Abstract
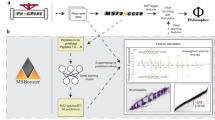
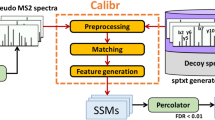
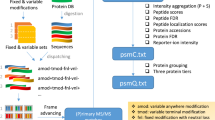
Peptide fragmentation spectra are routinely predicted in the interpretation of mass-spectrometry-based proteomics data. However, the generation of fragment ions has not been understood well enough for scientists to estimate fragment ion intensities accurately. Here, we demonstrate that machine learning can predict peptide fragmentation patterns in mass spectrometers with accuracy within the uncertainty of measurement. Moreover, analysis of our models reveals that peptide fragmentation depends on long-range interactions within a peptide sequence. We illustrate the utility of our models by applying them to the analysis of both data-dependent and data-independent acquisition datasets. In the former case, we observe a q-value-dependent increase in the total number of peptide identifications. In the latter case, we confirm that the use of predicted tandem mass spectrometry spectra is nearly equivalent to the use of spectra from experimental libraries.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The mass spectrometry proteomics data including summary tables have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD010382.
Code availability
We offer the trained DeepMass:Prism model for use via the Google Cloud ML platform (https://github.com/verilylifesciences/deepmass/tree/master/prism). To obtain the trained DeepMass:Prism model to run locally, please contact the corresponding authors. A user-friendly interface will be made available in the future MaxQuant releases.
References
Cottrell, J. S. Protein identification using MS/MS data. J. Proteom. 74, 1842–1851 (2011).
Sinitcyn, P., Rudolph, J. D. & Cox, J. Computational methods for understanding mass spectrometry-based shotgun proteomics data. Annu. Rev. Biomed. Data Sci. 1, 207–234 (2018).
Mitchell Wells, J. & McLuckey, S. A. Collision-induced dissociation (CID) of peptides and proteins. Methods Enzym. 402, 148–185 (2005).
Olsen, J. V. et al. Higher-energy C-trap dissociation for peptide modification analysis. Nat. Methods 4, 709–712 (2007).
Coon, J. J., Syka, J., Shabanowitz, J. & Hunt, D. F. Tandem mass spectrometry for peptide and protein sequence analysis. Biotechniques 38, 519–521 (2005).
Good, D. M., Wirtala, M., McAlister, G. C. & Coon, J. J. Performance characteristics of electron transfer dissociation mass spectrometry. Mol. Cell. Proteomics 6, 1942–1951 (2007).
Steen, H. & Mann, M. The ABC’s (and XYZ’s) of peptide sequencing. Nat. Rev. Mol. Cell Biol. 5, 699–711 (2004).
Boyd, R. & Somogyi, Á. The mobile proton hypothesis in fragmentation of protonated peptides: a perspective. J. Am. Soc. Mass Spectrom. 21, 1275–1278 (2010).
Arnold, R. J., Jayasankar, N., Aggarwal, D., Tang, H. & Radivojac, P. A machine learning approach to predicting peptide fragmentation spectra. Pac. Symp. Biocomput. 230, 219–230 (2006).
Degroeve, S., Martens, L. & Jurisica, I. MS2PIP: a tool for MS/MS peak intensity prediction. Bioinformatics 29, 3199–3203 (2013).
Dong, N. P. et al. Prediction of peptide fragment ion mass spectra by data mining techniques. Anal. Chem. 86, 7446–7454 (2014).
Park, J. et al. Informed-Proteomics: open-source software package for top-down proteomics. Nat. Methods 14, 909–914 (2017).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Wolters, D. A., Washburn, M. P. & Yates, J. R. An automated multidimensional protein identification technology for shotgun proteomics. Anal. Chem. 73, 5683–5690 (2001).
Doerr, A. DIA mass spectrometry. Nat. Methods 12, 35–35 (2014).
Graves, A. et al. A novel connectionist system for unconstrained handwriting recognition. IEEE Trans. Pattern Anal. Mach. Intell. 31, 855–868 (2009).
Garnier, J., Gibrat, J.-F. & Robson, B. GOR method for predicting protein secondary structure from amino acid sequence. Methods Enzymol. 266, 540–553 (1996).
Rost, B., Sander, C. & Schneider, R. PHD—an automatic mail server for protein secondary structure prediction. Bioinformatics 10, 53–60 (1994).
Vapnik, V. N. The Nature of Statistical Learning Theory (Springer, 1995).
Drucker, H., Burges, C. J. C., Kaufman, L., Smola, A. & Vapnik, V. Support vector regression machines. Adv. Neural Inf. Process. Syst. 9, 155–161 (1997).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Shao, C., Zhang, Y. & Sun, W. Statistical characterization of HCD fragmentation patterns of tryptic peptides on an LTQ Orbitrap Velos mass spectrometer. J. Proteomics 109, 26–37 (2014).
Sundararajan, M., Taly, A. & Yan, Q. Axiomatic attribution for deep networks. Preprint at https://arxiv.org/abs/1703.01365 (2017).
Schubert, O. T. et al. Building high-quality assay libraries for targeted analysis of SWATH MS data. Nat. Protoc. 10, 426–441 (2015).
Wu, J. X. et al. SWATH mass spectrometry performance using extended peptide MS/MS assay libraries. Mol. Cell. Proteomics 15, 2501–2514 (2016).
Tsou, C.-C. et al. DIA-Umpire: comprehensive computational framework for data-independent acquisition proteomics. Nat. Methods 12, 258–264 (2015).
Bruderer, R. et al. Optimization of experimental parameters in data-independent mass spectrometry significantly increases depth and reproducibility of results. Mol. Cell. Proteomics 16, 2296–2309 (2017).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Tyanova, S., Temu, T. & Cox, J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 11, 2301–2319 (2016).
Vizcaíno, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44, D447–D456 (2016).
Nanjappa, V. et al. Plasma proteome database as a resource for proteomics research: 2014 update. Nucleic Acids Res. 42, D959–D965 (2014).
Mallick, P. et al. Computational prediction of proteotypic peptides for quantitative proteomics. Nat. Biotechnol. 25, 125–131 (2007).
Sanders, W. S., Bridges, S. M., McCarthy, F. M., Nanduri, B. & Burgess, S. C. Prediction of peptides observable by mass spectrometry applied at the experimental set level. BMC Bioinformatics 8, S23 (2007).
Zolg, D. P. et al. Building proteometools based on a complete synthetic human proteome. Nat. Methods 14, 259–262 (2017).
Hochreiter, S. & Schmidhuber, J. J. Long short-term memory. Neural Comput. 9, 1–32 (1997).
Hahnioser, R. H. R., Sarpeshkar, R., Mahowald, M. A., Douglas, R. J. & Seung, H. S. Digital selection and analogue amplification coexist in a cortex-inspired silicon circuit. Nature 405, 947–951 (2000).
Abadi, M. et al. TensorFlow: a system for large-scale machine learning. In Proc. 12th USENIX Symposium on Operating Systems Design and Implementation (OSDI ’16) 265–284 (USENIX Association, 2016).
Golovin, D. et al. Google Vizier: a service for black-box optimization. In Proc. 23rd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining 1487–1495 (ACM, 2017).
Kingma, D. P. & Ba, J. L. Adam: a method for stochastic optimization. Preprint at https://arxiv.org/abs/1412.6980 (2015).
Hunt, D. F., Yates, J. R., Shabanowitz, J., Winston, S. & Hauer, C. R. Protein sequencing by tandem mass spectrometry. Proc. Natl Acad. Sci. USA 83, 6233–6237 (1986).
Kelstrup, C. D. et al. Performance evaluation of the q exactive hf-x for shotgun proteomics. J. Proteome Res. 17, 727–738 (2018).
Krokhin, O. V. Sequence-specific retention calculator. ALGORITHM for peptide retention prediction in ion-pair RP-HPLC: application to 300- and 100-Å pore size C18 sorbents. Anal. Chem. 78, 7785–7795 (2006).
Acknowledgements
This project has received funding from the European Union’s EU Framework Program for Research and Innovation Horizon 2020 under grant agreement no. 686547 (S.T.) and from FP7 grant agreement no. GA ERC-2012-SyG_318987–ToPAG (J.C.). J.C. and P.G. are supported by the Marie Skłodowska-Curie European Training Network TEMPERA, a project funded by the European Union’s EU Framework Program for Research and Innovation Horizon 2020 under grant agreement no. 722606. We thank E. Deutsch, J. Bingham, M. Liu, R. Perrone, P. Kheradpour, B. Brown, M. Edwards, L. Cao, N. Soltero, J. Lehar, T. Snyder, D. Glazer and T. Stanis for their help, support and suggestions.
Author information
Authors and Affiliations
Contributions
S.T., R.L., P.G., F.S.S., P.C. and J.C. designed and developed the code, and performed the analyses. M.B. and L.D. helped with deep learning architecture design, as well as with reviewing the code and analyses. A.B. helped with data ingestion and preprocessing. K.K.P. carried out the wet-laboratory experiments and the DIA data analysis. R.L., P.C. and J.C. wrote the manuscript and directed the project. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.L., A.B. and P.C. are employees of Verily Life Sciences. L.D. and M.B. are employees of Google LLC. Verily Life Sciences and Google LLC had no role in decisions related to the study/work, data collection or analysis of data described in this paper.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Summary of training, validation and testing datasets.
The charts show distribution of various peptide and acquisition properties in our training dataset (based on total of 1,263,431 unique sequence/charge/fragmentation/mass analyzer peptide combinations). Note that all counts on the vertical axes are on a logarithmic scale.
Supplementary Figure 2 Performance of DeepMass:Prism.
The box plots show distributions of correlation coefficients (PCC) between actual and predicted y- and b-ion peak intensities for each peptide in our testing dataset, stratified by peptide charge and length, and by mass analyzer and fragmentation type. Each box extends from the lower to upper quartile values of the data, with a line at the median. The whiskers extend to 1.5-multiple past the interquartile range between the low and high quartiles. Values beyond these ranges are considered outliers, and are plotted as diamonds. Precursor counts represented by each box (N) are listed in tables below each box plot.
Supplementary Figure 3 Performance of DeepMass:Prism with and without metadata.
The box plots show distributions of correlation coefficients (PCC) between actual and predicted y- and b-ion peak intensities for each peptide in our testing dataset, based on our final model with metadata features (precursor length, precursor charge, mass analyzer, fragmentation type; right), and a model with no metadata features (left). Each box extends from the lower to upper quartile values of the data, with a line at the median. The whiskers extend to 1.5-multiple past the interquartile range between the low and high quartiles. Values beyond these ranges are considered outliers, and are plotted as diamonds. The number of precursor (N) in both distributions is 69,888.
Supplementary Figure 4 Sliding window-based approach using different machine learning algorithms.
Distribution of PCCs of the window-based approach using different machine learning algorithms such as support vector regression (SVR), random forest (RF), RF layer on top of the output of SVR (SVR+RF) and two layer neural networks (NN). The comparison was done on CID+2 model and window size 8. Each box extends from the lower to upper quartile values of the data, with a line at the median. The whiskers extend to 1.5-multiple past the interquartile range between the low and high quartiles. Values beyond these ranges are considered outliers, and are plotted as diamonds. The boxplots contain 9,214 unique PSMs from the ProteomeTools dataset (PXD004732).
Supplementary Figure 5 Fragmentation efficiency between residue pairs.
Fragmentation efficiencies were calculated for b- and y- fragment ions generated between [X] (y-axis) and [Z] (x-axis) residues in A-A-A-[X]-[Z]-A-A-A-R peptides, for both CID and HCD fragmentation. The fragmentation efficiency is defined as a predicted peak intensity, normalized by the total sum of peak intensities of the same ion type. Similar to the previous findings (Shao et al., 2014), our model reports significantly higher fragmentation efficiency between [X]-Pro residues (where [X] can be any other residues), for both y- and b- ion types. The model also correctly predicts less efficient fragmentation between [X]-[Z] residue pairs where [X] is a hydrophobic residue. Furthermore, the model also correctly identified an increased fragmentation efficiency for b-ions between His-[Z] pairs.
Supplementary Figure 6 Long-range interactions in the training data.
In our dataset we found 63 peptide pairs with a single residue mismatch, but the same precursor charge, fragmentation, and mass analyzer types. We evaluated the effect of a position more than one residue away from the site of fragmentation by measuring standard deviations in peak intensities across all 63 peptide pairs (for example, between the y 3 -ion intensity in spectra for both peptides in the pair). The box plots show distributions of peak intensities when looking at positions that are N residues (x-axis) away from the site of the mismatch. Each box extends from the lower to upper quartile values of the data, with a red line at the median. The whiskers extend to 1.5-multiple past the interquartile range between the low and high quartiles. Data points beyond these ranges are considered outliers, and are plotted as diamonds. The numbers of pairs (N) for each box are shown in the table below the box plots.
Supplementary Figure 7 Window size dependent performance of the sliding window approach.
Insert figure caption here Distribution of PCCs for MS2PIP and for the window-based neural network model for sliding window sizes of 4, 8, 16 and 24 residues on CID+2 model. The predictive performance increases monotonically with window size. The window-based neural network model outperforms MS2PIP at sufficiently large window sizes. by deleting or overwriting this text; captions may run to a second page if necessary. Each box extends from the lower to upper quartile values of the data, with a line at the median. The whiskers extend to 1.5-multiple past the interquartile range between the low and high quartiles. Values beyond these ranges are considered outliers, and are plotted as diamonds. The boxplots contain 9214 unique PSMs from the ProteomeTools dataset (PXD004732).
Supplementary Figure 8 Interpretation of the DeepMass:Prism model.
Distributions of distance between peak intensity and major attributions are shown for y (top) and b (bottom) ions. Major attributions are attribution values (heat map pixels in top panels of Fig. 4) with absolute values greater than or equal to 0.7. Directional distances are in amino-acid residue counts from the cleavage site.
Supplementary Figure 9 Application to spectral library generation for DIA data analysis.
Left: The scatter plots show the correlation between log-transformed peptide peak intensities as identified using spectral libraries based on DDA experiments and on MS2PIP (top), DeepMass:Prism (middle), or DeepMass:Prism+Drip (bottom). Pearson correlation coefficients are shown in the upper-left corner of each correlation plot. Right (top): The histograms show the distributions of log-transformed Q-values for peptides detected in DDA-based spectral library searches, but that were missed in searches with spectral libraries based on MS2PIP, DeepMass:Prism, and DeepMass:Prism+Drip. As illustrated, a majority of the missed peptides have a Q-value worse than 10-3. Right (bottom): The histogram shows the distributions of log-transformed peak intensities for peptides detected with the DDA-based spectral library, but missed in results with spectral libraries based on MS2PIP, DeepMass:Prism, and DeepMass:Prism+Drip. The numbers of data points (N) in these plots are based on the number of identified peptides: 5,248, 5,181, 3,976 for the DeepMass:Prism-, DeepMass:Prism+Drip-, and MS2PIP-based spectral libraries, respectively. The Q-values in the upper-right chart are as reported by Spectronaut.
Supplementary Figure 10 Top and bottom 5 predictions by DeepMass:Prism from our testing set.
The five best and worst predictions are shown. For each example, the predicted MS/MS spectra and the actual MS/MS spectra are shown above and below the x-axis, respectively. Badly predicted spectra tend to have highly intense fragments at the beginning or the end of a series. The correlation analyses with the peptides from the testing sets were only performed once.
Supplementary Figure 11 Performance of the sliding window approach split by precursor charge and fragmentation type.
Distribution of PCCs for the sliding window-based neural network model with a window size of 24 (orange) and MS2PIP (blue) for comparison. The neural network approach results in better performance in all subclasses. Each box extends from the lower to upper quartile values of the data, with a line at the median. The whiskers extend to 1.5-multiple past the interquartile range between the low and high quartiles. Values beyond these ranges are considered outliers, and are plotted as diamonds. The boxplots contain 9,214, 9,317, 6,540 and 6,773 unique PSMs, respectively, for CID+2, HCD+2, CID+3 and HCD+3 test data from the ProteomeTools dataset (PXD004732).
Supplementary Figure 12 Per-residue distances of major attributions.
Each row represents a single histogram of the type displayed in Fig. 3, on a per-residue basis. Specifically, each row represents the distribution of distances over which a given residue has a strong influence (absolute attribution value ≥ 0.7) on a peak’s predicted intensity. Each row is frequency-normalized such that the area under the distribution sums to 1.0. Rows of each set of distributions were clustered via single linkage of PCC. Most rows resemble the overall trend, and notable exceptions are discussed in the text. The distributions in the heatmaps are based on 1000 randomly selected peptide sequences.
Supplementary Figure 13 Example I for correct PSM recovery by intensity prediction.
Including intensity predictions for theoretical spectra in the Andromeda score calculation enables the correct PSM to move from the second best score to the top scoring position. For example, before considering intensity predictions, the sequence of two adjacent amino acids is incorrect (a), but when including intensity information in the score, the sequence with the correct order becomes the highest-scoring PSM (b).
Supplementary Figure 14 Example II for correct PSM recovery by intensity prediction.
Including intensity predictions for theoretical spectra in the Andromeda score calculation enables a completely different peptide sequence to become the top-scoring PSM. For example, before considering intensity predictions, the peptide sequence matched to a particular MS/MS spectrum was low but passing (?) (a); however, after including intensity predictions, a new sequence with more matches to higher intense peaks and a more complete ion-series identification was obtained (b).
Supplementary information
Supplementary Information
Supplementary Figs. 1–14 and Supplementary Table 1
Rights and permissions
About this article
Cite this article
Tiwary, S., Levy, R., Gutenbrunner, P. et al. High-quality MS/MS spectrum prediction for data-dependent and data-independent acquisition data analysis. Nat Methods 16, 519–525 (2019). https://doi.org/10.1038/s41592-019-0427-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0427-6
This article is cited by
-
Coherence mapping to identify the intermediates of multi-channel dissociative ionization
Communications Chemistry (2024)
-
Deep-time phylogenetic inference by paleoproteomic analysis of dental enamel
Nature Protocols (2024)
-
Deep learning prediction of glycopeptide tandem mass spectra powers glycoproteomics
Nature Machine Intelligence (2024)
-
Prediction of glycopeptide fragment mass spectra by deep learning
Nature Communications (2024)
-
SeFilter-DIA: Squeeze-and-Excitation Network for Filtering High-Confidence Peptides of Data-Independent Acquisition Proteomics
Interdisciplinary Sciences: Computational Life Sciences (2024)