Abstract
The European XFEL (EuXFEL) is a 3.4-km long X-ray source, which produces femtosecond, ultrabrilliant and spatially coherent X-ray pulses at megahertz (MHz) repetition rates. This X-ray source has been designed to enable the observation of ultrafast processes with near-atomic spatial resolution. Time-resolved crystallographic investigations on biological macromolecules belong to an important class of experiments that explore fundamental and functional structural displacements in these molecules. Due to the unusual MHz X-ray pulse structure at the EuXFEL, these experiments are challenging. Here, we demonstrate how a biological reaction can be followed on ultrafast timescales at the EuXFEL. We investigate the picosecond time range in the photocycle of photoactive yellow protein (PYP) with MHz X-ray pulse rates. We show that difference electron density maps of excellent quality can be obtained. The results connect the previously explored femtosecond PYP dynamics to timescales accessible at synchrotrons. This opens the door to a wide range of time-resolved studies at the EuXFEL.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Data has been deposited with the Coherent X-ray Imaging Data Bank73 with CXIDB ID 100. This includes: stream files for all data and for data separated into each time delay, MTZ and PDB files for all time delays, including the dark/reference structures. We have deposited data (mtz-files and structures) for the 10 ps, 30 ps and 80 ps time delays, as well as the dark3 (30 ps) and pure dark reference structures, with the Protein Data Bank, with deposition codes 6P4I, 6P5D, 6P5E, 6P5G and 6P5F, respectively.
Code availability
Linux scripts and Fortran source codes for the calculation of weighted difference maps, extrapolated electron density maps and the integration of negative densities within a spherical volume are included in a demonstration, which is available online as Supplementary Data.
References
Moffat, K. Time-resolved biochemical crystallography: a mechanistic perspective. Chem. Rev. 101, 1569–1581 (2001).
Schmidt, M. Time-resolved macromolecular crystallography at modern X-ray sources. Methods Mol. Biol. 1607, 273–294 (2017).
Aquila, A. et al. Time-resolved protein nanocrystallography using an X-ray free-electron laser. Opt. Express 20, 2706–2716 (2012).
Tenboer, J. et al. Time-resolved serial crystallography captures high-resolution intermediates of photoactive yellow protein. Science 346, 1242–1246 (2014).
Chapman, H. N. et al. Femtosecond X-ray protein nanocrystallography. Nature 470, 73–77 (2011).
Boutet, S. et al. High-resolution protein structure determination by serial femtosecond crystallography. Science 337, 362–364 (2012).
Lomb, L. et al. Radiation damage in protein serial femtosecond crystallography using an X-ray free-electron laser. Phys. Rev. B 84, 214111 (2011).
Nass, K. et al. Indications of radiation damage in ferredoxin microcrystals using high-intensity X-FEL beams. J. Synchrotron Radiat. 22, 225–238 (2015).
Suga, M. et al. Light-induced structural changes and the site of O=O bond formation in PSII caught by XFEL. Nature 543, 131–135 (2017).
Chreifi, G. et al. Crystal structure of the pristine peroxidase ferryl center and its relevance to proton-coupled electron transfer. Proc. Natl Acad. Sci. USA 113, 1226–1231 (2016).
Wiedorn, M. O. et al. Megahertz serial crystallography. Nat. Commun. 9, 4025 (2018).
Grünbein, M. L. et al. Megahertz data collection from protein microcrystals at an X-ray free-electron laser. Nat. Commun. 9, 3487 (2018).
Barends, T. R. et al. Direct observation of ultrafast collective motions in CO myoglobin upon ligand dissociation. Science 350, 445–450 (2015).
Pande, K. et al. Femtosecond structural dynamics drives the trans/cis isomerization in photoactive yellow protein. Science 352, 725–729 (2016).
Meyer, T. E., Yakali, E., Cusanovich, M. A. & Tollin, G. Properties of a water-soluble, yellow protein isolated from a halophilic phototrophic bacterium that has photochemical activity analogous to sensory rhodopsin. Biochemistry 26, 418–423 (1987).
Genick, U. K. et al. Structure of a protein photocycle intermediate by millisecond time-resolved crystallography. Science 275, 1471–1475 (1997).
Ihee, H. et al. Visualizing reaction pathways in photoactive yellow protein from nanoseconds to seconds. Proc. Natl Acad. Sci. USA 102, 7145–7150 (2005).
Kort, R. et al. Evidence for trans–cis isomerization of the p-coumaric acid chromophore as the photochemical basis of the photocycle of photoactive yellow protein. FEBS Lett. 382, 73–78 (1996).
Polli, D. et al. Conical intersection dynamics of the primary photoisomerization event in vision. Nature 467, 440–443 (2010).
Mathes, T. et al. Femto- to microsecond photodynamics of an unusual bacteriophytochrome. J. Phys. Chem. Lett. 6, 5 (2014).
Ali, A. M. et al. Optogenetic inhibitor of the transcription factor CREB. Chem. Biol. 22, 1531–1539 (2015).
Schotte, F. et al. Watching a signaling protein function in real time via 100-ps time-resolved Laue crystallography. Proc. Natl Acad. Sci. USA 109, 19256–19261 (2012).
Creelman, M., Kumauchi, M., Hoff, W. D. & Mathies, R. A. Chromophore dynamics in the PYP photocycle from femtosecond stimulated Raman spectroscopy. J. Phys. Chem. B 118, 659–667 (2014).
Palmer, G. et al. Pump-probe laser system at the FXE and SPB/SFX instruments of the European X-ray free-electron laser facility. J. Synchrotron Radiat. 26, 328–332 (2019).
Schmidt, M. et al. Protein energy landscapes determined by five-dimensional crystallography. Acta Crystallogr. D 69, 2534–2542 (2013).
Prokhorenko, V. I. et al. Coherent control of retinal isomerization in bacteriorhodopsin. Science 313, 1257–1261 (2006).
Mancuso, A. P. et al. The single particles, clusters and biomolecules and serial femtosecond crystallography instrument of the European XFEL: initial installation. J. Synchrotron Radiat. 26, 660–676 (2019).
Allahgholi, A. et al. The adaptive gain integrating pixel detector at the European XFEL. J. Synchrotron Radiat. 26, 74–82 (2019).
Stan, C. A. et al. Liquid explosions induced by X-ray laser pulses. Nat. Phys. 12, 966–971 (2016).
Tripathi, S., Srajer, V., Purwar, N., Henning, R. & Schmidt, M. pH dependence of the photoactive yellow protein photocycle investigated by time-resolved crystallography. Biophys. J. 102, 325–332 (2012).
Schmidt, M. in Ultrashort Laser Pulses in Medicine and Biology (eds Braun, M. et al.) 201–241 (Springer, 2008).
Schmidt, M. Time-resolved macromolecular crystallography at pulsed X-ray sources. Int. J. Mol. Sci. 20, 1401 (2019).
Jung, Y. O. et al. Volume-conserving trans–cis isomerization pathways in photoactive yellow protein visualized by picosecond X-ray crystallography. Nat. Chem. 5, 212–220 (2013).
Hutchison, C. D. M. & van Thor, J. J. Populations and coherence in femtosecond time resolved X-ray crystallography of the photoactive yellow protein. Int, Rev. Phys. Chem. 36, 117–143 (2017).
Groenhof, G. et al. Photoactivation of the photoactive yellow protein: why photon absorption triggers a trans-to-cis isomerization of the chromophore in the protein. J. Am. Chem. Soc. 126, 4228–4233 (2004).
Markovitch, O. & Agmon, N. Structure and energetics of the hydronium hydration shells. J. Phys. Chem. A 111, 2253–2256 (2007).
Levantino, M. et al. Ultrafast myoglobin structural dynamics observed with an X-ray free-electron laser. Nat. Commun. 6, 6772 (2015).
DePonte, D. P. et al. Gas dynamic virtual nozzle for generation of microscopic droplet streams. J. Phys. D 41, 195505 (2008).
Schmidt, M., Rajagopal, S., Ren, Z. & Moffat, K. Application of singular value decomposition to the analysis of time-resolved macromolecular X-ray data. Biophys. J. 84, 2112–2129 (2003).
Rajagopal, S., Schmidt, M., Anderson, S., Ihee, H. & Moffat, K. Analysis of experimental time-resolved crystallographic data by singular value decomposition. Acta Crystallogr. D 60, 860–871 (2004).
Kang, Y. et al. Crystal structure of rhodopsin bound to arrestin by femtosecond X-ray laser. Nature 523, 561–567 (2015).
Kupitz, C. et al. Structural enzymology using X-ray free electron lasers. Struct. Dyn. 4, 044003 (2017).
Olmos, J. L. Jr. et al. Enzyme intermediates captured “on the fly” by mix-and-inject serial crystallography. BMC Biol. 16, 59 (2018).
Paul, K. et al. Coherent control of an opsin in living brain tissue. Nat. Phys. 13, 1111–1116 (2017).
Wang, J. et al. Time-resolved protein activation by proximal decaging in living systems. Nature 569, 509–513 (2019).
Pandey, S., Bean, R., Sato, T., Mancuso, A. P. & Schmidt, M. Time-resolved serial femtosecond crystallography at the European X-ray free electron laser. Prot. Exch. https://doi.org/10.21203/rs.2.14634/v1 (2019).
Mancuso, A. P., Aquila, A., Borchers, G., Giewekemeyer, K. & Reimers, N. Technical Design Report: Scientific Instrument Single Particles, Clusters, and Biomolecules (SPB) https://doi.org/10.3204/XFEL.EU/TR-2013-004 (XFEL.EU, 2013).
Echelmeier, A. et al. 3D printed droplet generation devices for serial femtosecond crystallography enabled by surface coating. J. Appl. Cryst. 52, 997–1008 (2019).
Hutchison, C. D. M. et al. Photocycle populations with femtosecond excitation of crystalline photoactive yellow protein. Chem. Phys. Lett. 654, 63–71 (2016).
Nass Kovacs, G. et al. Three-dimensional view of ultrafast dynamics in photoexcited bacteriorhodopsin. Nat. Comm. 10, 3177 (2019).
Bean, R. J., Aquila, A., Samoylova, L. & Mancuso, A. P. Design of the mirror optical systems for coherent diffractive imaging at the SPB/SFX instrument of the European XFEL. J. Opt. 18, 074011 (2016).
Greiffenberg, D. The AGIPD detector for the European XFEL. J. Instrum. 7, CO1103 (2012).
Fangohr, H. et al. Data analysis support in Karabo at European XFEL. In Proc. of International Conference on Accelerator and Large Experimental Control Systems (eds Costa, I. et al.) 245–252 (inSPIRE, 2017).
Boukhelef, D., Szuba, J., Wrona, K. & Youngman, C. Software Development for High Speed Data Recording and Processing (JACoW2014).
Kirkwood, H. J. et al. Initial observations of the femtosecond timing jitter at the European XFEL. Opt. Lett. 44, 1650–1653 (2019).
Mariani, V. et al. OnDA: online data analysis and feedback for serial X-ray imaging. J. Appl. Crystallogr. 49, (1073–1080 (2016).
Barty, A. et al. Cheetah: software for high-throughput reduction and analysis of serial femtosecond X-ray diffraction data. J. Appl. Crystallogr. 47, 1118–1131 (2014).
White, T. A. et al. Recent developments in CrystFEL. J. Appl. Cryst. 49, 680–689 (2016).
Gevorkov, Y. et al. XGANDALF – extended gradient descent algorithm for lattice finding. Acta Cryst. Found. Adv. 75, 694–704 (2019).
Yefanov, O. et al. Accurate determination of segmented X-ray detector geometry. Opt. Express 23, 28459–28470 (2015).
Brehm, W. & Diederichs, K. Breaking the indexing ambiguity in serial crystallography. Acta Crystallogr. D 70, 101–109 (2014).
Glownia, J. M. et al. Time-resolved pump-probe experiments at the LCLS. Opt. Express 18, 17620–17630 (2010).
Harmand, M. et al. Achieving few-femtosecond time-sorting at hard X-ray free-electron lasers. Nat. Photonics 7, 215–218 (2013).
Bionta, M. R. et al. Spectral encoding of X-ray/optical relative delay. Opt. Express 19, 21855–21865 (2011).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D 67, 355–367 (2011).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011).
Ren, Z. et al. A molecular movie at 1.8 Å resolution displays the photocycle of photoactive yellow protein, a eubacterial blue-light receptor, from nanoseconds to seconds. Biochemistry 40, 13788–13801 (2001).
Drenth, J. Principles of Protein X-Ray Crystallography (Springer, 1999)..
Terwilliger, T. C. & Berendzen, J. Bayesian difference refinement. Acta Crystallogr. D 52, 1004–1011 (1996).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Nogly, P. et al. Retinal isomerization in bacteriorhodopsin captured by a femtosecond X-ray laser. Science 361, eaat0094 (2018).
Richards, F. M. & Kundrot, C. E. Identification of structural motifs from protein coordinate data—secondary structure and 1st-level supersecondary structure. Proteins 3, 71–84 (1988).
Maia, F. R. The coherent X-ray imaging data bank. Nat. Methods 9, 854–855 (2012).
Acknowledgements
We acknowledge European XFEL in Schenefeld, Germany, for provision of X-ray free-electron laser beamtime at Scientific Instrument SPB/SFX and thank the instrument group and facility staff for their assistance. This work was supported by NSF Science and Technology Centers (grant no. NSF-1231306; Biology with X-ray Lasers) to H.N.C., P.F., A.O., A.R., P.S. and M.S.). CFEL (H.N.C.) is supported by the Gottfried Wilhelm Leibniz Program of the DFG; the project ‘X-probe’ funded by the European Union’s 2020 Research and Innovation Program under the Marie Sklodowska-Curie grant agreement (no. 637295); the European Research Council, ‘Frontiers in Attosecond X-ray Science: Imaging and Spectroscopy (AXSIS)’ (no. ERC-2013-SyG 609920, together with P.F.); and the Human Frontiers Science Program grant (no. RGP0010 2017). P.F. and A.R. acknowledge the support of funding from the Biodesign Center for Applied Structural Discovery at Arizona State University and NSF award (no. 1565180). Funding from the National Institutes of Health (grant nos. R01GM095583 to P.F. and R01GM117342 to M.F.) is also acknowledged.
Author information
Authors and Affiliations
Contributions
S.P., I.P. and M.S. expressed, purified and crystallized the protein. R.B., T.S., J.B., V.B., M.E., G.G., M.J., Y.K., H.K., A.K., R.L., L.M., T.M., G.P., M.R., A.S., J.S.-D. and A.P.M. operated the SPB/SFX instrument. S.L., J.K., R.S. and H.N.C. provided injector nozzles. J.C.V., C.K., M.H., M.H.A., J.K., F.H.M.K., S.L., V.Maz., D.M., R.S. and A.T. collected the data. S.P., I.P., O.Y., V.Mar., T.A.W., Y.G., A.O., P.S., A.T. and A.B. processed the data. S.P., I.P., P.S., A.O. and M.S. analyzed the data. C.K., M.H.A., R.F. and P.F. logged the experiment. J.C.V., A.E., D.D., D.K. and A.R. conceived and operated the oil co-flow. R.B., T.S., M.F., H.N.C., A.R., A.B., P.F., A.P.M. and M.S. designed the experiment. S.P., S.B., A.B., P.F., A.P.M. and M.S. wrote the manuscript with input from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Arunima Singh and Allison Doerr were the primary editors on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
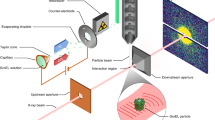
Supplementary Fig. 1 Setup of a MHz TR-SFX experiment at the EuXFEL (modified from Wiedorn et al.11).
X-ray pulses arrive in 1.13 MHz bursts which repeat every 100 ms. There are 176 X-ray pulses in the burst. The KB-mirror system focuses the X-ray beam to a 2–3 μm focal spot. The fs-laser delivers 376 kHz pulses (λ=420 nm, blue) synchronized to the X-ray pulses. The laser focus is 42 μm Ø in the X-ray interaction region (dotted circle). The microcrystals are mixed with fluorinated oil and injected by a GDVN. The jet produced by the GDVN, the laser beam as well as the X-ray pulses precisely intersect. The time-resolved diffraction patterns are collected by the AGIPD. Diffraction patterns with common time-delays were separated based on the pulse ID (see also Fig. 2b) and combined to datasets.
Supplementary Fig. 2 Hit and indexing rates.
a, Hit rates (red) and indexing rates (black) with 1.13 MHz X-ray pulse repetition rate. Note, the strong drop of the hit-rate after the first pulse from 2% to 1%. 472,528 total patterns, 41,559 hits and 24,815 indexed patterns were separated on the basis of pulse IDs. From these, hit rates and indexing rates were calculated. b, Hit rates (red) and indexing rates (black) with 564 kHz X-ray pulse repetition. The overall hit rate is about 2%. 52,495,158 total patterns, 304,673 hits and 142,948 indexed patterns were separated on the basis of pulse IDs from which hit rates and indexing rates were calculated. Blue solid line in a and b, X-ray pulse energy (on arbitrary scale). The indexing rate varies only slightly and is about 40% - 60%.
Supplementary Fig. 4 Excitation and ultrafast displacements in PYP.
a, Structure of PYP. Some important residues in the chromophore (pCA) binding pocket are marked. The M41-71 moiety (residues 41 to 71) is marked in red. Helix H74-88 is marked. b, Dark state spectra of PYP. Black: measured in solution, red: in the crystal. The wavelength at the absorption maximum is marked. Excitation has been achieved with 240 fs laser pulses with λ=420 nm. c, Solid spheres: root mean square displacements of 31 Cα atoms in M41-71 relative to the dark (reference) structure, red spheres: from data measured at EuXFEL. Dashed line: fit by a function consisting of an exponential, a strongly damped, phase shifted cosine function and a straight line as outlined in the text.
Supplementary Fig. 5 Difference distance matrices evaluated for Cα atoms of residues 42 to 93.
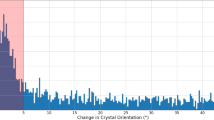
The green line denotes the M41-71 moiety. The scale on top is in Å. a - d, Difference distance matrices derived from structures at 10 ps, 30 ps, 80 ps and 100 ps relative to that at 3 ps, respectively. Difference distances are also shown for helix H74-88.
Supplementary Fig. 6 Signal levels in the DED map at the 30 ps delay.
The DED map at 30 ps is overlaid on the entire PYP and contoured from +/- 2σ to +/- 4σ in steps of 0.5σ. Red: negative DED, green: positive DED. The 3σ level, c, is the best compromise to distinguish the signal, for example on the pCA chromophore, from spurious noise features distributed within the protein volume.
Supplementary Fig. 7 Method to determine the factor N and the population transfer (PT).
The factor N has been determined to calculate extrapolated, conventional maps from data collected at various X-ray sources. Black spheres: summed absolute negative DED in a sphere of R = 4 Å centered on the PCA chromophore double bond. Red dotted lines: the more horizontal line follows the initial slope of the data; the second line delineates the constant incline with larger Ns. The Next (in brackets) can be estimated from the intersection of the two lines. a, 3ps data from CXI at LCLS collected with fs laser excitation in the absorption maximum (Pande et al. 14). Factor N = 16, PT = 12.5 %, insert: 1 μs data collected with ns laser excitation. N = 4, and PT = 50% (Tenboer et al.4). b, c, and d, Factors N for the 10 ps, 30 ps, and 80 ps data collected at the EXFEL with fs laser excitation outside the absorption maximum. PT is about 7 % throughout. Insert in d, 100 ps data collected at APS (about 6% PT, Jung et al.33). 13,214, 13,542, 13,722, 13,142, 13,014 and 12,889 observed difference amplitudes are used to determine extrapolated maps for the 100ps, 1 μs, 3ps, 10ps, 30ps and 80ps time delays, respectively.
Supplementary Fig. 8 Observed and calculated difference electron densities (DED) near the pCA chromophore.
Left panels: observed difference electron density (blue: 3 σ, red: -3 σ contour levels). Right panels: calculated difference electron density (blue: 4 σ, red: -4 σ contour levels). Yellow model: structure of the dark (reference) state; blue model: structure at a particular time delay. a, 10 ps; b, 30 ps, c, 80 ps. In panel b pairwise difference density features are marked with α (negative) and β (positive). The feature γ shows the signal caused by the Cys-69 sulfur. The marked DED features can be readily detected at the other time delays. 13,142, 13,014 and 12,889 difference amplitudes were used to calculate the observed DED maps for a, b and c, respectively.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8 and Tables 1–7.
Supplementary Software
demo.tar.zip. Compressed repository containing a demonstration and software for difference map calculation and structure determination. After the TR-SFX experiment datasets of reference (dark) and time-dependent intensities are available. This demonstration guides through the processes of difference map calculation and structure determination from extrapolated electron density maps.
Rights and permissions
About this article
Cite this article
Pandey, S., Bean, R., Sato, T. et al. Time-resolved serial femtosecond crystallography at the European XFEL. Nat Methods 17, 73–78 (2020). https://doi.org/10.1038/s41592-019-0628-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0628-z
This article is cited by
-
Thermoelastic effects in Bragg reflectors as a potential bottleneck for XFELs with megahertz repetition rate
Communications Physics (2024)
-
Helium-electrospray improves sample delivery in X-ray single-particle imaging experiments
Scientific Reports (2024)
-
Directed ultrafast conformational changes accompany electron transfer in a photolyase as resolved by serial crystallography
Nature Chemistry (2024)
-
Heterogeneity in M. tuberculosis β-lactamase inhibition by Sulbactam
Nature Communications (2023)
-
Watching the release of a photopharmacological drug from tubulin using time-resolved serial crystallography
Nature Communications (2023)