Abstract
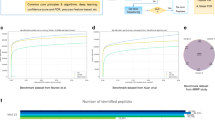
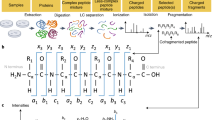
We present DeepNovo-DIA, a de novo peptide-sequencing method for data-independent acquisition (DIA) mass spectrometry data. We use neural networks to capture precursor and fragment ions across m/z, retention-time, and intensity dimensions. They are then further integrated with peptide sequence patterns to address the problem of highly multiplexed spectra. DIA coupled with de novo sequencing allowed us to identify novel peptides in human antibodies and antigens.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
Data and a pretrained model are publicly available in the MassIVE repository under accession number MSV000082368. Source data for Fig. 2 are available online.
References
Ott, P. A. et al. Nature 547, 217–221 (2017).
Sahin, U. et al. Nature 547, 222–226 (2017).
Anonymous. Nat. Biotechnol. 35, 97 (2017).
Vitiello, A. & Zanetti, M. Nat. Biotechnol. 35, 815–817 (2017).
Bassani-Sternberg, M. et al. Nat. Commun. 7, 13404 (2016).
Venable, J. D., Dong, M. Q., Wohlschlegel, J., Dillin, A. & Yates, J. R. Nat. Methods 1, 39–45 (2004).
Röst, H. L. et al. Nat. Biotechnol. 32, 219–223 (2014).
Egertson, J. D., MacLean, B., Johnson, R., Xuan, Y. & MacCoss, M. J. Nat. Protoc. 10, 887–903 (2015).
Tsou, C. C. et al. Nat. Methods 12, 258–264 (2015).
Ting, Y. S. et al. Nat. Methods 14, 903–908 (2017).
Tran, N. H., Zhang, X., Xin, L., Shan, B. & Li, M. Proc. Natl Acad. Sci. USA 114, 8247–8252 (2017).
Zhang, J. et al. Mol. Cell. Proteomics. 11, M111.010587 (2012).
Muntel, J. et al. J. Proteome. Res. 14, 4752–4762 (2015).
Bruderer, R. et al. Mol. Cell. Proteomics. 14, 1400–1410 (2015).
Tan, J. et al. Nature 529, 105–109 (2016).
Caron, E. et al. eLife 4, e07661 (2015).
Lin, T. Y., Goyal, P., Girshick, R., He, K. & Dollár, P. Focal loss for dense object detection. arXiv Preprint at https://arxiv.org/abs/1708.02002 (2017).
Tyanova, S., Temu, T. & Cox, J. Nat. Protoc. 11, 2301–2319 (2016).
Acknowledgements
This work was funded in part by NSERC (grant OGP0046506), China’s Research and Development Program (grants 2016YFB1000902 and 2018YFB1003202), the NSFC (grant 61832019), and the Canada Research Chair program for M.L. N.H.T. was supported by the Mitacs Elevate Fellowship. The authors thank N. Keshav, K.P. Choi, and K. Xiong for discussions and proofreading of the manuscript.
Author information
Authors and Affiliations
Contributions
M.L., B.S., and N.H.T. conceived the research idea. N.H.T. designed the model, implemented the software, and analyzed the results. R.Q. and X.C. contributed to the model design, software development, and data analysis. M.L., B.S., and L.X. supervised the research project. C.L., X.Z., and A.G. contributed to the data analysis. N.H.T., M.L., and R.Q. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
L.X., X.C., and B.S. are employees of Bioinformatics Solutions Inc., Waterloo, Ontario, Canada.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Comparison of unique peptides identified by DeepNovo, PECAN, and Spectronaut from the plasma dataset.
Note that the number of 3,268 de novo peptides reported here by DeepNovo have yet been validated (see Supplementary Note 1)
Supplementary Figure 2
Distribution of de novo confidence score versus peptide abundance of all peptides reported by DeepNovo from the plasma dataset
Supplementary Figure 3
Distribution of retention times of 1,143 de novo peptides reported by DeepNovo and database peptides reported by PECAN and Spectronaut from the plasma dataset
Supplementary Figure 4
Distribution of amino acids of 1,143 de novo peptides reported by DeepNovo and database peptides reported by PECAN and Spectronaut from the plasma dataset
Supplementary Figure 5 Unique peptides identified by DeepNovo, PECAN, and Spectronaut from the plasma dataset.
(a) Original model trained with the urine dataset. (b) Model retrained with part of the plasma dataset. Note that we have removed from DeepNovo the features that were used to retrain the model, so the numbers of DeepNovo peptides in a are less than those reported in Supplementary Fig. 1. DeepNovo peptides have not been filtered by sequencing errors and augmented database search (see Supplementary Note 1)
Supplementary Figure 6
Example of three de novo peptides aligned to the variable region of a recently published human antibody for malaria vaccine design
Supplementary Figure 7
Unique peptides identified by DeepNovo, OpenSWATH, and Spectronaut from the dataset Jurkat-Oxford
Supplementary Figure 8
Abundance distribution of 130 de novo peptides versus 102 peptides identified by DeepNovo and OpenSWATH or Spectronaut from the dataset Jurkat-Oxford
Supplementary Figure 9
DeepNovo sequencing framework
Supplementary Figure 10
Ion-CNN model
Supplementary Figure 11
Spectrum-CNN model
Supplementary information
Supplementary Information
Supplementary Figures 1–11 and Supplementary Notes 1 and 2
Supplementary Protocol
Documentation for using DeepNovo
Supplementary Software
Scripts and Swiss-Prot database FASTA file
Supplementary Data 1
Examples of DIA spectra from the plasma dataset that contain multiple precursors with at least one low-abundance, novel peptide identified by DeepNovo but not by other database search tools
Supplementary Data 2
Twelve examples from the plasma dataset showing that the low-abundance, novel peptides identified by DeepNovo have better supporting fragment ions than those candidate sequences returned by the database search engine
Supplementary Data 3
Evidence of supporting fragment ions (left column), coelution profiles of fragment ions and precursor ion (right column), and antibody protein ID for 30 low-abundance, novel peptides identified by DeepNovo from the plasma dataset. The database search engine was not able to find any candidate sequences that matched these 30 precursors
Supplementary Data 4
Twelve examples of low-abundance, novel HLA peptides that were identified by DeepNovo but not by other database search tools
Supplementary Table 1
Summary of training and testing datasets in our study
Supplementary Table 2
List of 2,753 unique peptides predicted by DeepNovo from the plasma dataset
Supplementary Table 3
Novel peptides that were identified by DeepNovo from the plasma dataset and were found in variable regions of human immunoglobulin light chains
Supplementary Table 4
Novel peptides that were identified by DeepNovo from the plasma dataset and were found in variable regions of human immunoglobulin heavy chains
Supplementary Table 5
Novel peptides that were identified by DeepNovo from the plasma dataset and contained human natural variants
Supplementary Table 6
List of 304 unique peptides predicted by DeepNovo from the Jurkat-Oxford dataset
Source data
Rights and permissions
About this article
Cite this article
Tran, N.H., Qiao, R., Xin, L. et al. Deep learning enables de novo peptide sequencing from data-independent-acquisition mass spectrometry. Nat Methods 16, 63–66 (2019). https://doi.org/10.1038/s41592-018-0260-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-018-0260-3
This article is cited by
-
MARS an improved de novo peptide candidate selection method for non-canonical antigen target discovery in cancer
Nature Communications (2024)
-
Python workflow for the selection and identification of marker peptides—proof-of-principle study with heated milk
Analytical and Bioanalytical Chemistry (2024)
-
SeFilter-DIA: Squeeze-and-Excitation Network for Filtering High-Confidence Peptides of Data-Independent Acquisition Proteomics
Interdisciplinary Sciences: Computational Life Sciences (2024)
-
Redefining serological diagnostics with immunoaffinity proteomics
Clinical Proteomics (2023)
-
InvitroSPI and a large database of proteasome-generated spliced and non-spliced peptides
Scientific Data (2023)