Abstract
Single-cell RNA sequencing (scRNA-seq) data are noisy and sparse. Here, we show that transfer learning across datasets remarkably improves data quality. By coupling a deep autoencoder with a Bayesian model, SAVER-X extracts transferable gene−gene relationships across data from different labs, varying conditions and divergent species, to denoise new target datasets.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The HCA dataset was downloaded from the HCA data portal (https://preview.data.humancellatlas.org/) and the PBMC data14 were downloaded from the 10X website (https://support.10xgenomics.com/single-cell-gene-expression/datasets, Supplementary Table 2). The breast cancer data16 were downloaded from the Gene Expression Omnibus (GEO) (GSE114725). The developing midbrain data12 were downloaded from GEO (GSE76381). For the other mouse developing brain datasets in Fig. 3, we included cells from neonatal and fetal brain tissues in the Mouse Cell Atlas7 data (GSE108097). For the non-UMI human developing brain datasets in Supplementary Fig. 8, we included GSE75140 (ref. 18) GSE104276 (ref. 19) and SRP041736 (ref. 17). No gene or cell filtering was done on the original dataset.
A complete list of the pretraining datasets used for pretraining the models on the SAVER-X website is provided in Supplementary Table 2.
Code availability
SAVER-X is publicly available at http://singlecell.wharton.upenn.edu/saver-x/, where users can currently upload their data for cloud computing and choose from models pretrained on 31 mouse tissues and human immune cells. Models jointly pretrained on cells from both species are also available for brain and pancreatic tissues. The R package and source code of SAVER-X was also released at https://github.com/jingshuw/SAVERX.
References
Huang, M. et al. Nat. Methods 15, 539–542 (2018).
Li, W. V. & Li, J. J. Nat. Commun. 9, 1–9 (2018).
van Dijk, D. et al. Cell 174, 716–729 (2018).
Eraslan, G., Simon, L. M., Mircea, M., Mueller, N. S. & Theis, F. J. Nat. Commun. 10, 390 (2019).
Gong, W., Kwak, I., Pota, P., Koyano-nakagawa, N. & Garry, D. J. BMC Bioinforma. 19, 1–10 (2018).
Lopez, R., Regier, J., Cole, M. B., Jordan, M. I. & Yosef, N. Nat. Methods 15, 1053–1058 (2018).
Han, X. et al. Cell 172, 1091–1097 (2018).
Tabula, T. & Consortium, M. Nature 562, 367–372 (2018).
Regev, A. et al. eLife 6, 1–30 (2017).
Hinton, G. E. & Salakhutdinov, R. R. Science 313, 504–507 (2006).
Andrews, T. S., Hemberg, M. & Hicks, S. F1000Research 7, 1740 (2018).
La Manno, G. et al. Cell 167, 566–580 (2016).
Nguyen, A. et al. Front. Immunol. 9, 1553 (2018).
Zheng, G. X. Y. et al. Nat. Commun. 8, 1–12 (2017).
Stoeckius, M. et al. Nat. Methods 14, 865–868 (2017).
Azizi, E. et al. Cell 174, 1293–1308 (2018).
Pollen, A. A. et al. Nat. Biotechnol. 32, 1053–1058 (2014).
Camp, J. G. et al. Proc. Natl Acad. Sci. USA 112, 15672–15677 (2015).
Zhong, S. et al. Nature 555, 524–528 (2018).
Wang, J. et al. Proc. Natl Acad. Sci. USA 115, E6437–E6446 (2018).
Kim, J. K. et al. Nat. Commun. 6, 8687 (2015).
Acknowledgements
We thank H. MacMullan IV, V. Conley and S. Zamechek from Wharton Computing’s Research & Analytics Team (https://research-it.wharton.upenn.edu/) for their valuable assistance in implementing the code in a scalable fashion and integrating the scalable code solution into a backend service for our website. We also thank the National Institute of Health for the award 5R01-HG006137 (for D.A., Z.Z., and N.Z.), the National Science Foundation for the award DMS-1562665 (to J.W., N.Z.), the Blavatnik Family Foundation’s Graduate Student Fellowship awarded to D.A., NSF Graduate Fellowship DGE-1321851 awarded to M.H., and the Natural Science Foundation of Tianjin for grant 18JCYBJC24900 to G.H. This work used the Extreme Science and Engineering Discovery Environment (XSEDE), which is supported by National Science Foundation grant number ACI-1548562. Specifically, it used the Bridges system, which is supported by NSF award number ACI-1445606, at the Pittsburgh Supercomputing Center (PSC).
Author information
Authors and Affiliations
Contributions
J.W. and N.Z. conceptualized the study, designed the model and planned the case studies. J.W. developed the algorithm, implemented the SAVER-X software and led the data analysis. D.A. constructed the SAVER-X website and helped with data analysis. M.H. performed benchmarking with other methods and helped with algorithm development. G.H. helped with algorithm development and model design. Z.Z. tested SAVER-X website and software and helped with data analysis. C.Y. conducted the analysis of CITE-seq data. J.W., D.A. and N.Z. wrote the paper with feedback from M.H. and Z.Z.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Nicole Rusk was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Gene-gene correlation recovery.
Gene-gene Pearson correlations of all gene pairs are calculated in the denoised matrix and compared with the gene-gene correlations of the reference normalized count matrix. These are density plots of the difference between the denoised matrix and the reference across all gene-gene pairs for four different datasets. We compare five different methods: SAVER-X, DCA, scVI and the latter two with empirical Bayes shrinkage as in SAVER-X (DCA + EB, scVI + EB). These plots show that our final empirical Bayes shrinkage step in SAVER-X is essential to reduce bias in the denoised data regardless of the denoising method. See supplementary note 5 for more details.
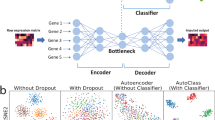
Supplementary Figure 2 Architecture of the autoencoder.
The autoencoder allows cells from both human and mouse, both with and without UMI, to be used for pre-training. For each species, there are ~20,000 input nodes accepting raw gene expression values; two-thirds of these are shared between mouse and humans by accounting for genes with homologs. See supplementary note 1 for more details.
Supplementary Figure 3 Comparison of denoising methods under four scenarios with varying numbers of cells and the sequencing depth.
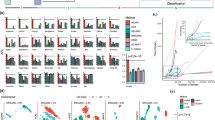
The performance of SAVER-X is benchmarked against existing denoising methods using t-SNE visualization and clustering adjusted rand index (ARI). The number at the right-bottom corner of each plot is the ARI, which is summarized also in the last row.
Supplementary Figure 4 More scenarios on PBMC T cell transfer learning.
The other two scenarios of the in-silico experiments on the 600 PBMC T cell transfer learning that is not shown in Fig. 2c. More details see supplementary note 4 for more details.
Supplementary Figure 5 Benchmarking scRNA-seq data denoising methods using the CITE-seq data (Stoeckius, M. et al, 2017).
Pearson correlation were calculated between proteins and corresponding mRNA levels in the CITE-seq PBMC (a) and CBMC (b) datasets. The mRNA measurements were denoised using SAVER-X and other benchmarking denoising methods (X-axis). The effect of transfer learning shows significantly when the information available from the target data is limited.
Supplementary Figure 6 Transfer learning from normal to breast cancer immune cells.
a) Infiltrating immune cells in resected breast carcinoma from two breast cancer patient tumors from Azizi et al (2018). The three panels show visualizations using original data, denoised values by SAVER-X without pretraining, and denoised values by SAVER-X pretrained on immune cells (HCA and 10X PBMC data). The t-SNE plots show separation between cell types (cell labels are obtained from the original paper). Feature plots show the expression of some known marker genes, and a darker red color represents a relatively higher expression level in some cells compared with the rest of cells. b) t-SNE plots for tumors of the other 6 breast cancer patients.
Supplementary Figure 7 Marker genes of the tumor-specific cell group in tumor BC8 of Azizi et al (2018).
a) tSNE plot and feature plots of the BC8 tumor myeloid cells (1046 cells) using the original raw data. The highly enriched expressions of immunoglobulin genes confirm that this is a tumor-related cell population. b) The t-SNE plot and feature plots of the BC8 tumor myeloid cells using the SAVER-X denoised data, after pretraining with the normal immune cells. A darker red color represents a relatively higher expression level in some cells compared with the rest of cells.
Supplementary Figure 8 SAVER-X analyses of the La Manno et. al. (2016) data.
a) Illustration of the complete design and data use. 1977 human cells are randomly split into two groups. The 1000 cells in group 1 are further down sampled. We consider pretraining with 1907 mouse cells in the same paper, 977 original human cells, 7187 mouse cells from MCA and a total of 3344 non-UMI human developmental brain human cells. b) t-SNE plots of the 1000 down-sampled cells for other denoising models not shown in Fig. 3b. Cell labels are the computed labels from the original paper. The numbers at the right corner are the ARI for each plot. Cell types are colored the same as in Fig. 3b. c) Log fold changes between human and mouse data of cell-type-specific differentially expressed genes. X axis uses the original human data and Y axis uses the denoised down-sampled human data which is denoised using SAVER-X pretrained with the paired mouse cells from La Manno et. al. (2016), but without gene filtering or empirical Bayes shrinkage. Each dot is a differentially expressed gene between human and mouse in that cell type. Genes that have notable bias are highlighted with blue circles. d) the heatmaps of the denoised gene expressions for a set of known marker genes for 5 human brain cell types.
Supplementary Figure 9 Data alignment pre- and post-denoising yields consistent clustering results.
Panels (a) and (b) show the results of data alignment using the raw and denoised results, respectively, when we align the CBMC and PBMC scRNAs-seq datasets of ~8000 cells each. The panels on the left show the results of data alignment using 20 canonical components in SeuratCCA v2, whereas the panels on the right demonstrate the cell type identities of the clusters identified after alignment. See supplementary note 6 for more details.
Supplementary information
Supplementary Information
Supplementary Figures 1–9, Supplementary Tables 1 and 2, Supplementary Notes 1–6.
Rights and permissions
About this article
Cite this article
Wang, J., Agarwal, D., Huang, M. et al. Data denoising with transfer learning in single-cell transcriptomics. Nat Methods 16, 875–878 (2019). https://doi.org/10.1038/s41592-019-0537-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0537-1
This article is cited by
-
scNAT: a deep learning method for integrating paired single-cell RNA and T cell receptor sequencing profiles
Genome Biology (2023)
-
scBridge embraces cell heterogeneity in single-cell RNA-seq and ATAC-seq data integration
Nature Communications (2023)
-
SPACEL: deep learning-based characterization of spatial transcriptome architectures
Nature Communications (2023)
-
Two-Stage Training of Graph Neural Networks for Graph Classification
Neural Processing Letters (2023)
-
WaveCNNs-AT: Wavelet-based deep CNNs of adaptive threshold for signal recognition
Applied Intelligence (2023)