Abstract
Human cortical organoids (hCOs), derived from human embryonic stem cells (hESCs), provide a platform to study human brain development and diseases in complex three-dimensional tissue. However, current hCOs lack microvasculature, resulting in limited oxygen and nutrient delivery to the inner-most parts of hCOs. We engineered hESCs to ectopically express human ETS variant 2 (ETV2). ETV2-expressing cells in hCOs contributed to forming a complex vascular-like network in hCOs. Importantly, the presence of vasculature-like structures resulted in enhanced functional maturation of organoids. We found that vascularized hCOs (vhCOs) acquired several blood-brain barrier characteristics, including an increase in the expression of tight junctions, nutrient transporters and trans-endothelial electrical resistance. Finally, ETV2-induced endothelium supported the formation of perfused blood vessels in vivo. These vhCOs form vasculature-like structures that resemble the vasculature in early prenatal brain, and they present a robust model to study brain disease in vitro.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Single-cell transcriptome data are available in the Gene Expression Omnibus under accession code GSE134049.
Code availability
The DCMRSoft code that support the findings of this study are available on request from the corresponding author.
References
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373 (2013).
Paşca, A. M. et al. Functional cortical neurons and astrocytes from human pluripotent stem cells in 3D culture. Nat. Methods 12, 671 (2015).
Qian, X. et al. Brain-region-specific organoids using mini-bioreactors for modeling ZIKV exposure. Cell 165, 1238–1254 (2016).
Pașca, S. P. The rise of three-dimensional human brain cultures. Nature 553, 437 (2018).
Quadrato, G., Brown, J. & Arlotta, P. The promises and challenges of human brain organoids as models of neuropsychiatric disease. Nat. Med. 22, 1220 (2016).
Heide, M., Huttner, W. B. & Mora-Bermudez, F. Brain organoids as models to study human neocortex development and evolution. Curr. Opin. Cell Biol. 55, 8–16 (2018).
Lancaster, M. A. & Knoblich, J. A. Generation of cerebral organoids from human pluripotent stem cells. Nat. Protoc. 9, 2329 (2014).
Yin, X. et al. Engineering stem cell organoids. Cell Stem Cell 18, 25–38 (2016).
Shen, Q. et al. Endothelial cells stimulate self-renewal and expand neurogenesis of neural stem cells. Science 304, 1338–1340 (2004).
Mansour, A. A. et al. An in vivo model of functional and vascularized human brain organoids. Nat. Biotechnol. 36, 432 (2018).
Lee, S. et al. Direct reprogramming of human dermal fibroblasts into endothelial cells using ER71/ETV2. Circulation Res., Circresaha. 116, 309833 (2016).
Patsch, C. et al. Generation of vascular endothelial and smooth muscle cells from human pluripotent stem cells. Nat. Cell Biol. 17, 994–1003 (2015).
Lee, S., Kim, J. E., Johnson, B. A., Andukuri, A. & Yoon, Y.-S. Direct reprogramming into endothelial cells: a new source for vascular regeneration. Regen. Med. 12, 317–320 (2017).
Morita, R. et al. ETS transcription factor ETV2 directly converts human fibroblasts into functional endothelial cells. Proc. Natl Acad. Sci. USA 112, 160–165 (2015).
Engelhardt, B. & Liebner, S. Novel insights into the development and maintenance of the blood–brain barrier. Cell Tissue Res. 355, 687–699 (2014).
Hogan, K. A., Ambler, C. A., Chapman, D. L. & Bautch, V. L. The neural tube patterns vessels developmentally using the VEGF signaling pathway. Development 131, 1503–1513 (2004).
Norman, M. G. & O’kusky, J. R. The growth and development of microvasculature in human cerebral cortex. J. Neuropathol. Exp. Neurol. 45, 222–232 (1986).
Paredes, I., Himmels, P. & de Almodóvar, C. R. Neurovascular communication during CNS development. Developmental Cell 45, 10–32 (2018).
Jin, K. et al. Vascular endothelial growth factor (VEGF) stimulates neurogenesis in vitro and in vivo. Proc. Natl Acad. Sci. USA 99, 11946–11950 (2002).
Quadrato, G. et al. Cell diversity and network dynamics in photosensitive human brain organoids. Nature 545, 48 (2017).
Xiang, Y. et al. Fusion of regionally specified hPSC-derived organoids models human brain development and interneuron migration. Cell Stem Cell 21, 383–398 (2017). e387.
Xiang, Y. et al. hESC-derived thalamic organoids form reciprocal projections when fused with cortical organoids. Cell Stem Cell 24, 487–497 (2019). e487.
Darmanis, S. et al. A survey of human brain transcriptome diversity at the single cell level. Proc. Natl Acad. Sci. USA 112, 7285–7290 (2015).
Xu, B. et al. The endothelial cell-specific antibody PAL-E identifies a secreted form of vimentin in the blood vasculature. Mol. Cell Biol. 24, 9198–9206 (2004).
Morikawa, Y. & Cserjesi, P. Extra-embryonic vasculature development is regulated by the transcription factor HAND1. Development 131, 2195–2204 (2004).
Zhong, S. et al. A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex. Nature 555, 524–528 (2018).
Obermeier, B., Daneman, R. & Ransohoff, R. M. Development, maintenance and disruption of the blood-brain barrier. Nat. Med. 19, 1584 (2013).
Daneman, R., Zhou, L., Kebede, A. A. & Barres, B. A. Pericytes are required for blood–brain barrier integrity during embryogenesis. Nature 468, 562 (2010).
Winkler, E. A., Bell, R. D. & Zlokovic, B. V. Central nervous system pericytes in health and disease. Nat. Neurosci. 14, 1398 (2011).
Lippmann, E. S. et al. Derivation of blood-brain barrier endothelial cells from human pluripotent stem cells. Nat. Biotechnol. 30, 783 (2012).
Cho, C.-F. et al. Blood-brain-barrier spheroids as an in vitro screening platform for brain-penetrating agents. Nat. Commun. 8, 15623 (2017).
Wan, W. et al. Aβ1–42 oligomer‐induced leakage in an in vitro blood–brain barrier model is associated with up‐regulation of RAGE and metalloproteinases, and down‐regulation of tight junction scaffold proteins. J. Neurochem. 134, 382–393 (2015).
Almutairi, M. M., Gong, C., Xu, Y. G., Chang, Y. & Shi, H. Factors controlling permeability of the blood–brain barrier. Cell. Mol. Life Sci. 73, 57–77 (2016).
Deane, R. et al. RAGE mediates amyloid-β peptide transport across the blood-brain barrier and accumulation in brain. Nat. Med. 9, 907 (2003).
Cakir, B., Xiang, Y. & Park, I. H. Generation of vascularized human brain organoids Prot. Exch. https://doi.org/10.21203/rs.2.13464/v1 (2019).
Mandegar, M. A. et al. CRISPR interference efficiently induces specific and reversible gene silencing in human iPSCs. Cell Stem Cell 18, 541–553 (2016).
Yuan, Y., Altalhi, W. A., Ng, J. J. & Courtman, D. W. Derivation of human peripheral blood derived endothelial progenitor cells and the role of osteopontin surface modification and eNOS transfection. Biomaterials 34, 7292–7301 (2013).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Masselink, W. et al. Broad applicability of a streamlined ethyl cinnamate-based clearing procedure. Development 146, dev.166884 (2019).
Acknowledgements
I.-H.P. was partly supported by the NIH (grant nos. GM111667-01, R01AA025080-01, R01CA203011-2), CSCRF (grant nos. 14-SCC-YALE-01 and 16-RMB-YALE-04), Kavli Foundation, Simons Foundation and the KRIBB/KRCF research initiative program (grant no. NAP-09-3). This work was supported by the College of Medicine, University of Arkansas for Medical Sciences to Sang-Hun Lee, the Core Facilities of the Center for Translational Neuroscience, an award (no. P30 GM110702) from the IDeA program at NIGMS. Y.-S.Y was partly supported by the NIH (grant nos. R01HL127759 and DP3DK108245) and the Korea Health Technology R&D Project through the Korea Health Industry Development Institute, Republic of Korea (grant nos. HI15C2782 and HI16C2211). L.E.N was partly supported by NIH (1R21 EB024889, 1R01 HL148819), M.P. was partly supported by Fonds de recherche du Québec – Santé in Canada, and F.H. was partly supported by NIH (EB023366-02, MH067528-02). Computation time was provided by Yale University Biomedical High Performance Computing Center.
Author information
Authors and Affiliations
Contributions
B.C. and I.-H.P. conceived the study. B.C., Y.X., B.P., Y.-J.K., Y.T., K.-Y.K., P.S., Sang-Ho Lee and Y.-S.Y. performed the experiments. M.H.K., M.S.B.R. and L.E.N. performed perfusion setup and analysis. Y.Y. performed ECIS analysis. M.P. and F.H. performed MRI analysis. C.-S.H. performed surgeries. K.C., J.D. and P.P. performed TEER analysis. Y.-J.K. and S.-H.L. performed electrophysiological recordings. B.C., Y.X. and I.-H.P. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
I.-H.P. serves on the scientific advisory boards of ELGEN Therapeutics and INSTEMCARE with financial interest. However, the work presented in this manuscript is not related to ELGEN Therapeutics or INSTEMCARE.
Additional information
Peer review information Nina Vogt was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Characterization of ETV2-induced cells in hCOs.
(a) Left, immunostaining for SOX2 and TBR1 in control hCOs and vhCOs (30-day old). Right, immunostaining for CTIP2 and SATB2 in 120-day old control and vhCOs. Data are representative images of 5 organoids from three independent experiments. (b) Expression of pluripotency and neural genes from control hCOs and vhCOs (30- and 70-day old). Gene expression was measured relative to HES3 hESCs and normalized to β-Actin. Data represent the mean ± SEM (n=5, from three independent batches). (c) Quantification of vascularization in vhCOs and control hCOs at different time points. CD31 labeled row shows the projections of z-stacks of CD31 stained red whole mount images. Angiotool analysis of the CD31 projections, thick red lines show the vascular paths, blue dots represent vascular junction points and the thin white line demonstrate the entire area over which vascular structures formed within organoid. Data represent the mean ± SEM (n=3, from three independent batches). (d-e) Co-immunostaining for CD31 and mCherry (d), CDH5 and mCherry (e) in 70-day old hCOs. Data are representative images of three independent experiments. (f) Representative electron microscopy images of vhCO at day 30 and 180. The E represents the endothelial cells, black arrowhead points at tight junction and white arrowhead indicates the astrocytic end feet. (n=2, from two independent batches). The scale bar represents 50 μm in a, c and d, and 100 μm in e, and 2 and 1 μm at day 30 and 180, respectively in f.
Supplementary Figure 2 Characterization of ETV2-induction in hCOs.
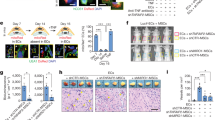
(a) Immunostaining for endothelial markers, CD31 and CDH5, and tight junction marker, α-ZO1, in 30-day old H1-derived hCOs. Data are representative images of 5 organoids from two independent experiments. (b) Image of the FITC-dextran perfusion system for organoids via connected tubes. (c) Left, Image of FITC-dextran and CDH5 in perfused hCOs with confocal microscopy. Right, the percentage of lumens containing FITC-dextran in vhCOs and control hCOs. Data represent the mean ± SEM (n=5, from three independent batches); (T=9.761 DF=4 and *p=0.0001). (d) Left, HIF-1α staining of organoids after 30-,70- and 120-day culture. Right, quantification of HIF-1α +/DAPI+ cells in control hCOs and vhCOs at days 30, 70 and 120. Data represent the mean ± SEM (n=3, from three independent batches). (D70: T=7.91 DF=4 and *p=0.00138, D120: T=16.07 DF=4 and **p=0.000087). (e) Left, expression of the synaptic marker SYN1 in control hCOs and vhCOs. Right, quantification of SYNP +/DAPI+ cells in control hCOs and vhCOs at days 30, 70. Data represent the mean ± SEM (n=3, from three independent batches). (D70: T=11.18 DF=4 and *p=0.00036, D120: T=7.372 DF=4 and **p=0.0018). The unpaired two-tail t-test was used for all comparisons. The scale bar represents 100 μm in d, e, and 50 μm in a, and c.
Supplementary Figure 3 Classification, annotation and evaluation of cell clusters from vhCOs.
(a) tSNE plot of single cells colored by functional clusters. Data are representative of 20026 cells. (b) Schematic representation of cluster annotation. (c) Expression patterns of genes related with neuronal growth cone, early neurogenesis, proteoglycans and EMT. Data are representative of 20026 cells. (d) Comparison of total UMI counts, mitochondria-derived reads, GO enrichment and unique markers for lineage commitment and cellular event among clusters. (e) Exclusive expression pattern of TBR1 and GFAP genes in single cells. Cells co-expressing both genes (colored by blue) were considered doublets. (f) Histogram of gene expression related to vasculogenesis. Two-sided t-test p-value=1.91e-02 (VTN) and 2.2e-16 (HAND1). Data are representative of 2257 cells.
Supplementary Figure 4 Functional characterization of vhCOs.
(a) Immunostaining for α-ZO1 in hCOs and vhCOs at day 70. (b-d) Immunostaining for β-Catenin (b), OCLN and KDR (c), and GFAP and S100β (d) in 30-day old hCOs and vhCOs. Arrows show co-localization of OCLN and KDR in vhCOs. Data are representative images of 5 organoids from three independent experiments in a, b, c, and d. (e) Co-immunostaining for GFAP and PDGFRβ in control and vhCOs (day 70). Arrows indicate regions where pericytes and astrocytes are in close juxtaposition. (n=2, from two independent batches) (f) Expression of tight junctions and transporters with and without Aβ1-42-oligo treatment of vhCOs. Gene expressions were measured relative to control hCOs and normalized to β-Actin. Data represent the mean ± SEM (n=3, from three independent batches). The scale bar represents 50 μm in a, b, c, d and e.
Supplementary information
Supplementary Information
Supplementary Figs. 1–4, Notes 1–3 and Protocol.
Supplementary Video 1
Vascular networks in vhCOs at day 30.
Supplementary Video 2
Vascular networks in hCOs at day 30.
Supplementary Video 3
FITC-dextran perfusion of a vhCO at day 30.
Supplementary Video 4
FITC-dextran perfusion of vhCO explant stained for human-specific CD31 and hNUC.
Supplementary Video 5
FITC-dextran perfusion of vhCO explant.
Supplementary Video 6
FITC-dextran perfusion of control hCO explant.
Supplementary Video 7
Mouse brain explant with FITC-dextran perfusion.
Rights and permissions
About this article
Cite this article
Cakir, B., Xiang, Y., Tanaka, Y. et al. Engineering of human brain organoids with a functional vascular-like system. Nat Methods 16, 1169–1175 (2019). https://doi.org/10.1038/s41592-019-0586-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0586-5
This article is cited by
-
Application status and optimization suggestions of tumor organoids and CAR-T cell co-culture models
Cancer Cell International (2024)
-
Chronic hypoxia remodels the tumor microenvironment to support glioma stem cell growth
Acta Neuropathologica Communications (2024)
-
The molecular determinants of microglial developmental dynamics
Nature Reviews Neuroscience (2024)
-
Profiling human brain vascular cells using single-cell transcriptomics and organoids
Nature Protocols (2024)
-
A single-nuclei paired multiomic analysis of the human midbrain reveals age- and Parkinson’s disease–associated glial changes
Nature Aging (2024)