Abstract
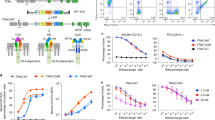
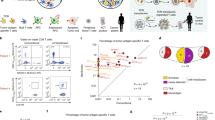
CD8+ T cells recognize and eliminate tumors in an antigen-specific manner. Despite progress in characterizing the antitumor T cell repertoire and function, the identification of target antigens remains a challenge. Here we describe the use of chimeric receptors called signaling and antigen-presenting bifunctional receptors (SABRs) in a cell-based platform for T cell receptor (TCR) antigen discovery. SABRs present an extracellular complex comprising a peptide and major histocompatibility complex (MHC), and induce intracellular signaling via a TCR-like signal after binding with a cognate TCR. We devised a strategy for antigen discovery using SABR libraries to screen thousands of antigenic epitopes. We validated this platform by identifying the targets recognized by public TCRs of known specificities. Moreover, we extended this approach for personalized neoantigen discovery.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request. The raw data for Figs. 1–5 and Supplementary Figs. 1, 3, 4, 7, and 8 can be found in the Source Data files. The list of epitopes in the SABR libraries can be found in Supplementary Tables 3 and 4. The plasmids for HLA-A*0201-SABR backbone (pCCLc-MND-A0201-SABR-Backbone; ID 119050), HLA-B*2705-SABR backbone (pCCLc-MND-B2705-SABR-Backbone; ID 119051), A2-Mart1-SABR (pCCLc-MND-A0201-Mart1-SABR; ID 119052), and B27-KK10-SABR (pCCLc-MND-B2705-KK10-SABR; ID 119053) are available through Addgene. The sequencing data have been deposited in the Sequence Read Archive (SRR8207921, amplicon sequencing of A2-SABR-library co-incubated with F5 TCR; SRR8207922, amplicon sequencing of A2-SABR-library co-incubated with SL9 TCR; SRR8207923, amplicon sequencing of A2-SABR-library co-incubated with no TCR; SRR8207924, amplicon sequencing of A2-NeoAg-library co-incubated with neoTCR; SRR8207925, amplicon sequencing of A2-NeoAg-library co-incubated with no TCR). The code used to analyze sequences has been deposited in GitHub (https://github.com/Baltimore-Lab/nat-methods-SABR-trogo).
References
Shankaran, V. et al. IFN-γ and lymphocytes prevent primary tumour development and shape tumour immunogenicity. Nature 410, 1107–1111 (2001).
Lollini, P. L., Cavallo, F., Nanni, P. & Forni, G. Vaccines for tumour prevention. Nat. Rev. Cancer 6, 204–216 (2006).
Leach, D. R., Krummel, M. F. & Allison, J. P. Enhancement of antitumor immunity by CTLA-4 blockade. Science 271, 1734–1736 (1996).
Dong, H. et al. Tumor-associated B7-H1 promotes T-cell apoptosis: a potential mechanism of immune evasion. Nat. Med. 8, 793–800 (2002).
Yee, C. et al. Adoptive T cell therapy using antigen-specific CD8+ T cell clones for the treatment of patients with metastatic melanoma: in vivo persistence, migration, and antitumor effect of transferred T cells. Proc. Natl Acad. Sci. USA 99, 16168–16173 (2002).
Davis, M. M. & Bjorkman, P. J. T-cell antigen receptor genes and T-cell recognition. Nature 334, 395–402 (1988).
Weiss, A. & Littman, D. R. Signal transduction by lymphocyte antigen receptors. Cell 76, 263–274 (1994).
Bethune, M. T. & Joglekar, A. V. Personalized T cell-mediated cancer immunotherapy: progress and challenges. Curr. Opin. Biotechnol. 48, 142–152 (2017).
Woodsworth, D. J., Castellarin, M. & Holt, R. A. Sequence analysis of T-cell repertoires in health and disease. Genome Med. 5, 98 (2013).
Buchholz, V. R., Schumacher, T. N. & Busch, D. H. T cell fate at the single-cell level. Annu. Rev. Immunol. 34, 65–92 (2016).
Klenerman, P., Cerundolo, V. & Dunbar, P. R. Tracking T cells with tetramers: new tales from new tools. Nat. Rev. Immunol. 2, 263–272 (2002).
Castle, J. C. et al. Exploiting the mutanome for tumor vaccination. Cancer Res. 72, 1081–1091 (2012).
Boon, T. & van der Bruggen, P. Human tumor antigens recognized by T lymphocytes. J. Exp. Med. 183, 725–729 (1996).
Matsushita, H. et al. Cancer exome analysis reveals a T-cell-dependent mechanism of cancer immunoediting. Nature 482, 400–404 (2012).
Gee, M. H. et al. Antigen identification for orphan T cell receptors expressed on tumor-infiltrating lymphocytes. Cell 172, 549–563 (2018).
Birnbaum, M. E. et al. Deconstructing the peptide-MHC specificity of T cell recognition. Cell 157, 1073–1087 (2014).
Yu, Y. Y., Netuschil, N., Lybarger, L., Connolly, J. M. & Hansen, T. H. Cutting edge: single-chain trimers of MHC class I molecules form stable structures that potently stimulate antigen-specific T cells and B cells. J. Immunol. 168, 3145–3149 (2002).
Morgan, R. A. et al. Cancer regression in patients after transfer of genetically engineered lymphocytes. Science 314, 126–129 (2006).
Joglekar, A. V. et al. T cell receptors for the HIV KK10 epitope from patients with differential immunologic control are functionally indistinguishable. Proc. Natl Acad. Sci. USA 115, 1877–1882 (2018).
Bennett, M. S., Joseph, A., Ng, H. L., Goldstein, H. & Yang, O. O. Fine-tuning of T-cell receptor avidity to increase HIV epitope variant recognition by cytotoxic T lymphocytes. AIDS 24, 2619–2628 (2010).
Bethune, M. T., Comin-Anduix, B., Hwang Fu, Y. H., Ribas, A. & Baltimore, D. Preparation of peptide-MHC and T-cell receptor dextramers by biotinylated dextran doping. Biotechniques 62, 123–130 (2017).
Sahin, U. et al. Personalized RNA mutanome vaccines mobilize poly-specific therapeutic immunity against cancer. Nature 547, 222–226 (2017).
Vita, R. et al. The immune epitope database (IEDB) 3.0. Nucleic Acids Res. 43, D405–D412 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Yokomaku, Y. et al. Impaired processing and presentation of cytotoxic-T-lymphocyte (CTL) epitopes are major escape mechanisms from CTL immune pressure in human immunodeficiency virus type 1 infection. J. Virol. 78, 1324–1332 (2004).
Dorrell, L. et al. Distinct recognition of non-clade B human immunodeficiency virus type 1 epitopes by cytotoxic T lymphocytes generated from donors infected in Africa. J. Virol. 73, 1708–1714 (1999).
Li, G. S. W. et al. T cell antigen discovery via trogocytosis. Nat. Methods https://doi.org/10.1038/s41592-018-0305-7 (2019).
Peakman, M. et al. T cell clones generated from patients with type 1 diabetes using interleukin-2 proliferate to human islet antigens. Autoimmunity 17, 31–39 (1994).
Tang, Q. & Bluestone, J. A. The Foxp3+ regulatory T cell: a jack of all trades, master of regulation. Nat. Immunol. 9, 239–244 (2008).
Garcia, K. C. et al. Structural basis of plasticity in T cell receptor recognition of a self peptide-MHC antigen. Science 279, 1166–1172 (1998).
Suwandi, J. S., Nikolic, T. & Roep, B. O. Translating mechanism of regulatory action of tolerogenic dendritic cells to monitoring endpoints in clinical trials. Front. Immunol. 8, 1598 (2017).
Bentzen, A. K. & Hadrup, S. R. Evolution of MHC-based technologies used for detection of antigen-responsive T cells. Cancer Immunol. Immunother. 66, 657–666 (2017).
Tran, E. et al. Cancer immunotherapy based on mutation-specific CD4+ T cells in a patient with epithelial cancer. Science 344, 641–645 (2014).
Baltimore, D. et al. A cell-based platform for T cell antigen discovery: engineered antigen presenting cells expressing signaling and antigen presenting bifunctional receptors (SABRs). Protocol Exchange https://doi.org/10.1038/protex.2018.126 (2019).
Acknowledgements
We thank I. Antoshechkin at the Millard and Muriel Jacobs Genetics and Genomics Laboratory for Illumina sequencing, and A. Spalla at the Analytical Cytometry Core at the City of Hope for help with FACS. NFAT-GFP-Jurkat cells were a gift from A. Weiss (University of California, San Francisco, San Francisco, CA, USA) and Y. Chen (University of California, Los Angeles, Los Angeles, CA, USA). GXR-B27+ cells were a gift from B.D. Walker (Ragon Institute of Massachusetts General Hospital, Massachusetts Institute of Technology, and Harvard, Cambridge, MA, USA). J3 chimeric antigen receptor was a gift from P. Wang (University of Southern California, Los Angeles, CA, USA). The pCCLc-MND-X backbone and pCMV-RD8.9 were gifts from D.B. Kohn (University of California, Los Angeles, Los Angeles, CA, USA). MSCV-based shuttle plasmid was a gift from R.A. Morgan (National Institutes of Health, Bethesda, MD, USA). This work was funded by the California Institute for Regenerative Medicine (award DISC2-09123 to D.B.), the Caltech Rothenberg Innovation Initiative (to D.B.), and the US National Cancer Institute (grant 1U54 CA199090-01 to J.R.H.).
Author information
Authors and Affiliations
Contributions
A.V.J. designed and performed experiments, analyzed and interpreted the data, and wrote the manuscript. M.T.L. designed and performed experiments, performed computational analyses, and interpreted the data. M.S. and J.D.J. designed and performed experiments, and analyzed the data. G.L., S.W., S.P., J.M.Z., and M.T.B. designed and performed experiments, and contributed reagents. J.R.H. and A.R. contributed reagents and supervised experiments. D.B. supervised the experiments, analyzed and interpreted the data, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.V.J., M.T.L., M.T.B., and D.B. are named as co-inventors on a patent application concerning the described technology. D.B. is a consultant of PACT and head of their scientific advising board. J.R.H. and A.R. are directors and consultants of PACT; M.T.B. and S.P. are employees of PACT; J.M.Z. is a consultant of PACT; and each of the foregoing individuals has equity interests in PACT.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Variants of SABR constructs.
a. Schematics showing the SABR-F and SABR-E constructs. SABR-F contains the transmembrane domain from HLA (black rectangle), whereas SABR-E contains the transmembrane domain from CD3ζ (teal rectangle). The two horizontal lines indicate two leaflets of the plasma membrane. b. Representative flow cytometry plots showing GFP expression in coculture assays from Fig 1b. The experiments were performed at n = 3 biologically independent cell culture replicates. c. GFP expression in coculture assays comparing SABR-F and SABR-E. The lines and error bars indicate mean ± s.d. from n = 12 biologically independent cell culture replicates.
Supplementary Figure 2 Time course of GFP expression induced by SABRs.
Representative flow cytometry plots from the F5+A2-MART1 experiment enumerated in Fig 1e are shown. The rectangle in the right bottom corners shows the gate for counting GFP+ cells. The time at which each sample was collected is shown as hours. The frequency of cells in the GFP+ gate is indicated as a percentage.
Supplementary Figure 3 Recognition of low-affinity TCRs by SABRs.
GFP expression in coculture assays using F5 and M1 TCRs with A2-MART1-SABR. The dots indicate individual values from n = 2 biologically independent cell culture replicates.
Supplementary Figure 4 Empty SABR vector constructs and recognition of low-affinity pMHC–TCR interactions.
a. Schematic showing SCT, SABRs, empty SABRs, and TMGs. EP, epitope; S, signal sequence; MITD, MHC class I trafficking signal; numbers 1–6 indicate Gly–Ser linkers. b. Correlation of functional avidity of interaction of EC27 TCR with variants of KK10 peptides with their ability to initiate signal through SABRs. The indicated peptides are variants of the KK10 epitope. R2T, KTWIILGLNK; R2I, KIWIILGLNK; R2G, KGWIILGLNK; R2K, KKWIILGLNK; WT, KRWIILGLNK; R2Q, KQWIILGLNK.
Supplementary Figure 5 Strategy to clone custom oligonucleotides into SABR vectors.
SABR vector constructs with a stuffer fragment showing BsmBI sites (top), and cloning strategy using double-stranded oligonucleotides with encoding the epitope flanked by overlaps.
Supplementary Figure 6 SABR library screen for antigen discovery.
a. Schematic showing the pipeline to construct custom SABR libraries. EP, epitope. The left panel shows the procedure to obtain and synthesize a list of epitopes. The right panel shows the schematic of the SABR library. b. Schematic showing coculture experiment to select cells from SABR library that are recognized by an orphan TCR. Left panel shows a SABR library presenting numerous unique epitopes. The middle panel shows antigen-presenting cells (APCs) showing reporter expression induced by SABRs presenting the cognate epitope for the orphan TCR. Right panel shows processing of the selected cells. c. Flowchart showing the computational analysis pipeline.
Supplementary Figure 7 Enrichment of EAAGIGILTV and SLYNTVATL analogs in SABR library screen.
a. Average ranks for all the EAAGIGILTV analogs in the A2-SABR library. b. Average ranks for all the SLYNTVATL analogs in the A2-SABR library. The ranks were calculated as described in the manuscript. The data are averaged from three biologically independent cell culture replicates.
Supplementary Figure 8 Enrichment of USP7 neoepitopes in SABR library screen.
Average ranks for all the USP7-derived epitopes in the NeoAg-SABR library. The data are averaged from three biologically independent cell culture replicates.
Supplementary Figure 9 Gating strategy used in flow cytometry.
a. Gating strategy used in coculture assays to measure GFP expression. b. Gating strategy used in coculture assays to measure GFP and CD69 expression. c. Gating strategy used in cytotoxicity assays.
Supplementary information
Supplementary Text and Figures
Supplementary Figs. 1–9 and Supplementary Tables 1 and 2
Supplementary Protocol
A cell-based platform for T cell antigen discovery: engineered antigen-presenting cells expressing signaling and antigen-presenting bifunctional receptors (SABRs).
Supplementary Table 3
The list of epitopes in the A2-SABR library.
Supplementary Table 4
The list of epitopes in the A2-NeoAg library.
Rights and permissions
About this article
Cite this article
Joglekar, A.V., Leonard, M.T., Jeppson, J.D. et al. T cell antigen discovery via signaling and antigen-presenting bifunctional receptors. Nat Methods 16, 191–198 (2019). https://doi.org/10.1038/s41592-018-0304-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-018-0304-8
This article is cited by
-
Novel insights into TCR-T cell therapy in solid neoplasms: optimizing adoptive immunotherapy
Experimental Hematology & Oncology (2024)
-
Neoantigen-targeted TCR-engineered T cell immunotherapy: current advances and challenges
Biomarker Research (2023)
-
The screening, identification, design and clinical application of tumor-specific neoantigens for TCR-T cells
Molecular Cancer (2023)
-
Identification of patient-specific CD4+ and CD8+ T cell neoantigens through HLA-unbiased genetic screens
Nature Biotechnology (2023)
-
Can we predict T cell specificity with digital biology and machine learning?
Nature Reviews Immunology (2023)