Abstract
Achieving a comprehensive understanding of brain function requires multiple imaging modalities with complementary strengths. We present an approach for concurrent widefield optical and functional magnetic resonance imaging. By merging these modalities, we can simultaneously acquire whole-brain blood-oxygen-level-dependent (BOLD) and whole-cortex calcium-sensitive fluorescent measures of brain activity. In a transgenic murine model, we show that calcium predicts the BOLD signal, using a model that optimizes a gamma-variant transfer function. We find consistent predictions across the cortex, which are best at low frequency (0.009–0.08 Hz). Furthermore, we show that the relationship between modality connectivity strengths varies by region. Our approach links cell-type-specific optical measurements of activity to the most widely used method for assessing human brain function.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The raw MR and optical-imaging data generated during the current study are available from the corresponding author upon reasonable request. Data are not available in a public repository at the time of this publication due to ongoing work by the authors on these data set. The Allen Brain Atlas was downloaded from http://www.brain-map.org.
Code availability
Custom MATLAB code for fMRI preprocessing and MR and Ca2+ data post-processing (parcellation and computation of connectivity matrices) is available from the corresponding author upon reasonable request. For analyses of Ca2+ data, refer to: https://github.com/bioimagesuiteweb/bisweb/tree/calcium. The analysis tools for multi-modal data registration and analysis are available BioImageSuite Web at www.bioimagesuite.org.
Change history
23 June 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41592-021-01217-0
References
Ross, W. N. Understanding calcium waves and sparks in central neurons. Nat. Rev. Neurosci. 13, 157–168 (2012).
Barson, D. et al. Simultaneous mesoscopic and two-photon imaging of neuronal activity in cortical circuits. Nat. Methods 17, 107–113 (2020).
Ogawa, S. et al. Brain magnetic resonance imaging with contrast dependent on blood oxygenation. Proc. Natl Acad. Sci. USA 87, 9868–9872 (1990).
Albers, F. et al. Multimodal functional neuroimaging by simultaneous BOLD fMRI and fiber-optic calcium recordings and optogenetic control. Mol. Imaging Biol. 20, 171–182 (2018).
Hyder, F. et al. Neurovascular and neurometabolic couplings in dynamic calibrated fMRI: transient oxidative neuroenergetics for block-design and event-related paradigms. Front. Neuroenerg. 2, 1–11 (2019).
Ma, Y. et al. Resting-state hemodynamics are spatiotemporally coupled to synchronized and symmetric neural activity in excitatory neurons. Proc. Natl Acad. Sci. USA 113, E8463–E8471 (2016).
Keilholz, S. D. The neural basis of time-varying resting-state functional connectivity. Brain Connect 4, 769–779 (2014).
Mateo, C. et al. Entrainment of arteriole vasomotor fluctuations by neural activity is a basis of blood-oxygenation-level-dependent ‘resting-state’ connectivity. Neuron 96, 936–948 (2017).
Desjardins, M. et al. Awake mouse imaging: from two-photon microscopy to blood oxygen level-dependent functional magnetic resonance imaging. Biol. Psychiatry Cogn. Neurosci. Neuroimaging 4, 533–542 (2019).
Kennerley, A. J. et al. Is optical imaging spectroscopy a viable measurement technique for the investigation of the negative BOLD phenomenon? A concurrent optical imaging spectroscopy and fMRI study at high field (7 T). Neuroimage 61, 10–20 (2012).
Kennerley, A. J. et al. Concurrent fMRI and optical measures for the investigation of the hemodynamic response function. Magn. Reson. Med. 54, 354–365 (2005).
Schulz, K. et al. Simultaneous BOLD fMRI and fiber-optic calcium recording in rat neocortex. Nat. Methods 9, 597–602 (2012).
Liang, Z. et al. Simultaneous GCaMP6-based fiber photometry and fMRI in rats. J. Neurosci. Methods 289, 31–38 (2017).
Schwalm, M. et al. Cortex-wide BOLD fMRI activity reflects locally-recorded slow oscillation-associated calcium waves. eLife 6, e27602 (2017).
Wang, M. et al. Brain-state dependent astrocytic Ca2+ signals are coupled to both positive and negative BOLD-fMRI signals. Proc. Natl Acad. Sci. USA 115, E1647–E1656 (2018).
Schlegel, F. et al. Fibre-optic implant for simultaneous fluorescence-based calcium recordings and BOLD fMRI in mice. Nat. Protoc. 13, 840–855 (2018).
Lien, E. S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168–176 (2007).
Schroeter, A. et al. Specificity of stimulus-evoked fMRI responses in the mouse: the influence of systemic physiological changes associated with innocuous stimulation under four different anesthetics. Neuroimage 94, 372–384 (2014).
Zhang, S. & Murphy, T. H. Imaging the impact of cortical microcirculation on synaptic structure and sensory-evoked hemodynamic responses in vivo. PLoS Biol. 5, e119 (2007).
Cramer, J. V. et al. In vivo widefield calcium imaging of the mouse cortex for analysis of network connectivity in health and brain disease. Neuroimage 199, 750–584 (2019).
Turner, R. How much cortex can a vein drain? Downstream dilution of activation-related cerebral blood oxygenation changes. Neuroimage 16, 1062–1067 (2002).
Ma, Y. et al. Wide-field optical mapping of neural activity and brain haemodynamics: considerations and novel approaches. Philos. Trans. R. Soc. Lond. B Biol. Sci. 371, 1–17 (2016).
Bauer, A. Q. et al. Optical imaging of disrupted functional connectivity following ischemic stroke in mice. Neuroimage 99, 388–401 (2014).
White, B. R. et al. Imaging of functional connectivity in the mouse brain. PLoS ONE 6, e16322 (2011).
Fox, M. D. et al. The human brain is intrinsically organized into dynamic, anticorrelated functional networks. Proc. Natl Acad. Sci. USA 102, 9673–9678 (2005).
Shen, X., Papademetris, X. & Constable, R. T. Graph-theory based parcellation of functional subunits in the brain from resting-state fMRI. Neuroimage 50, 1027–1035 (2010).
van Oort, E. S. B. et al. Functional parcellation using time courses of instantaneous connectivity. Neuroimage 170, 31–40 (2018).
Arslan, S. et al. Human brain mapping: a systematic comparison of parcellation methods for the human cerebral cortex. Neuroimage 170, 5–30 (2018).
Eickhoff, S. B., Yeo, B. T. T. & Genon, S. Imaging-based parcellations of the human brain. Nat. Rev. Neurosci. 19, 672–686 (2018).
Shen, X. et al. Groupwise whole-brain parcellation from resting-state fMRI data for network node identificatio. Neuroimage 82, 403–415 (2013).
Uhlirova, H. et al. The roadmap for estimation of cell-type-specific neuronal activity from non-invasive measurements. Philos. Trans. R. Soc. Lond. B Biol. Sci. 371, 20150356 (2016).
Scaglione, A. et al. Trial-totrial variability in the responses of neurons carries information about stimulus location in the rat whisker thalamus. Proc. Acad. Sci. USA 108, 14956–14961 (2011).
Lee, J. H. Informing brain connectivity with optogenetic functional magnetic resonance imaging. Neuroimage 62, 2244–2249 (2012).
Miyamoto, D. & Murayama, M. The fibre-optic imaging and manipulation of neural activity during animal behavior. Neurosci. Res. 103, 1–9 (2016).
Palmer, H. S. Optogenetic fMRI sheds light on the neural basis of the BOLD signal. J. Neurophysiol. 104, 1838–1840 (2010).
Logothetis, N. K. The neural basis of the blood-oxygen-level-dependent functional magnetic resonance imaging signal. Philos. Trans. R. Soc. Lond. B Biol. Sci. 357, 1003–1037 (2002).
Buxton, R. B. The physis of functional magnetic resonance imaging (fMRI). Rep. Prog. Phys. 79, 096601 (2013).
Gao, Y. R. et al. Time to wake up: studying neurovascular coupling and brain-wide circuit function in the un-anesthetized animal. Neuroimage 153, 382–398 (2017).
Daigle, T. L. et al. A suite of transgenic driver and reporter mouse lines with enhanced brain cell type targeting and functionality. Cell 174, 465–480 (2018).
Madisen, L. et al. Transgenic mice for intersectional targeting of neural sensors and effectors with high specificity and performance. Neuron 85, 942–958 (2015).
Zeng, H. et al. An inducible and reversible mouse genetic rescue system. PLoS Genet. 4, e1000069 (2008).
Harris, J. A. et al. Anatomical characterization of Cre driver mice for neural circuit mapping and manipulation. Front. Neural Circuits 8, 76 (2014).
Allen, W. E. et al. Global representations of goal-directed behavior in distinct cell types of mouse neocortex. Neuron 94, 891–907 (2017).
Valley, M. T. et al. Separation of hemodynamic signals from GCaMP fluorescence measured with widefield imaging. J. Neurophysiol. 123, 256–366 (2020).
Shen, Q., Huang, S. & Duong, T. Q. Ultra-high\spatial resolution basal and evoked cerebral blood flow MRI of the rat brain. Brain Res. 1599, 126–136 (2014).
Wekselblatt, J. B. et al. Large-scale imaging of cortical dynamics during sensory perception and behavior. J. Neurophysiol. 115, 2852–2866 (2016).
Cox, R. W. AFNI: software for analysis and visulization of functional magnetic resonance neuroimages. Comput. Biomed. Res. 29, 162–173 (1996).
Tsurugizawa, T., Djemai, B. & Zalesky, A. The impact of fasting on resting state brain networks in mice. Sci. Rep. 9, 2976 (2019).
Pais-Roldán, P. et al. Identifying respiration-related aliasing artifacts in the rodent resting-state fMRI. Front. Neurosci. 12, 788 (2018).
Li, Q. et al. Resting-state functional MRI reveals altered brain connectivity and its correlation with motor dysfunction in a mouse model of Huntington’s disease. Sci. Rep. 7, 16742 (2017).
Lake, E. M. R. et al. Modulation of the peri-infarct neurogliovascular function by delayed COX-1 inhibition. J. Magn. Reson Imaging 46, 505–517 (2017).
Shen, X. et al. Groupwise whole-brain parcellation from resting-state fMRI data for network node identification. Neuroimage 82, 403–415 (2013).
Acknowledgements
We would like to thank all members of the Multiscale Imaging and Spontaneous Activity in Cortex (MISAC) collaboration at Yale University for their valuable contributions to this project. We thank P. Brown for valuable input on the design and building the RF saddle coil and the design and building of the telecentric lens holder. We thank Joel Greenwood and the Neurotechnology Core for modifying and building optics associated with the telecentric lens. We thank A. DeSimone, P. Brown and the Yale School of Medicine electronics and machine shop for help with rebuilding the telecentric lens. We thank C. Lacadie for help with data registration. This work was supported by funding from the NIH R01 MH111424 to R.T.C., M.C.C. and F.H., as well as U01 N2094358 to M.C.C. and R.T.C.
Author information
Authors and Affiliations
Contributions
All authors contributed to the overall study design. E.M.R.L. and X.G. designed and constructed the imaging apparatus and collected the data. E.M.R.L. and X.G. analyzed the data. E.M.R.L., X.G., X.S., D.S. and X.P. contributed code for the analysis of the data. P.H., EMRL, and XG designed the surgical protocol for longitudinal dual-imaging experiments. PH conducted these surgeries. E.M.R.L., X.G., F.H., J.A.C., M.J.H., M.C.C. and R.T.C. wrote the manuscript. M.C.C. and R.T.C. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nina Vogt was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Assembly of MR saddle coil, mouse head-plate, and Ca2+ imaging optical apparatus. Custom saddle coil and imaging apparatus design.
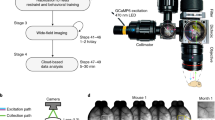
a, Removable saddle coil, with case that protects hardware. b, Coil in place on dual imaging sled. Coil (white) is mounted on a support system (blue) which is fixed to the sled (semi-transparent blue). A support system for the mouse (orange) is attached to the sled to which the mouse head plate (red) attaches. c, Assembled dual imaging apparatus. The telecentric lens (for Ca2+ imaging) is secured above the mouse and saddle coil. The position of the telecentric lens (and housing) can be adjusted (yellow) along the magnet Bo axis to focus the Ca2+ image.
Extended Data Fig. 2 A cross section of the optical apparatus.
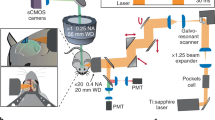
A diagram of the light path overlaid on a cross section of our optical apparatus. The light enters the system via a flexible liquid light guide. At the base of the telecentric lens, the light is bent by 90° degrees to enter the telecentric lens. Upon entering the telecentric lens, the excitation light reflects off of the dichroic mirror and is redirected along the length of the telecentric lens and into the prism at the end of the apparatus. The prism redirects the excitation light onto the mouse cortex. The emission light is similarly re-directed by the prism along the length of the telecentric lens (this time traveling in the opposite direction) and passes through the dichroic mirror. The fiber bundle array is mounted onto the end of the telecentric lens and transmits the light to the room neighboring the magnet where the camera is housed. To focus the Ca2+ image, the fiber bundle moves relative to the telecentric lens (red arrows).
Extended Data Fig. 3 Videos of Ca2+ data pre- and post-image processing.
Representative frames from example videos (Supplementary Video 1 & 2) a, Raw (unprocessed) fluorescence signal (cyan wavelength). A fluorescent bead placed within the dental cement at the right anterior edge of the surgical preparation is indicated (white arrow). The bead is used for right and left identification and motion correction. b, Data from a) after processing. c, Estimated motion parameters based on position of fluorescent bead.
Extended Data Fig. 4 Ray-casting algorithm to create the TOF MR angiogram-projected surface image for multi-modal image registration.
a, Three example views of the raw 3D MR angiogram data. Blood vessels have high MR signal intensity. b, Example of a maximum intensity projection (MIP) image (left) and a schematic of our ray-casting approach (right). The MIP is generated following masking which removes signal from anatomy outside of the brain. To show the curvature of the brain surface, and to isolate the blood vessels on the surface of the brain, we use the ray-casting algorithm. We project the MR data along the axis perpendicular to the optical imaging plane. Each pixel is shaded based on brain curvature. c, The resulting 2D projection of the MR image.
Extended Data Fig. 5 Average responses to unilateral hind-paw stimulation.
a, Average responses to nine stimuli across N = 6 mice. Stimulus onset is denoted by black triangles. Ca2+ data (top) and the corresponding fMRI data (bottom) are plotted. The fMRI signal is normalized to the mean. The standard deviation within the responding ROI is shown as shading. b, The average normalized (to peak amplitude), stimulus response across N = 6 mice, n = 9 stimuli each. This shows the different temporal dynamics of these two modalities. The fMRI signal is delayed, relative to stimulus presentation and the Ca2+ signal. c) A zoomed in view of the Ca2+ and BOLD signals. Since the Ca2+ signal is collected at a relatively high temporal resolution (10 Hz), it appears in to be noisy. By zooming in the fast kinetics of these data are shown. No filtering of the Ca2+ signal has been applied.
Extended Data Fig. 6 Localization of Ca2+ and fMRI responses to stimuli.
a, A surface projection of the down-sampled Allen Atlas overlaid on the optical data of an example mouse. The ROI expected to respond to the presented unilateral hind limb stimuli is indicated with a dotted line. b, Responding ROIs from all mice from both modalities normalized to the maximum response amplitude. Calculated as follows if we had two instead of six mice: if voxel (i,j) for mouse #1 on average showed a 40% response relative to the maximum responding voxel for that mouse, and voxel (i,j) for mouse #2 on average showed a 60% response relative to the maximum responding voxel for mouse #2, then voxel (i,j) would be color-coded to 50%. The expected responding ROI from the Allen Atlas is shown as a dotted line. c, An example responding ROI from one mouse overlaid on the optical data. d, The same example responding ROIs (from the same mouse) overlaid on the projected MRI data.
Extended Data Fig. 7 Gamma-variant convolution model applied within responding ROIs identified by GLM.
a, Three example 50-second windows from N = 3 mice (left and middle panels). Ca2+ signal, averaged within the responding ROI, before (green) and after (blue) applying the gamma-variant convolution. BOLD signal (orange), averaged within the responding ROI. The average predicted hemodynamic response function (HRF) from these three examples (right panel). Goodness of fit assessed by correlating (Fisher’s Z transformed Pearson’s correlation) the Ca2+ signal convolved with the predicted HRF (blue) and the BOLD signal (orange). b, The correlations for N = 6 mice (n = 4 session, n = 11 windows, that is 44 data points per mouse) using filtered [0.04-0.1 Hz] (left) or unfiltered (middle) data. Each 50-second window contains the presentation of one unilateral hind-paw stimulus. The Ca2+ and BOLD responding ROIs are not fixed across mice, as illustrated (right) [reproduced from Extended Data Fig. 6]. For the boxplots, the central line is the median, the minima and maxima of the box extends to the 25th and 75th percentiles, whiskers extend to all data points, and outliers (data points beyond the 25th to 75th percentiles) are denoted by ‘+’.
Extended Data Fig. 8 Convolution model applied within Allen Atlas ROIs is not affected by window, frequency band, scan number or mouse.
Correlation between Ca2+ and BOLD signals. a, Correlation strengths compared across nine scans spanning the duration of our acquisitions. Scans where no stimulus are presented (grey), and during unilateral hind-paw stimulation (green), are different from the null (BOLD time points scrambled). b, Correlation strengths compared across mice. c, Correlation strengths compared between windows. d, Correlation strengths compared within different frequency filters. All show the same relationship to the null using a two-samples t-test. For boxplots, the central line is the median, the minima and maxima of the box extends to the 25th and 75th percentiles, whiskers extend to all data points, and outliers (data points beyond the 25th to 75th percentiles) are denoted by ‘+’. No correction for multiple comparisons was applied.
Extended Data Fig. 9 Parcellation results are independent of frequency filter across mice and parcel, variance is caused by parcel size.
a, Variance across parcels for each mouse for n = 5 frequency filters. b, Variance due to parcel for each filter for N = 6 mice. Neither frequency filter, nor mouse, captures the variance in the Dice coefficient observed. c, Variance is highly correlated with parcel size. d, Variance across frequency filters for N = 6 mice. For the boxplots, the central line is the median, the minima and maxima of the box extends to the 25th and 75th percentiles, whiskers extend to all data points, and outliers (data points beyond the 25th to 75th percentiles) are denoted by ‘+’.
Extended Data Fig. 10 Inter- vs. intra-hemisphere connectivity strength patterns between Ca2+ and BOLD vary regionally and by brain functional area.
a, and b, are reproduced for reference from Fig. 6. c, and d, are equivalent plots to b) showing the same information (inter-, purple, and intra-, grey, hemisphere regional connectivity strengths) for the three parcellations shown in a). b) shows results from the Allen atlas, c) shows results from the Ca2+ parcellation, and d) shows results from the BOLD parcellation. For the boxplots, the central line is the median, the minima and maxima of the box extends to the 25th and 75th percentiles, whiskers extend to all data points, and outliers (data points beyond the 25th to 75th percentiles) are denoted by ‘+’.
Supplementary information
Supplementary Information
Supplementary Figures 1–10
Supplementary Video 1
Accompanies Supplementary Fig. 3 (a). Shows raw (unprocessed) fluorescence signal (cyan wavelength) collected during a dual-imaging experiment. Video plays at 30 frames per second.
Supplementary Video 2
Accompanies Supplementary Fig. 3 (b). These are the same data as shown in Supplementary Fig 3.a. after preprocessing. In this example, unilateral hind-limb stimulation is delivered. The stimulus is ‘ON’ when the red square in the upper right corner of the video appears. Video plays at 30 frames per second.
Source data
Source Data Fig. 3
Statistical source data
Source Data Fig. 4
Statistical source data
Source Data Fig. 5
Statistical source data
Source Data Fig. 6
Statistical source data
Source Data Extended Data Fig. 5
Statistical source data
Source Data Extended Data Fig. 7
Statistical source data
Source Data Extended Data Fig. 8
Statistical source data
Source Data Extended Data Fig. 9
Statistical source data
Source Data Extended Data Fig. 10
Statistical source data
Rights and permissions
About this article
Cite this article
Lake, E.M.R., Ge, X., Shen, X. et al. Simultaneous cortex-wide fluorescence Ca2+ imaging and whole-brain fMRI. Nat Methods 17, 1262–1271 (2020). https://doi.org/10.1038/s41592-020-00984-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-00984-6
This article is cited by
-
Distinct neurochemical influences on fMRI response polarity in the striatum
Nature Communications (2024)
-
Global spatiotemporal synchronizing structures of spontaneous neural activities in different cell types
Nature Communications (2024)
-
Rapid fluctuations in functional connectivity of cortical networks encode spontaneous behavior
Nature Neuroscience (2024)
-
Modular subgraphs in large-scale connectomes underpin spontaneous co-fluctuation events in mouse and human brains
Communications Biology (2024)
-
Cortex-wide transcranial localization microscopy with fluorescently labeled red blood cells
Nature Communications (2024)