Abstract
We introduce a liquid application method for time-resolved analyses (LAMA), an in situ mixing approach for serial crystallography. Picoliter-sized droplets are shot onto chip-mounted protein crystals, achieving near-full ligand occupancy within theoretical diffusion times. We demonstrate proof-of-principle binding of GlcNac to lysozyme, and resolve glucose binding and subsequent ring opening in a time-resolved study of xylose isomerase.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Moffat, K. Annu Rev. Biophys. Chem. 18, 309–332 (1989).
Levantino, M., Yorke, B. A., Monteiro, D. C., Cammarata, M. & Pearson, A. R. Curr. Opin. Struct. Biol. 35, 41–48 (2015).
Gati, C. et al. IUCrJ 1, 87–94 (2014).
Yamamoto, M. et al. IUCrJ 4, 529–539 (2017).
Oghbaey, S. et al. Acta Crystallogr. Sect. D. 72, 944–955 (2016).
Owen, R. L. et al. Acta Crystallogr. Sect. D. 73, 373–378 (2017).
Schulz, E. C. et al. Nat. Methods 15, 901–904 (2018).
Coquelle, N. et al. Acta Crystallogr. Sect. D. 71, 1184–1196 (2015).
Bar-Even, A. et al. Biochemistry 50, 4402–4410 (2011).
Chapman, H. N. Annu. Rev. Biochem. 88, 35–58 (2019).
Stagno, J. R. et al. Nature 541, 242–246 (2017).
Beyerlein, K. R. et al. IUCrJ 4, 769–777 (2017).
Schmidt, M. Adv. Condens. Matter Phys. 2013, 167276 (2013).
Makinen, M. W. & Fink, A. L. Annu. Rev. Biophys. Bioeng. 6, 301–343 (1977).
Olmos, J. L. et al. BMC Biol. 16, 1–15 (2018).
Russi, S. et al. J. Synchrotron Radiat. 24, 73–82 (2017).
Fischer, M., Shoichet, B. K. & Fraser, J. S. ChemBioChem 16, 1560–1564 (2015).
Keedy, D. A. et al. Structure 22, 899–910 (2014).
Hochster, R. M. & Watson, R. W. J. Am. Chem. Soc. 75, 3284–3285 (1953).
Hogue Angeletti, R. A. J. Biol. Chem. 250, 7814–7818 (1975).
Chauthaiwale, J. & Rao, M. Appl. Environ. Microbiol. 60, 4495–4499 (1994).
Cha, J. & Batt, C. A. Mol. Cells 8, 374–382 (1998).
Toteva, M. M., Silvaggi, N. R., Allen, K. N. & Richard, J. P. Biochemistry 50, 10170–10181 (2011).
Chanitnun, K. & Pinphanichakarn, P. Braz. J. Microbiol. 43, 1084–1093 (2012).
Dauter, Z., Dauter, M., Hemker, J., Witzel, H. & Wilson, K. S. FEBS Lett. 247, 1–8 (1989).
Bhosale, S. H., Rao, M. B. & Deshpande, V. V. Microbiol. Rev. 60, 280–300 (1996).
Lavie, A., Allen, K. N., Petsko, G. A. & Ringe, D. Biochemistry 33, 5469–5480 (1994).
Kovalevsky, A. Y. et al. Structure 18, 688–699 (2010).
Martin, R. W. & Zilm, K. W. J. Magn. Reson. 165, 162–174 (2003).
Kabsch, W. Acta Crystallogr. Sect. D 70, 2204–2216 (2014).
White, T. A. et al. J. Appl. Crystallogr. 45, 335–341 (2012).
McCoy, A. J. et al. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P. D. et al. J. Synchrotron Radiat. 11, 53–55 (2004).
Emsley, P. & Cowtan, K. Acta Crystallogr. Sect. D 60, 2126–2132 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Acta Crystallogr. Sect. D 66, 486–501 (2010).
Liebschner, D. et al. Acta Crystallogr. Sect. D 73, 148–157 (2017).
Lang, P. T., Holton, J. M., Fraser, J. S. & Alber, T. Proc. Natl Acad. Sci. USA 111, 237–242 (2014).
The PyMOL Molecular Graphics System, v.1.8. (Schrödinger LLC, 2015).
Acknowledgements
The authors thank D. Oberthuer (CFEL, DESY Hamburg) for the provision of the GlcNAc3 substrate immediately before data collection. The LAMA TR-SSX data were collected at beamline P14 and P14-2 (T-REXX) operated by EMBL Hamburg at the PETRA III storage ring (DESY, Hamburg, Germany). Construction of T-REXX was funded by the BMBF (Verbund Forschungsprojekt 05K16GU1). The authors are grateful to R. Dousza, L. Bultema and J. Besaw for their technical assistance during data collection. T-REXX beamtime was awarded as part of the EMBL BAG MX-660. The authors gratefully acknowledge the support provided by the Max Planck Society and the excellence cluster ‘The Hamburg Centre for Ultrafast Imaging—Structure, Dynamics and Control of Matter at the Atomic Scale’ of the Deutsche Forschungsgemeinschaft EXC 1074 project ID 194651731 (R.J.D.M.). P.M. was supported by the Alexander von Humboldt-Stiftung for postdoctoral researchers.
Author information
Authors and Affiliations
Contributions
P.M. and E.C.S. designed the experiment. E.C. S. and P. M. performed the experiments with support from F.T., H.S, J.P.L., S.H, D.v.S., G.B. and M.A. P.M. prepared the protein crystals. F.T. and P.M designed the integration of the droplet injector with support from E.C.S. J.P.L. designed and built the humidity nozzle. E.C.S. and P.M. processed and analyzed the diffraction data. S.H., M.A., G.B., D.v.S., A.R.P. and T.R.S. designed and built the T-REXX endstation to enable these experiments. R.J.D.M, P.M. and E.C.S. wrote the manuscript. All authors discussed and corrected the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Allison Doerr was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
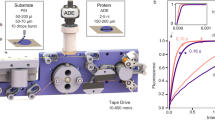
Supplementary Figure 1 The LAMA setup.
a) Overview CAD render of the LAMA setup. The humidifier is connected to the translation stage with a hose represented in orange. The diffractometer, chip holder and translation stages were previously described in detail1. b) Closeup view of a) displays the injector nozzle (beige) mounted at a fixed point above the chip (black). The landing point of the droplets intersects with the X-ray beam at the position of each individual protein microcrystal inside the chip features as the chip is rastered across the X-ray beam. The humidified air nozzle is 3D-printed to provide a homogenous humidity stream over chip, preventing crystal dehydration. c) A ~150 pl droplet generated by a single pulse waveform of the piezo transducer (microdrop Technologies). d) A ~75 pl droplet generated by a triple pulse waveform of the piezo transducer (microdrop Technologies).
Supplementary Figure 2 Absolute value electron density maps for the different Lysozyme and Xylose isomerase time-points.
(a–c) GlcNac3 is shown in a ball-and-stick representation. Absolute-value maps contoured at 0.5 e− Å−3 from 0.05 s—1 s display a gradual increase of the electron density as a function of time. (d–i) Coloring and representation as in Fig. 2. All absolute value electron density maps are shown with the density contoured at 0.7 e− Å−3.
Supplementary Figure 3 Detector/Stages/Droplet generator pulse sequence.
The plot shows the on/off times of the various components of the LAMA setup. Initially the stage moves into position and there is a pre-exposure time of 1 ms during which the detector is triggered, and a t0 image is taken. Subsequently a TTL pulse is sent to the droplet generator; the droplet has a time-of-flight of ~0.5 ms which is encompassed in the post-exposure time (4 ms). After droplet injection additional images are recorded at different feature positions. Finally, the sequence is repeated without any further droplet injection. The total time delay corresponds to a HARE number1.
Supplementary Figure 4 XI structure comparison.
a) The XI active site of the 0 s structure is clearly empty and shows only strong density for the metal binding sites. Metals were modelled as cobalt (green sphere) and magnesium (pink sphere) since these matched the 2Fo-Fc electron density best. The POLDER-OMIT map for the metals is shown at 3σ. b) Superimposition of XI with a previously determined neutron structure of XI in complex with glucose in closed ring conformation (cyan, 3KCL). c) Superimposition of XI with a previously determined neutron structure of XI in complex with glucose in open ring conformation (cyan, 3KBN).
Supplementary Figure 5 Superposition of XI structures over time.
a) Cartoon representation of the 7 XI-monomer structures superimposed with an overall r.m.s.d of 0.09 Å2. indicating that differences between all the XI structures are within error. b) The active site of all the superimposed XI structures showing that the side chains as well as the structural waters within a 7 Å radius around the ligand also show near identical conformations from one time point to the next.
Supplementary information
Supplementary Information
Supplementary Figures 1–5, Supplementary Note, Supplementary Tables 1–3
Rights and permissions
About this article
Cite this article
Mehrabi, P., Schulz, E.C., Agthe, M. et al. Liquid application method for time-resolved analyses by serial synchrotron crystallography. Nat Methods 16, 979–982 (2019). https://doi.org/10.1038/s41592-019-0553-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0553-1
This article is cited by
-
Millisecond cryo-trapping by the spitrobot crystal plunger simplifies time-resolved crystallography
Nature Communications (2023)
-
Using macromolecular electron densities to improve the enrichment of active compounds in virtual screening
Communications Chemistry (2023)
-
Heterogeneity in M. tuberculosis β-lactamase inhibition by Sulbactam
Nature Communications (2023)
-
Serial femtosecond crystallography
Nature Reviews Methods Primers (2022)
-
Beef tallow injection matrix for serial crystallography
Scientific Reports (2022)