Abstract
The open-source de novo protein-level assembler, Plass (https://plass.mmseqs.com), assembles six-frame-translated sequencing reads into protein sequences. It recovers 2–10 times more protein sequences from complex metagenomes and can assemble huge datasets. We assembled two redundancy-filtered reference protein catalogs, 2 billion sequences from 640 soil samples (soil reference protein catalog) and 292 million sequences from 775 marine eukaryotic metatranscriptomes (marine eukaryotic reference catalog), the largest free collections of protein sequences.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The assembled protein sequence sets are available in FASTA format under a Creative Commons Attribution CC-BY 4.0 License at https://plass.mmseqs.com. All scripts and benchmark data including command-line parameters necessary to reproduce the benchmark and analysis results presented are available at https://github.com/martin-steinegger/plass-analysis.
Code availability
Plass is GPLv3-licensed open-source software. The source code and binaries for Plass can be downloaded at https://github.com/soedinglab/plass.
References
Howe, A. C. et al. Proc. Natl. Acad. Sci. USA 111, 4904–4909 (2014).
Li, J. et al. Nat. Biotechnol. 32, 834–841 (2014).
Sunagawa, S. et al. Science 348, 1261359 (2015).
Nielsen, H. B. et al. Nat. Biotechnol. 32, 822–828 (2014).
Zerbino, D. & Birney, E. Genome Res. 18, 821–829 (2008).
Li, D. et al. Bioinformatics 31, 1674–1676 (2015).
Nurk, S. et al. Genome Res. 27, 824–834 (2017).
Ye, Y. & Tang, H. J. Bioinform. Comput. Biol. 7, 455–471 (2009).
Yang, Y. et al. Bioinformatics 31, 1833–1835 (2015).
Steinegger, M. & Söding, J. Nat. Commun. 9, 2542 (2018).
Sczyrba, A. et al. Nat. Methods 14, 1063–1071 (2017).
Kashtan, N. et al. Science 344, 416–420 (2014).
Berube, P. M. et al. Sci. Data 5, 180154 (2018).
Hyatt, D. et al. BMC Bioinforma. 11, 119 (2010).
van der Walt, A. J. et al. BMC Genom. 18, 521 (2017).
Huerta-Cepas, J. et al. Mol. Biol. Evol. 34, 2115–2122 (2017).
Tian, W. & Skolnick, J. J. Mol. Biol. 333, 863–882 (2003).
Lee, S. T. M. et al. Microbiome 5, 50 (2017).
Carradec, Q. et al. Nat. Commun. 9, 373 (2018).
Ovchinnikov, S. et al. Science 355, 294–298 (2017).
Magoc, T. & Salzberg, S. L. Bioinformatics 27, 2957–2963 (2011).
Sheetlin, S. et al. Bioinformatics 32, 304–305 (2016).
Mirdita, M. et al. Nucleic Acids Res. 45, D170–D176 (2017).
Kanehisa, M. et al. Nucleic Acids Res. 45, D353–D361 (2016).
Steinegger, M. & Söding, J. Nat. Biotechnol. 35, 1026–1028 (2017).
Huerta-Cepas, J. et al. Nucleic Acids Res. 44, D286–D293 (2016).
Frith, M. Nucleic Acids Res. 39, E23 (2011).
Hingamp, P. ISME J. 7, 1678–1695 (2013).
Acknowledgements
We are grateful to C. Notredame and C. Seok for hosting M.S. at the Centre for Genomic Regulation and Seoul National University for 12 and 30 months, respectively. We thank S. Sunagawa, F. Meyer and A. Sczyrba for helpful discussions, and T. Brown for his early analysis and detailed feedback on Plass results. We thank all who contributed metagenomic datasets used to build SRC and MERC, in particular contributors to the TARA ocean project and the US Department of Energy Joint Genome Institute (http://www.jgi.doe.gov). This work was supported by the EU’s Horizon 2020 Framework Programme (Virus-X, grant no. 685778).
Author information
Authors and Affiliations
Contributions
M.S. and J.S. designed the research study. M.S. and M.M. developed code and performed the analyses. M.S. and J.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Lei Tang was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Schematic comparison of a nucleotide- and a protein assembly.
On top is the final protein assembly followed by the stacked overlapping protein reads. The small gray section highlights the multiple protein sequence alignment of the overlapping reads and below the respective nucleotide alignment. Less ambiguity is visible on the protein level due to conservative mutations (mutations with similar biochemical properties) compared to the nucleotide level, resulting in an assembly that is more robust to microdiversity in the population.
Supplementary Figure 2 Overlap of assemblies of Plass with Megahit and metaSPAdes.
(a) left: A fraction 38.3% of amino acids in the Plass-assembled proteins of set 1 is covered by alignments to proteins in the Megahit assembly at a minimum sequence identity cut-off of 99%. Conversely, 83.2% of proteins in the Megahit-assembled set 1 is covered by alignments with Plass-assembled proteins. Right: Same as on the left but comparing the Plass assembly with the metaSPAdes assembly. (b) Same as (a) but for protein set 2.
Supplementary Figure 3 Effect of neural network filter to remove wrong translation frames.
Sensitivity and precision in set 1 (a) and set 2 (b). Top: assembly sensitivity is the fraction of reference sequence amino acids that matches to an assembled protein sequence. Bottom: assembly precision is the fraction of assembled amino acids that matches to a reference protein at the minimum sequence identity on the x-axis. Plass uses a minimum sequence identity for merging fragments of 90%, Plass-97 uses a threshold of 97%.
Supplementary Figure 4 Comparison of Megahit assignment using the 2bLCA protocol.
The MMseqs2 taxonomy assignment workflow uses three steps to assign a taxonomic label to a query sequence. (1) We search with the query sequence against a reference database and extract the aligned subsequence of the best hit. (2) This sequence is matched again against the reference database. Each hit with an E-value smaller than the best hit E-value from the previous search is accepted. (3) We compute the lowest common ancestor based on the taxonomic labels of all accepted hits.
Supplementary Figure 5 Plass ORF extraction and start codon prediction (ORF calling).
Plass extracts two sets of ORFs. ORF set 1 contains all translated ORFs with at least 45 codons. ORF set 2 contains all translated ORFs with at least 20 codons starting with a putative ATG start codon that is the first ATG codon after a stop codon in the same frame. (Start codon prediction) Plass predicts start codons with a consensus method using a multiple sequence alignment of ORF set 1 and 2. Wherever at least 20% of all methionines in one column are marked by a prepended asterisk, it removes the preceding residues from all other sequences and prepends an asterisk to all sequences to mark the start.
Supplementary Figure 6 Taxonomy evaluation of the soil metagenome assembly.
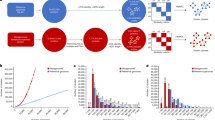
(a) We investigate the taxonomic composition of the 8 most abundant taxa (all other taxa are pooled in ‘Others’) in the soil assemblies from Fig. 2d (blue: Megahit, red: Plass) and the assemblies of the 12 soil samples from Fig. 2e (light blue: Megahit, light red: Plass). On top we show the read count ratios between Plass and Megahit, for both the single and 12 soil assemblies. The inset gives the fraction of reads in the single and the 12 soil samples that could be mapped to an assembled protein sequence. (b) We show the count of assembled amino acids within various coverage ranges for Megahit (blue) and Plass (red) in the single soil sample.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6
Source data
Rights and permissions
About this article
Cite this article
Steinegger, M., Mirdita, M. & Söding, J. Protein-level assembly increases protein sequence recovery from metagenomic samples manyfold. Nat Methods 16, 603–606 (2019). https://doi.org/10.1038/s41592-019-0437-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0437-4
This article is cited by
-
Protein embedding based alignment
BMC Bioinformatics (2024)
-
Improving deep learning protein monomer and complex structure prediction using DeepMSA2 with huge metagenomics data
Nature Methods (2024)
-
Streptomyces umbrella toxin particles block hyphal growth of competing species
Nature (2024)
-
Application of Computational Techniques in Antibody Fc-Fused Molecule Design for Therapeutics
Molecular Biotechnology (2024)
-
AttSec: protein secondary structure prediction by capturing local patterns from attention map
BMC Bioinformatics (2023)