Abstract
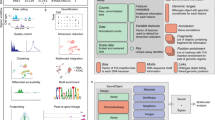
Recent technological advancements have enabled the profiling of a large number of genome-wide features in individual cells. However, single-cell data present unique challenges that require the development of specialized methods and software infrastructure to successfully derive biological insights. The Bioconductor project has rapidly grown to meet these demands, hosting community-developed open-source software distributed as R packages. Featuring state-of-the-art computational methods, standardized data infrastructure and interactive data visualization tools, we present an overview and online book (https://osca.bioconductor.org) of single-cell methods for prospective users.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
11 December 2019
A Correction to this paper has been published: https://doi.org/10.1038/s41592-019-0700-8
References
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 12, 115–121 (2015).
Robinson, M. D. et al. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Aryee, M. J. et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 30, 1363–1369 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Serratì, S. et al. Next-generation sequencing: advances and applications in cancer diagnosis. Onco. Targets Ther. 9, 7355–7365 (2016).
Nakato, R. & Shirahige, K. Recent advances in ChIP-seq analysis: from quality management to whole-genome annotation. Brief. Bioinform. 18, 279–290 (2017).
Kukurba, K. R. & Montgomery, S. B. RNA sequencing and analysis. Cold Spring Harb. Protoc. 2015, 951–969 (2015).
Kolodziejczyk, A. A., Kim, J. K., Svensson, V., Marioni, J. C. & Teichmann, S. A. The technology and biology of single-cell RNA sequencing. Mol. Cell 58, 610–620 (2015).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–401 (2014).
Tirosh., I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Karaayvaz, M. et al. Unravelling subclonal heterogeneity and aggressive disease states in TNBC through single-cell RNA-seq. Nat. Commun. 9, 3588 (2018).
Jean Fan. et al. Linking transcriptional and genetic tumor heterogeneity through allele analysis of single-cell RNA-seq data. Genome Res. 28, 1217–1227 (2018).
Levitin, H. M., Yuan, J. & Sims, P. A. Single-cell transcriptomic analysis of tumor heterogeneity. Trends Cancer 4, 264–268 (2018).
Paulson, K. G. et al. Acquired cancer resistance to combination immunotherapy from transcriptional loss of class I HLA. Nat. Commun. 9, 3868 (2018).
Zeisel, A. et al. Brain structure: cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Deng, Q., Ramsköld, D., Reinius, B. & Sandberg, R. Single-cell RNA-seq reveals dynamic, random monoallelic gene expression in mammalian cells. Science 343, 193–196 (2014).
Kiselev, V. Y., Andrews, T. S. & Hemberg, M. Challenges in unsupervised clustering of single-cell RNA-seq data. Nat. Rev. Genet. 20, 273–282 (2019).
Cannoodt, R., Saelens, W. & Saeys, Y. Computational methods for trajectory inference from single-cell transcriptomics. Eur. J. Immunol. 46, 2496–2506 (2016).
Regev, A. et al. The Human cell atlas. eLife 6, e27041 (2017).
Rozenblatt-Rosen, O., Stubbington, M. J. T., Regev, A. & Teichmann, S. A. The human cell atlas: from vision to reality. Nature 550, 451–453 (2017).
Han, X. et al. Mapping the mouse cell atlas by microwell-seq. Cell 173, 1307 (2018).
McDavid, A. et al. Data exploration, quality control and testing in single-cell qPCR-based gene expression experiments. Bioinformatics 29, 461–467 (2013).
Hicks, S. C., Townes, F. W., Teng, M. & Irizarry, R. A. Missing data and technical variability in single-cell RNA-sequencing experiments. Biostatistics 19, 562–578 (2018).
Kharchenko, P. V., Silberstein, L. & Scadden, D. T. Bayesian approach to single-cell differential expression analysis. Nat. Methods 11, 740–742 (2014).
Finak, G. et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 278 (2015).
Lun, A. T. L., Bach, K. & Marioni, J. C. Pooling across cells to normalize single-cell RNA sequencing data with many zero counts. Genome Biol. 17, 75 (2016).
Ji, Z. & Ji, H. TSCAN: Pseudo-time reconstruction and evaluation in single-cell RNA-seq analysis. Nucleic Acids Res. 44, e117 (2016).
Risso, D., Perraudeau, F., Gribkova, S., Dudoit, S. & Vert, J.-P. A general and flexible method for signal extraction from single-cell RNA-seq data. Nat. Commun. 9, 284 (2018).
Chambers, J. M. Object-oriented programming, functional programming and R. Stat. Sci. 29, 167–180 (2014).
Tian, L. et al. scPipe: a flexible R/Bioconductor preprocessing pipeline for single-cell RNA-sequencing data. PLoS Comput. Biol. 14, e1006361 (2018).
Wang, Z., Hu, J., Johnson, W. E. & Campbell, J. D. scruff: an R/Bioconductor package for preprocessing single-cell RNA-sequencing data. BMC Bioinform. 20, 222 (2019).
Lun, AaronT. L. et al. Emptydrops: distinguishing cells from empty droplets in droplet-based single-cell RNA sequencing data. Genome Biol. 20, 63 (2019).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Melsted, P. et al. Modular and efficient pre-processing of single-cell rna-seq. Preprint at bioRxiv https://doi.org/10.1101/673285 (2019).
Srivastava, A., Malik, L., Smith, T., Sudbery, I. & Patro, R. Alevin efficiently estimates accurate gene abundances from dscRNA-seq data. Genome Biol. 20, 65 (2019).
Griffiths, J. A., Richard, A. C., Bach, K., Lun, A. T. L. & Marioni, J. C. Detection and removal of barcode swapping in single-cell RNA-seq data. Nat. Commun. 9, 2667 (2018).
Bais, A. S. & Kostka, D. scds: computational annotation of doublets in single cell RNA sequencing data. Bioinformatics https://doi.org/10.1093/bioinformatics/btz698 (2019).
Ilicic, T. et al. Classification of low quality cells from single-cell RNA-seq data. Genome Biol. 17, 29 (2016).
McCarthy, D. J., Campbell, K. R., Lun, A. T. L. & Wills, Q. F. Scater: pre-processing, quality control, normalization and visualization of single-cell RNA-seq data in R. Bioinformatics 33, 1179–1186 (2017).
Vallejos, C. A., Risso, D. R., Scialdone, A., Dudoit, S. & Marioni, J. C. Normalizing single-cell RNA sequencing data: challenges and opportunities. Nat. Methods 14, 565–571 (2017).
Vallejos, C. A., Richardson, S. & Marioni, J. C. Beyond comparisons of means: understanding changes in gene expression at the single-cell level. Genome Biol. 17, 70 (2016).
Huang, M. et al. SAVER: gene expression recovery for single-cell RNA sequencing. Nat. Methods 15, 539–542 (2018).
Li, W. V. & Li, J. L. An accurate and robust imputation method scImpute for singlecell RNA-seq data. Nat. Commun. 9, 997 (2018).
Svensson, V. Droplet scRNA-seq is not zero-inflated. Preprint bioRxiv https://doi.org/10.1101/582064 (2019).
Vieth, B., Ziegenhain, C., Parekh, S., Enard, W. & Hellmann, I. powsimR: power analysis for bulk and single cell RNA-seq experiments. Bioinformatics 33, 3486–3488 (2017).
Townes, F. W., Hicks, S. C., Aryee, M. J. & Irizarry, R. A. Feature selection and dimension reduction for single cell RNA-seq based on a multinomial model. Preprint at bioRxiv https://doi.org/10.1101/574574 (2019).
Andrews, T. & Hemberg, M. False signals induced by single-cell imputation. F1000Res. https://doi.org/10.12688/f1000research.16613.2 (2019).
Andrews, T. & Hemberg, M. M3Drop: Dropout-based feature selection for scRNASeq. Bioinformatics 35, 2865–2867 (2019).
Yip, S. H., Sham, P. C. & Wang, J. Evaluation of tools for highly variable gene discovery from single-cell RNA-seq data. Brief. Bioinform. 20, 1583–1589 (2018).
Lun, A. T. L., McCarthy, D. J. & Marioni, J. C. A step-by-step workflow for low-level analysis of single-cell RNA-seq data with Bioconductor. F1000Res. 5, 2122 (2016).
van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Melville, J., McInnes, L. & Healy, J. UMAP: uniform manifold approximation and projection for dimension reduction. Preprint at arXiv https://arxiv.org/abs/1802.03426 (2018).
Angerer., P. et al. Destiny: diffusion maps for large-scale single-cell data in R. Bioinformatics 32, 1241–1243 (2016).
Haghverdi, L., Lun, A. T. L., Morgan, M. D. & Marioni, J. C. Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors. Nat. Biotechnol. 36, 421–427 (2018).
Lin, Y. et al. scMerge leverages factor analysis, stable expression, and pseudoreplication to merge multiple single-cell RNA-seq datasets. Proc. Natl. Acad. Sci. USA 116, 9775–9784 (2019).
Kiselev, V. Y., Yiu, A. & Hemberg, M. scmap: projection of single-cell RNA-seq data across data sets. Nat. Methods 15, 359–362 (2018).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Wang, B., Zhu, J., Pierson, E., Ramazzotti, D. & Batzoglou, S. Visualization and analysis of single-cell RNA-seq data by kernel-based similarity learning. Nat. Methods 14, 414–416 (2017).
Kiselev, V. Y. et al. SC3: consensus clustering of single-cell RNA-seq data. Nat. Methods 14, 483–486 (2017).
Risso, D. et al. clusterExperiment and RSEC: a bioconductor package and framework for clustering of singlecell and other large gene expression datasets. PLoS Comp. Biol. 14, e1006378–16 (2018).
Van den Berge, K. et al. Observation weights unlock bulk RNA-seq tools for zero inflation and single-cell applications. Genome Biol. 19, 24 (2018).
Korthauer, K. D. et al. A statistical approach for identifying differential distributions in single-cell RNA-seq experiments. Genome Biol. 17, 222 (2016).
Soneson, C. & Robinson, M. D. Bias, robustness and scalability in single-cell differential expression analysis. Nat. Methods 15, 255–261 (2018).
Wang, T., Li, B., Nelson, C. E. & Nabavi, S. Comparative analysis of differential gene expression analysis tools for single-cell RNA sequencing data. BMC Bioinform. 20, 40 (2019).
Crowell, H. L. et al. On the discovery of population-specific state transitions from multi-sample multi-condition single-cell RNA sequencing data. Preprint at bioRxiv https://doi.org/10.1101/713412 (2019).
Andrews, T. S. & Hemberg, M. Identifying cell populations with scRNASeq. Mol. Asp. Med. 59, 114–122 (2018).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Campbell, K. R. & Yau, C. switchde: inference of switch-like differential expression along single-cell trajectories. Bioinformatics 33, 1241–1242 (2017).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 19, 477 (2018).
duVerle, D. A., Yotsukura, S., Nomura, S., Aburatani, H. & Tsuda, K. CellTree: an R/bioconductor package to infer the hierarchical structure of cell populations from single-cell RNA-seq data. BMC Bioinform. 17, 363 (2016).
Campbell, K. R. & Yau, C. Probabilistic modeling of bifurcations in single-cell gene expression data using a bayesian mixture of factor analyzers. Wellcome Open Res. 2, 19 (2017).
Saelens, W., Cannoodt, R., Todorov, H. & Saeys, Y. A comparison of single-cell trajectory inference methods. Nat. Biotechnol. 37, 547 (2019).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, 353–361 (2017).
Fabregat, A. et al. The reactome pathway knowledgebase. Nucleic Acids Res. 44, 481–487 (2015).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Geistlinger, L., Csaba, G. & Zimmer, R. Bioconductor’s EnrichmentBrowser: seamless navigation through combined results of set and network-based enrichment analysis. BMC Bioinform. 17, 45 (2016).
Alhamdoosh, M. et al. Combining multiple tools outperforms individual methods in gene set enrichment analyses. Bioinformatics 33, 414–424 (2017).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Buettner, F., Pratanwanich, N., McCarthy, D. J., Marioni, J. C. & Stegle, O. fscLVM: scalable and versatile factor analysis for single-cell RNA-seq. Genome Biol. 18, 212 (2017).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Zappia, L., Phipson, B. & Oshlack, A. Splatter: simulation of single-cell RNA sequencing data. Genome Biol. 18, 174 (2017).
Kimes, P. K. & Reyes, A. Reproducible and replicable comparisons using SummarizedBenchmark. Bioinformatics 35, 137–139 (2019).
Tian, L. et al. Benchmarking single cell RNA-sequencing analysis pipelines using mixture control experiments. Nat. Methods 16, 479–487 (2019).
Rue-Albrecht, K., Marini, F., Soneson, C. & Lun, A. T. L. iSEE: interactive SummarizedExperiment Explorer. F1000Res. 7, 741 (2018).
Peterson, V. M. et al. Multiplexed quantification of proteins and transcripts in single cells. Nat. Biotechnol. 35, 936–939 (2017).
Dey, S. S., Kester, L., Spanjaard, B., Bienko, M. & van Oudenaarden, A. Integrated genome and transcriptome sequencing of the same cell. Nat. Biotechnol. 33, 285–289 (2015).
Macaulay, IainC. et al. Separation and parallel sequencing of the genomes and transcriptomes of single cells using GT-seq. Nat. Protoc. 11, 2081–2103 (2016).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017).
Shahi, P., Kim, S. C., Haliburton, J. R., Gartner, Z. J. & Abate, A. R. Abseq: ultrahighthroughput single cell protein profiling with droplet microfluidic barcoding. Sci. Rep. 7, 44447 (2017).
Angermueller, C. et al. Parallel single-cell sequencing links transcriptional and epigenetic heterogeneity. Nat. Methods 13, 229–232 (2016).
Cao, J. et al. Joint profiling of chromatin accessibility and gene expression in thousands of single cells. Science 361, 1380–1385 (2018).
Clark, S. J. et al. scNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells. Nat. Commun. 9, 781 (2018).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Eddelbuettel, D. & François, R. Rcpp: seamless R and C++ integration. J. Stat. Softw. 40, 1–18 (2011).
Acknowledgements
Bioconductor is supported by the National Human Genome Research Institute (NHGRI) and National Cancer Institute (NCI) of the National Institutes of Health (NIH) (grant no. U41HG004059, U24CA180996), the European Union (EU) H2020 Personalizing Health and Care Program Action (contract number 633974) and the SOUND Consortium. In addition, M.M., S.C.H., R.G., W.H., A.T.L.L. and D.R. are supported by the Chan Zuckerberg Initiative (CZI) DAF (grant no. 2018-183201, 2018-183560), an advised fund of Silicon Valley Community Foundation. D.R., W.H., M.M. and S.C.H. are supported by 2019-002443 from the CZI. S.C.H. is supported by the NIH/NHGRI (grant no. R00HG009007). R.A.A. and R.G. are supported by the Integrated Immunotherapy Research Center at Fred Hutch. M.M. is supported by the NCI/NHGRI (grant no. U24CA232979). L.G. is supported by a research fellowship from the German Research Foundation (grant no. GE3023/1-1). L.W. and V.J.C. are supported by the NCI (grant no. U24CA18099). V.J.C. is additionally supported by NCI U01 CA214846 and Chan Zuckerberg Initiative DAF (grant no. 2018-183436). ATLL received support from CRUK (grant no. A17179) and the Wellcome Trust (grant no. WT/108437/Z/15). F.M. is supported by the German Federal Ministry of Education and Research (grant no. BMBF 01EO1003). M.L.S. is supported by the German Network for Bioinformatics Infrastructure (grant no. 031A537B). D.R. is supported by the Programma per Giovani Ricercatori Rita Levi Montalcini from the Italian Ministry of Education, University and Research. H.P. is supported by the NIH Bioconductor grant (no. U41HG004059).
Author information
Authors and Affiliations
Contributions
E.B., V.J.C., L.N.C., L.G., F.M., K.R., D.R., C.S. and L.W. contributed equally to this work. S.C.H. and R.G. contributed equally to the supervision of this work. S.C.H. and R.G. conceptualized the manuscript. R.A.A., A.T.L.L., S.C.H. and R.G. wrote the manuscript with contributions and input from all authors. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.G. declares ownership in CellSpace Biosciences.
Additional information
Peer review information Lei Tang was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Amezquita, R.A., Lun, A.T.L., Becht, E. et al. Orchestrating single-cell analysis with Bioconductor. Nat Methods 17, 137–145 (2020). https://doi.org/10.1038/s41592-019-0654-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0654-x
This article is cited by
-
ReUseData: an R/Bioconductor tool for reusable and reproducible genomic data management
BMC Bioinformatics (2024)
-
A comparison of marker gene selection methods for single-cell RNA sequencing data
Genome Biology (2024)
-
cytoviewer: an R/Bioconductor package for interactive visualization and exploration of highly multiplexed imaging data
BMC Bioinformatics (2024)
-
Library size confounds biology in spatial transcriptomics data
Genome Biology (2024)
-
Single-cell RNA sequencing reveals peripheral blood leukocyte responses to spinal cord injury in mice with humanised immune systems
Journal of Neuroinflammation (2024)