: circuit function in the larval zebrafish. Curr. Opin. Neurobiol. 23, 119–126 (2013).
Friedrich, R. W., Jacobson, G. A. & Zhu, P. Circuit neuroscience in Zebrafish. Curr. Biol. 20, R371–R381 (2010).
Muto, A., Ohkura, M., Abe, G., Nakai, J. & Kawakami, K. Real-time visualization of neuronal activity during perception. Curr. Biol. 23, 307–311 (2013).
Kim, D. H. et al. Pan-neuronal calcium imaging with cellular resolution in freely swimming zebrafish. Nat. Methods 14, 1107–1114 (2017).
Buske, C. & Gerlai, R. Shoaling develops with age in Zebrafish (Danio rerio). Prog. Neuropsychopharmacol. Biol. Psychiatry 35, 1409–1415 (2011).
Frank, T., Mönig, N. R., Satou, C., Higashijima, S. & Friedrich, R. W. Associative conditioning remaps odor representations and modifies inhibition in a higher olfactory brain area. Nat. Neurosci. 22, 1844–1856 (2019).
Gerlach, G. & Lysiak, N. Kin recognition and inbreeding avoidance in zebrafish, Danio rerio, is based on phenotype matching. Anim. Behav. 71, 1371–1377 (2006).
Valente, A., Huang, K.-H., Portugues, R. & Engert, F. Ontogeny of classical and operant learning behaviors in zebrafish. Learn. Mem. 19, 170–177 (2012).
Wanner, A. A., Genoud, C., Masudi, T., Siksou, L. & Friedrich, R. W. Dense EM-based reconstruction of the interglomerular projectome in the zebrafish olfactory bulb. Nat. Neurosci. 19, 816–825 (2016).
Friedrich, R. W., Genoud, C. & Wanner, A. A. Analyzing the structure and function of neuronal circuits in zebrafish. Front. Neural Circuits 7, 71 (2013).
Zhu, P., Fajardo, O., Shum, J., Zhang Schärer, Y.-P. & Friedrich, R. W. High-resolution optical control of spatiotemporal neuronal activity patterns in zebrafish using a digital micromirror device. Nat. Protoc. 7, 1410–1425 (2012).
Rao, R. P. N. & Ballard, D. H. Predictive coding in the visual cortex: a functional interpretation of some extra-classical receptive-field effects. Nat. Neurosci. 2, 79–87 (1999).
Bastos, A. M. et al. Canonical microcircuits for predictive coding. Neuron 76, 695–711 (2012).
Keller, G. B. & Mrsic-Flogel, T. D. Predictive processing: a canonical cortical computation. Neuron 100, 424–435 (2018).
Dreosti, E., Lopes, G., Kampff, A. R. & Wilson, S. W. Development of social behavior in young zebrafish. Front. Neural Circuits 9, 39 (2015).
Stednitz, S. J. et al. Forebrain control of behaviorally driven social orienting in Zebrafish. Curr. Biol. 28, 2445–2451.e3 (2018).
Gerlai, R. Animated images in the analysis of zebrafish behavior. Curr. Zool. 63, 35–44 (2017).
Rupprecht, P., Prendergast, A., Wyart, C. & Friedrich, R. W. Remote z-scanning with a macroscopic voice coil motor for fast 3D multiphoton laser scanning microscopy. Biomed. Opt. Express 7, 1656 (2016).
Koide, T. et al. Olfactory neural circuitry for attraction to amino acids revealed by transposon-mediated gene trap approach in zebrafish. Proc. Natl Acad. Sci. USA 106, 9884–9889 (2009).
Lal, P. et al. Identification of a neuronal population in the telencephalon essential for fear conditioning in zebrafish. BMC Biol. 16, 45 (2018).
Mueller, T., Dong, Z., Berberoglu, M. A. & Guo, S. The dorsal pallium in zebrafish, Danio rerio (Cyprinidae, Teleostei). Brain Res. 1381, 95–105 (2011).
Aoki, T. et al. Imaging of neural ensemble for the retrieval of a learned behavioral program. Neuron 78, 881–894 (2013).
Rodrı́guez, F. et al. Conservation of spatial memory function in the pallial forebrain of reptiles and ray-finned fishes. J. Neurosci. 22, 2894–2903 (2002).
Berens, P. et al. Community-based benchmarking improves spike rate inference from two-photon calcium imaging data. PLoS Comput. Biol. 14, e1006157 (2018).
Koster-Hale, J. & Saxe, R. Theory of mind: a neural prediction problem. Neuron 79, 836–848 (2013).
Blakemore, S.-J., Wolpert, D. M. & Frith, C. D. Central cancellation of self-produced tickle sensation. Nat. Neurosci. 1, 635–640 (1998).
Cullen, K. E. Vestibular processing during natural self-motion: implications for perception and action. Nat. Rev. Neurosci. 20, 346–363 (2019).
Schultz, W., Dayan, P. & Montague, P. R. A neural substrate of prediction and reward. Science 275, 1593–1599 (1997).
Attinger, A., Wang, B. & Keller, G. B. Visuomotor coupling shapes the functional development of mouse visual cortex. Cell 169, 1291–1302.e14 (2017).
McElligott, M. B. & O’Malley, D. M. Prey tracking by larval Zebrafish: axial kinematics and visual control. Brain. Behav. Evol. 66, 177–196 (2005).
Wang, T. et al. Three-photon imaging of mouse brain structure and function through the intact skull. Nat. Methods 15, 789–792 (2018).
Ji, N. Adaptive optical fluorescence microscopy. Nat. Methods 14, 374–380 (2017).
Asakawa, K. et al. Genetic dissection of neural circuits by Tol2 transposon-mediated Gal4 gene and enhancer trapping in zebrafish. Proc. Natl Acad. Sci. USA 105, 1255–1260 (2008).
Han, X. et al. A high-light sensitivity optical neural silencer: development and application to optogenetic control of non-human primate cortex. Front. Syst. Neurosci. 5, 18 (2011).
Kwan, K. M. et al. The Tol2kit: a multisite gateway-based construction kit forTol2 transposon transgenesis constructs. Dev. Dyn. 236, 3088–3099 (2007).
Distel, M., Wullimann, M. F. & Koster, R. W. Optimized Gal4 genetics for permanent gene expression mapping in zebrafish. Proc. Natl Acad. Sci. USA 106, 13365–13370 (2009).
Leinweber, M. et al. Two-photon calcium imaging in mice navigating a virtual reality environment. J. Vis. Exp. 84, e50885 (2014).
Huang, K.-H., Ahrens, M. B., Dunn, T. W. & Engert, F. Spinal projection neurons control turning behaviors in Zebrafish. Curr. Biol. 23, 1566–1573 (2013).
Pologruto, T. A., Sabatini, B. L. & Svoboda, K. ScanImage: flexible software for operating laser scanning microscopes. Biomed. Eng. OnLine 2, 13 (2003).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Frey, B. J. & Dueck, D. Clustering by passing messages between data points. Science 315, 972–976 (2007).
Acknowledgements
We thank G. Keller and M. Leinweber for comments on the manuscript; P. Argast, P. Buchmann and P. Zmarz (Friedrich Miescher Institute for Biomedical Research, Basel, Switzerland) for technical support; A. Prendergast and C. Wyart (Institut du Cerveau et de la Moelle Épinière, Paris, France) for sharing Tg(neuroD:GCaMP6f) fish; E. Boyden (Massachusetts Institute of Technology, Boston, MA, USA), B. Roska (Friedrich Miescher Institute for Biomedical Research, Basel, Switzerland) for sharing reagents (ArchT-GFP), and the Friedrich group for stimulating discussions. This work was supported by the Novartis Research Foundation (R.W.F.), by a Novartis Institutes for Biomedical Research Presidential Postdoctoral Fellowship (K.H.H.), by the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation program (grant agreement no. 742576), by the Swiss National Science Foundation (R.W.F. grant nos. 310030B_152833/1 and 135196/1), by a fellowship from the Boehringer Ingelheim Fonds (P.R.) and by the NBRP and NBRP/Fundamental Technologies Upgrading Program from AMED (K.K.).
Author information
Authors and Affiliations
Contributions
K.H.H. developed the methodology, designed and performed experiments, analyzed data and wrote the manuscript. P.R. developed methodology and wrote the manuscript. T.F. and K.K. created and analyzed transgenic fish. T.B. supervised the project. R.W.F. supervised the project, analyzed data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nina Vogt was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Head fixation of adult zebrafish.
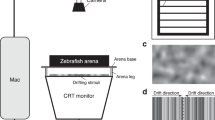
a, Location to glue L-shaped head bars. Skin removal should be avoided at two specific locations to prevent bleeding (red arrows). b, Photographic images showing five major sets of skull bones in adult zebrafish. Sets 1 - 4 are located on top of set 5. Tel: telencephalon, TeO: optic tectum, CB: cerebellum. c, Wrapping materials to keep skin moisturized and to stabilize the body during surgery. d, Setup to glue a head bar to the skull. e, Components of the VR chamber. Inset: grooves on the base plate (top) and post plate (bottom) facilitate stabilization of the posts in the VR chamber. f, Tools to align the yaw, roll and pitch angles of the fish on the mounting posts.
Supplementary Figure 2 Workflow of the closed-loop VR.
Photograph shows a fish in a custom-made holder for reversible mounting. In the LabVIEW interface, the animal’s behavior is recorded by the camera and tail curvature is analyzed in real-time. The gain for angular and forward update in the VR can be specified independently. The duration and the type of movies (visual stimuli) can be assigned automatically or manually. These settings are then sent to a Python-based environment (Panda 3D) to control the position and angle of virtual cameras in the VR as well as the display of dynamic texture for movie presentations. Collision between the virtual wall and virtual cameras is also calculated. The scene captured by the virtual cameras is presented to the animal using LED projectors, thus closing the loop from motor output to visual feedback. The “Hijack mode” in the LabVIEW interface uncouples the animal’s behavior from VR update. This feature can be used to introduce an open-loop condition by presenting a static VR or trigger optomotor response by rotating or linearly drifting the VR.
Supplementary Figure 3 Gain settings for VR update.
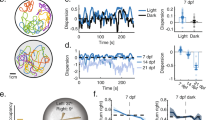
When a fish was introduced into the VR, translational and rotational gains were initially set to default values and subsequently fine-adjusted to minimize struggling and inactivity in each individual. a, Distribution of the mean angular gain in the VR (n = 26 fish). Box plot shows median (2.95), 25th and 75th percentiles (2.05 - 4.0); dot and whiskers show mean ± s.d. (3.4 ± 1.6). b, Variation of the angular gain across experiments in each fish, measured by the coefficient of variation (CV; s.d. normalized by the mean). Each plot symbol shows the CV of the angular gain in an individual fish (n = 26 fish). Box plot shows median, 25th and 75th percentiles; dot and whiskers show mean ± s.d. (median: 9.3%, 25th and 75th percentiles: 5.3% - 19.8%; mean ± s.d.: 13.7 ± 11.7%). c, Distribution of the mean forward gain in the VR (n = 26 fish). Box plot shows median (0.38), 25th and 75th percentiles (0.28 - 0.4); dot and whiskers show mean ± s.d. (0.37 ± 0.11). d, Variation of the forward gain across experiments in each fish, quantified by the CV (n = 26 fish). Box plot shows median, 25th and 75th percentiles; dot and whiskers show mean ± s.d.. CV: median: 2.1%, 25th and 75th percentiles: 0% - 4.5%; mean ± s.d.: 3.6 ± 5.0%.
Supplementary Figure 4 Swim patterns of adult zebrafish.
a, Video frames during a swim event showing different tail curvature. Red shading depicts the active swimming period detected by thresholding tail movement intensity, which was measured by the mean of the absolute pixel-wise difference in adjacent frames. Similar behavior was observed in head-fixed zebrafish (n = 26) and in freely swimming zebrafish (n = 8). b, Tail curvature as a function of time in a freely swimming fish. Red shading defined as in a. Experiments were repeated in 8 fish. c, Tail curvature as a function of time in a head-fixed fish in the closed-loop VR. Experiments were repeated in 26 fish. d, Fraction of time that freely swimming fish engaged in active swimming behavior in successive 30-minute time intervals. Active swimming was defined as in a. Box plots show median, 25th and 75th percentiles. Dots and error bars show mean ± s.d. (n = 8 fish). e, Fraction of time that head-fixed fish in a closed-loop VR engaged in active swimming. Number of fish in successive 30-min time bins: 26, 26, 24, 19, 12, 7, 1, 1 fish. In all fish, calcium imaging was performed during each time window. Box plots show median, 25th and 75th percentiles. Dots and error bars show mean ± s.d..
Supplementary Figure 5 Comparison of swimming behavior under head-fixation in a closed loop VR to free swimming.
a, Distribution of swim period, active period and inactive period of individual swim events performed by head-fixed adult zebrafish in a closed-loop VR (n = 26 fish, each recorded for >30 min). As illustrated in the inset, the swim period is the time between the onsets of successive swims and consists of an active period (active tail movement) followed by an inactive period (no tail movement). The medians of the swim period, active period and inactive period were 0.98 s (25th–75th percentile = 0.64 s–1.6 s), 0.46 s (25th–75th percentile = 0.24 s–0.86 s) and 0.36 s (25th–75th percentile = 0.22 s–0.62 s), respectively. To identify activity, the intensity of tail movements was quantified by the mean of the absolute pixel-wise difference in adjacent video frames and thresholded. b, Frequency distribution of swim events (median = 1.02 Hz, 25th–75th percentile = 0.63 HZ–1.56 Hz). In this and following plots, box plot shows median, 25th percentile and 75th percentile; dot and whiskers show mean ± s.d. (n = 26 fish). c, Distribution of the change in heading direction during individual swim events (median = 0.16°, 25th–75th percentile = -6.89° - 8.77°; n = 26 fish). d, Distribution of distances traveled during individual swim events. Note that absolute distance is not defined objectively in the VR because distance is represented by VR units that can be scaled arbitrarily relative to physical distance. We chose a scaling such that 1 VR unit corresponded to approximately 5 mm in physical distance because this resulted in a size of approximately 25 mm for animated fish when movies were displayed. However, it is obviously possible that this scaling does not represent the perceived distance in the VR. The median distance per swim event was 3.0 mm (25th–75th percentile = 0.6–7.6 mm; n = 26 fish). For technical reasons the detection of individual swim events by a fixed threshold of tail movement intensity resulted in the detection of short events that did not reflect real swim bouts (“spurious swim events”). To exclude such spurious events from the analysis of mean swim distance (reported below), we fitted the histogram by the sum of two distributions (red): pink noise (dashed grey line) to represent short spurious swim events and a Gaussian distribution (solid grey) to represent real swim events (mean ± s.d. = 5.2 ± 4.4 mm). e–h, Behavioral analysis of freely swimming adult zebrafish. The median swim period, active period and inactive period were 0.60 s (25th – 75th percentile: 0.45 s – 0.87 s), 0.18 s (25th – 75th percentile: 0.12 s – 0.36 s) and 0.33 s (25th – 75th percentile: 0.21 s – 0.48 s), respectively. The median swim frequency was 1.67 Hz (25th – 75th percentile: 1.15Hz – 2.22 Hz). The median change in heading angle was 0.41° (25th – 75th percentile: -16.8° - 17.6°) and the median distance traveled per swim was 26.4 mm (25th – 75th percentile: 17.8 - 38.5 mm; mean ± s.d. = 31.0 ± 20.1 mm). Data from n = 18 fish. The duration of the active period was significantly higher under head fixation (p < 0.001; two-sided Wilcoxon rank-sum test) while the swim frequency, the absolute angular change and the distance traveled per swim event were significantly lower (p < 0.001; two-sided Wilcoxon rank-sum test). As a consequence, the total distance travel per hour was lower under head-fixation (24.4 ± 1.8 m; spurious events excluded) than in freely swimming fish (207.2 ± 8.4 m). However, these distances cannot be compared directly because absolute distance is not defined objectively in the VR. Moreover, it is also difficult to directly compare changes in heading direction (“turning”) because this behavior is likely to strongly depend on the geometry of the environment.
Supplementary Figure 6 Behavioral response of freely swimming adult zebrafish to movies of conspecifics.
a, Schematic of the behavioral assay. Individual animals can swim freely in a rectangular chamber (20 cm by 10 cm by 15 cm, length by width by height; water depth = 10 cm) and observe movies of conspecifics through anti-reflection glass at two ends of the tank. b, Probability map of animal position during 20 s periods of no stimulus presentation (left) or fish movie presentation on one side (right; n = 64 fish). c, Stimulus-induced change in side preference index (SPI). The SPI is defined as the mean of the normalized position along the long axis (between -1 and 1) during stimulus presentation. The change in SPI is defined as ΔSPI = SPI(movie vs. empty tank) - SPI (empty tank vs. empty tank). Fish exhibited a significant change in side preference towards the movie of conspecifics (p = 10-10, two-sided t-test, n = 64 fish). Error bars represent mean ± s.e.m..
Supplementary Figure 7 Effect of the skull on optical resolution.
To explore conditions for optical imaging in the telencephalon of adult zebrafish we extracted telencephalic skulls of adult zebrafish and acquired z-stacks of fluorescent beads positioned below the skulls using a standard two-photon microscope. Column 1: Point spread function (PSF) of two-photon imaging determined by imaging fluorescent beads embedded in agarose at different depth. The width of the PSF in axial direction (bottom) and within the horizontal plane (top) was quantified by the full width at half maximum (FWHM). Columns 2 – 4: Axial and radial PSF as a function of depth below the bone after placing the telencephalic skull of adult zebrafish on top of agarose-embedded fluorescent beads. Three skulls (length by width = 2 mm by 1.5 mm) were collected from different animals. Red lines represent the linear regression of each distribution. Note that the PSF in scattering brain tissue would likely be broader. Based on these results we estimate that broadening of the PSF by the skull does not preclude high-resolution imaging of neuronal activity up to a depth of at least 200 μm (not including the thickness of the skull itself), at least in a subset of fish.
Supplementary Figure 8 Two-photon imaging through the intact skull.
In vivo two-photon imaging was performed through the skull in the telencephalon of Tg(Gad1b:loxP-DsREd-loxP-EGFP) fish which expresses GFP in GABAergic interneurons (n = 1 fish)61. Individual somata and neurites could be resolved up to a depth of >200 μm below the brain surface (not including the thickness of the skull). Images are averages over 50 frames acquired at 30 Hz without applying any spatial filtering or other image processing. Depth was measured from the surface of the brain (thickness of the skull not included). Laser power at 920 nm linearly increased from 18 mW at the brain surface to 45 mW at a depth of 275 μm (Nikon 16x objective, NA = 0.8). 61 Satou, C. et al. Transgenic tools to characterize neuronal properties of discrete populations of zebrafish neurons. Development 140, 3927–3931 (2013).
Supplementary Figure 9 Non-invasive imaging of GCaMP6f in the dorsal telencephalon of adult zebrafish.
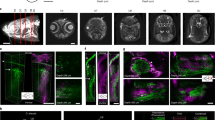
a, Schematic showing the location of the dorsal telencephalon (left) and its subdivisions (right). Tel: telencephalon; TeO: optic tectum; OB: olfactory bulb. b–m. In vivo two-photon imaging of fluorescence in Tg(neuroD:GCaMP6f) fish, which express the calcium indicator GCaMP6f in most pallial forebrain areas (n = 1 fish)36. The green channel shows expression of GCaMP6f in a dorsal-to-ventral series of two-photon images acquired through the intact skull of a head-fixed adult zebrafish. Each image is the mean of an image series of 8000 frames. Insets show enlarged subregions (40 μm x 40 μm). The magenta channel shows a maximum projection after subtraction of the mean, highlighting pixels where large changes in fluorescence occurred during the acquisition of the image series (indicating neuronal activity). Note that expression of GCaMP6f (green) is broad but spontaneous activity (magenta) was relatively sparse. Note also that individual somata and the activity of single neurons could be resolved throughout the dorsal telencephalon. Labels denote telencephalic brain areas. Dm is separated from Dc regions by the sulcus of ypsilonformis (sy). cDc was separated from rDc and Dl by an obvious anatomical boundary (dashed line in e). Dl is lateral to rDc at the rostral end of the forebrain but extends dorsally and covers rDc at caudal locations (d - f). z = 0 μm is defined by the dorsal surface of Dm. Depth measures do therefore not include the thickness of the skull.
Supplementary Figure 10 Expression pattern and integration site of SAGFF212C transgenics.
a, Expression of GFP in Tg(SAGFF212C:Gal4), Tg(UAS:GFP) zebrafish at five days post fertilization (dpf). Fish were created by a gene trap51 approach and express the Gal4 transactivator in a tissue-specific pattern37. b, Expression of GFP in the brain of an adult Tg(SAGFF212C:Gal4), Tg(UAS:GFP) fish. Prominent expression was detected in the telencephalon, the hypothalamus and the cerebellum. c, Coronal section through the adult telencephalon. d, Integration site of the transposon in Tg(SAGFF212C:Gal4) fish at the ppfia3 gene on chromosome 3.
Supplementary Figure 11 Intensity and duration of tail movement during different motor behaviors.
a, Mean tail movement intensity during regular swim bouts in the normal VR, during BREPs and during struggling. Tail movement intensity was averaged over the first 3 s of each event. Data from 102979 periodic swim events, 303 BREPs and 40 struggling events. Error bars show s.e.m.. p = 10-168 (normal VR vs. BREP), 10-66 (normal VR vs. struggling), 10-7 (BREP vs. struggling); two-sided t-test. Asterisks indicate p < 0.001 (***). b, Duration of tail movements during different motor behaviors. The duration of active swimming was significantly longer during BREPs (8.4 ± 0.2 s, mean ± s.e.m.) and struggling events (8.0 ± 3.7 s) than during regular swim events in the normal VR (0.76 ± 0.003 s; two-sided t-test, p < 0.001, respectively). The duration was determined by the time that tail movement intensity exceeded a fixed detection threshold. Data represents 102979 periodic swim events, 303 BREPs and 40 struggling events. Asterisks indicate p < 0.001 (***).
Supplementary Figure 12 Behavioral responses to visuo-motor perturbation.
a, Duration of active swimming during normal and perturbed visuo-motor coupling. Cumulative distribution of time engaged in active swimming of increasing duration. Active swimming time was elevated during visuo-motor perturbations (red) in comparison to swims during normal visuo-motor coupling (black). The fraction of prolonged active swimming was also increased during perturbations (dashed line, right-hand y-axis). Shading shows s.e.m. over animals (n = 26). b, Behavioral adaptation to visuo-motor perturbations. The strength of the behavioral response to a perturbation was quantified as (m2-m1) / m1, where m1 and m2 are the mean tail movement intensity during 10 s time windows immediately before and after perturbation onset, respectively. Linear regression was used to model the behavioral response as a function of the number of perturbations experienced. The histogram shows the distribution of regression slopes across fish (n = 26). The mean of the distribution was not significantly different from zero (p = 0.41, two-sided t-test). Hence, the analysis revealed no obvious sign of behavioral adaptation, possibly because the interval between perturbations was long (5.0 ± 4.8 min; mean ± s.d.). c, Distribution of pre-BREP periods. The median of the pre-BREP period was 2.1 s (25th – 75th percentile = 1.2 s – 4.1 s; 303 perturbation trials from 26 fish). The mean pre-BREP periods of individual fish are shown by red dots. Box plots show median, 25th and 75th percentiles. Dots and error bars show mean ± s.d.
Supplementary Figure 13 Neuronal responses to visual feedback mismatch.
a, Clustering of responses (882 out of 7090 neuron-perturbation pairs from 26 fish). Calcium fluorescence signals (ΔF/F) were aligned to the onset of the BREP and clustered by affinity propagation. Right: average ΔF/F of each cluster. b, Pre-BREP responses (left) and BREP responses (right) aligned to the onset of perturbations (dashed line) and sorted by the onset of the BREP (solid curve). c, Consistency of pre-BREP and BREP responses. The analysis included n = 558 neurons that were recorded in at least two positive trials (perturbation trials that resulted in a BREP). Each symbol represents a neuron. Most neurons responded predominantly during the pre-BREP period or during the BREP period (data points near the x-axis or y-axis). The inset shows the distribution of angular coordinates of each data point (illustrated for one data point in red; only neurons with at least two BREP responses were included). An angle of 0 ° (90 °) indicates that a neuron responded exclusively during the BREP (pre-BREP) period while an angle of 45 ° indicates equal response probabilities during both periods. The distribution indicates that most neurons responded selectively during the BREP or during the pre-BREP period. However, the number of observations per neuron was insufficient to conclude that individual neurons responded preferentially during the pre-BREP or the BREP phase although the results are consistent with this hypothesis. d, Examples of responses of a pre-BREP neuron in three trials. Response onset consistently occurred before the onset of the BREP (red dot). e, Mean AP probability of pre-BREP responses (green; n = 382 neuron-perturbation pairs) and BREP responses (red; n = 500 neuron-perturbation pairs), triggered on the onset of spontaneous swims in a closed-loop VR without perturbation. Shading indicates s.e.m. f, Accuracy of decoding behavior (occurrence of BREP) based on the activity of pre-BREP neurons. Subsets of neurons were selected randomly from the set of pre-BREP neurons (n = 65). For each neuron in a subset, individual responses representing a positive trial and a negative trial were selected randomly. Selected responses were then averaged over each subset and considered correct when the mean response was higher for the subset representing positive trials. The procedure was repeated 1000 times for each subset size.
Supplementary Figure 14 Estimate of false-positive rate for response detection.
For each neuron-perturbation pair (n = 7090), the detection of neuronal responses was triggered on a random time point instead of perturbation onset in the same image series. Plots show the probability of observing a significant change in ΔF/F (“response”) in different brain regions during fictive pre-BREP and BREP periods following the randomly selected time points. The onsets of fictive BREPs were determined by randomly sampling the onsets of BREPs triggered by perturbations. In each of the forebrain regions, the probability of detecting pre-BREP response and BREP response were significantly lower in comparison to the probability of detecting a response when the analysis was triggered on perturbations (Fig. 6f). Dm (pre-BREP response, p = 10-6; BREP response, p ≤ 10-16; X2 test, n = 2520); cDc (pre-BREP response, p = 10-4; BREP response, p = 10-16; X2 test, n = 805); rDc (pre-BREP response, p = 10-11; BREP response, p = 10-14; X2 test, n = 882); Dl (pre-BREP response, p = 10-12; BREP response, p ≤ 10-16; X2 test, n = 2883). Asterisk indicates p < 0.001 in the brain area.
Supplementary information
Supplementary Information
Supplementary Figs. 1–14 and Protocol
Supplementary Video 1
Responses of head-fixed adult zebrafish to visual access of conspecifics in the VR. Video shows tail movements of a head-fixed adult zebrafish and the concomitant projection of the virtual environment in closed-loop. The figure shows an overview of the VR and the time course of the experiment. Red shading depicts the period corresponding to the video.
Supplementary Video 2
Brain motion of head-fixed adult zebrafish during behaviour. Time series of two-photon images of a neuron (frame rate, 30 Hz) taken through the intact skull from a head-fixed adult zebrafish in vivo (top left) and concomitant tail movements (bottom left). Top right image shows an average of the image frames in the series. Bottom right. quantification of image motion and tail movement as a function of time.
Supplementary Video 3
Responses of head-fixed adult zebrafish to left-right reversals in the VR. Top, view of the virtual environment. Bottom left, concomitant neural activity in a field of view in the telencephalon. Bottom right, concomitant video imaging of the tail and quantification of tail movement and neural activity as a function of time.
Supplementary Video 4
Behavioral responses (BREPs) of head-fixed adult zebrafish to left-right reversals in the VR. Video imaging of tails in 24 fish during visuomotor perturbation. Open squares mark perturbation onset, filled squares mark manually annotated onsets of behavioural responses (BREPs).
Supplementary Video 5
Postsurgery behaviors: saccades, locomotion, feeding and social affiliation. Examples of different behaviors after attachment of head bars.
Rights and permissions
About this article
Cite this article
Huang, KH., Rupprecht, P., Frank, T. et al. A virtual reality system to analyze neural activity and behavior in adult zebrafish. Nat Methods 17, 343–351 (2020). https://doi.org/10.1038/s41592-020-0759-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0759-2
This article is cited by
-
An automated, low-latency environment for studying the neural basis of behavior in freely moving rats
BMC Biology (2023)
-
An optofluidic platform for interrogating chemosensory behavior and brainwide neural representation in larval zebrafish
Nature Communications (2023)
-
Extended reality for biomedicine
Nature Reviews Methods Primers (2023)
-
Precise visuomotor transformations underlying collective behavior in larval zebrafish
Nature Communications (2021)
-
Zebrafish capable of generating future state prediction error show improved active avoidance behavior in virtual reality
Nature Communications (2021)