Abstract
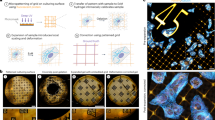
Many biological investigations require 3D imaging of cells or tissues with nanoscale spatial resolution. We recently discovered that preserved biological specimens can be physically expanded in an isotropic fashion through a chemical process. Expansion microscopy (ExM) allows nanoscale imaging of biological specimens with conventional microscopes, decrowds biomolecules in support of signal amplification and multiplexed readout chemistries, and makes specimens transparent. We review the principles of how ExM works, advances in the technology made by our group and others, and its applications throughout biology and medicine.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Dunn, R. C. Near-field scanning optical microscopy. Chem. Rev. 99, 2891–2928 (1999).
Dürig, U., Pohl, D. W. & Rohner, F. Near-field optical-scanning microscopy. J. Appl. Phys. 59, 3318–3327 (1986).
Hell, S.W. Far-field optical nanoscopy. In Proc. 2010 23rd Annual Meeting of the IEEE Photonics Society (eds Jagadish, C. et al.) 3–4 (IEEE, New York, 2010).
Huang, B., Bates, M. & Zhuang, X. Super-resolution fluorescence microscopy. Annu. Rev. Biochem. 78, 993–1016 (2009).
Chen, F., Tillberg, P. W. & Boyden, E. S. Expansion microscopy. Science 347, 543–548 (2015).
Tanaka, T. et al. Phase transitions in ionic gels. Phys. Rev. Lett. 45, 1636–1639 (1980).
Hausen, P. & Dreyer, C. The use of polyacrylamide as an embedding medium for immunohistochemical studies of embryonic tissues. Stain Technol. 56, 287–293 (1981).
Cohen, Y., Ramon, O., Kopelman, I. J. & Mizrahi, S. Characterization of inhomogeneous polyacrylamide hydrogels. J. Polym. Sci. B Polym. Phys. 30, 1055–1067 (1992).
Tillberg, P. W. et al. Protein-retention expansion microscopy of cells and tissues labeled using standard fluorescent proteins and antibodies. Nat. Biotechnol. 34, 987–992 (2016).
Chen, F. et al. Nanoscale imaging of RNA with expansion microscopy. Nat. Methods 13, 679–684 (2016).
Chang, J.-B. et al. Iterative expansion microscopy. Nat. Methods 14, 593–599 (2017).
Zhao, Y. et al. Nanoscale imaging of clinical specimens using pathology-optimized expansion microscopy. Nat. Biotechnol. 35, 757–764 (2017).
Chozinski, T. J. et al. Expansion microscopy with conventional antibodies and fluorescent proteins. Nat. Methods 13, 485–488 (2016).
Ku, T. et al. Multiplexed and scalable super-resolution imaging of three-dimensional protein localization in size-adjustable tissues. Nat. Biotechnol. 34, 973–981 (2016).
Truckenbrodt, S. et al. X10 expansion microscopy enables 25-nm resolution on conventional microscopes. EMBO Rep. 19, e45836 (2018).
Tsanov, N. et al. smiFISH and FISH-quant—a flexible single RNA detection approach with super-resolution capability. Nucleic Acids Res. 44, e165 (2016).
Asano, S. M. et al. Expansion microscopy: protocols for imaging proteins and RNA in cells and tissues. Curr. Protoc. Cell Biol. 80, e56 (2018).
Freifeld, L. et al. Expansion microscopy of zebrafish for neuroscience and developmental biology studies. Proc. Natl Acad. Sci. USA 114, E10799–E10808 (2017).
Migliori, B. et al. Light sheet theta microscopy for rapid high-resolution imaging of large biological samples. BMC Biol. 16, 57 (2018).
Hama, H. et al. Scale: a chemical approach for fluorescence imaging and reconstruction of transparent mouse brain. Nat. Neurosci. 14, 1481–1488 (2011).
Susaki, E. A. et al. Whole-brain imaging with single-cell resolution using chemical cocktails and computational analysis. Cell 157, 726–739 (2014).
Chung, K. et al. Structural and molecular interrogation of intact biological systems. Nature 497, 332–337 (2013).
Murakami, T. C. et al. A three-dimensional single-cell-resolution whole-brain atlas using CUBIC-X expansion microscopy and tissue clearing. Nat. Neurosci. 21, 625–637 (2018).
Zhang, Y. S. et al. Hybrid microscopy: enabling inexpensive high-performance imaging through combined physical and optical magnifications. Sci. Rep. 6, 22691 (2016).
Aoki, T., Tsuchida, S., Yahara, T. & Hamaue, N. Novel assays for proteases using green fluorescent protein-tagged substrate immobilized on a membrane disk. Anal. Biochem. 378, 132–137 (2008).
Nicholls, S. B. & Hardy, J. A. Structural basis of fluorescence quenching in caspase activatable-GFP. Protein Sci. 22, 247–257 (2013).
Deshpande, T. et al. Subcellular reorganization and altered phosphorylation of the astrocytic gap junction protein connexin43 in human and experimental temporal lobe epilepsy. Glia 65, 1809–1820 (2017).
Crittenden, J. R. et al. Striosome-dendron bouquets highlight a unique striatonigral circuit targeting dopamine-containing neurons. Proc. Natl Acad. Sci. USA 113, 11318–11323 (2016).
Decarreau, J. et al. The tetrameric kinesin Kif25 suppresses pre-mitotic centrosome separation to establish proper spindle orientation. Nat. Cell Biol. 19, 384–390 (2017).
Suofu, Y. et al. Dual role of mitochondria in producing melatonin and driving GPCR signaling to block cytochrome c release. Proc. Natl Acad. Sci. USA 114, E7997–E8006 (2017).
Wang, Y. et al. Combined expansion microscopy with structured illumination microscopy for analyzing protein complexes. Nat. Protoc. 13, 1869–1895 (2018).
Orth, A. et al. Super-multiplexed fluorescence microscopy via photostability contrast. Biomed. Opt. Express 9, 2943–2954 (2018).
Chozinski, T. J. et al. Volumetric, nanoscale optical imaging of mouse and human kidney via expansion microscopy. Sci. Rep. 8, 10396 (2018).
Tsai, A. et al. Nuclear microenvironments modulate transcription from low-affinity enhancers. eLife 6, e28975 (2017).
Jiang, N. et al. Superresolution imaging of Drosophila tissues using expansion microscopy. Mol. Biol. Cell 29, 1413–1421 (2018).
Cahoon, C. K. et al. Superresolution expansion microscopy reveals the three-dimensional organization of the Drosophila synaptonemal complex. Proc. Natl Acad. Sci. USA 114, E6857–E6866 (2017).
Sümbül, U. et al. Automated scalable segmentation of neurons from multispectral images. In Advances in Neural Information Processing Systems 29 (eds Lee, D. D. et al.) 1912–1920 (NIPS Foundation, La Jolla, CA, 2016).
Mosca, T. J., Luginbuhl, D. J., Wang, I. E. & Luo, L. Presynaptic LRP4 promotes synapse number and function of excitatory CNS neurons. eLife 6, 1–29 (2017).
Wang, I. E., Lapan, S. W., Scimone, M. L., Clandinin, T. R. & Reddien, P. W. Hedgehog signaling regulates gene expression in planarian glia. eLife 5, e16996 (2016).
Halpern, A. R., Alas, G. C. M., Chozinski, T. J., Paredez, A. R. & Vaughan, J. C. Hybrid structured illumination expansion microscopy reveals microbial cytoskeleton organization. ACS Nano 11, 12677–12686 (2017).
Artur, C. G. et al. Plasmonic nanoparticle-based expansion microscopy with surface-enhanced Raman and dark-field spectroscopic imaging. Biomed. Opt. Express 9, 603–615 (2018).
Villaseñor, R., Schilling, M., Sundaresan, J., Lutz, Y. & Collin, L. Sorting tubules regulate blood-brain barrier transcytosis. Cell Rep. 21, 3256–3270 (2017).
Li, R., Chen, X., Lin, Z., Wang, Y. & Sun, Y. Expansion enhanced nanoscopy. Nanoscale 10, 17552–17556 (2018).
Treweek, J. B. et al. Whole-body tissue stabilization and selective extractions via tissue-hydrogel hybrids for high-resolution intact circuit mapping and phenotyping. Nat. Protoc. 10, 1860–1896 (2015).
Unnersjö-Jess, D. et al. Confocal super-resolution imaging of the glomerular filtration barrier enabled by tissue expansion. Kidney Int. 93, 1008–1013 (2018).
Wei, L. et al. Super-multiplex vibrational imaging. Nature 544, 465–470 (2017).
Hu, F. et al. Supermultiplexed optical imaging and barcoding with engineered polyynes. Nat. Methods 15, 194–200 (2018).
Kumar, A. et al. Influenza virus exploits tunneling nanotubes for cell-to-cell spread. Sci. Rep. 7, 40360 (2017).
Gao, M. et al. Expansion stimulated emission depletion microscopy (ExSTED). ACS Nano 12, 4178–4185 (2018).
Elmore, J. G. et al. Diagnostic concordance among pathologists interpreting breast biopsy specimens. J. Am. Med. Assoc. 313, 1122–1132 (2015).
Choi, H. M. T. et al. Programmable in situ amplification for multiplexed imaging of mRNA expression. Nat. Biotechnol. 28, 1208–1212 (2010).
Choi, H. M. T., Beck, V. A. & Pierce, N. A. Next-generation in situ hybridization chain reaction: higher gain, lower cost, greater durability. ACS Nano 8, 4284–4294 (2014).
Lin, R. et al. A hybridization-chain-reaction-based method for amplifying immunosignals. Nat. Methods 15, 275–278 (2018).
Lee, J. H. et al. Highly multiplexed subcellular RNA sequencing in situ. Science 343, 1360–1363 (2014).
Ke, R. et al. In situ sequencing for RNA analysis in preserved tissue and cells. Nat. Methods 10, 857–860 (2013).
Yoon, Y.-G. et al. Feasibility of 3D reconstruction of neural morphology using expansion microscopy and barcode-guided agglomeration. Front. Comput. Neurosci. 11, 97 (2017).
Lubeck, E. & Cai, L. Single-cell systems biology by super-resolution imaging and combinatorial labeling. Nat. Methods 9, 743–748 (2012).
Lubeck, E., Coskun, A. F., Zhiyentayev, T., Ahmad, M. & Cai, L. Single-cell in situ RNA profiling by sequential hybridization. Nat. Methods 11, 360–361 (2014).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Moffitt, J. R. et al. High-throughput single-cell gene-expression profiling with multiplexed error-robust fluorescence in situ hybridization. Proc. Natl Acad. Sci. USA 113, 11046–11051 (2016).
Wang, G., Moffitt, J. R. & Zhuang, X. Multiplexed imaging of high-density libraries of RNAs with MERFISH and expansion microscopy. Sci. Rep. 8, 4847 (2018).
Shah, S., Lubeck, E., Zhou, W. & Cai, L. In situ transcription profiling of single cells reveals spatial organization of cells in the mouse hippocampus. Neuron 92, 342–357 (2016).
Wang, Y. et al. Rapid sequential in situ multiplexing with DNA-exchange-imaging in neuronal cells and tissues. Nano Lett. 17, 6131–6139 (2017).
Acknowledgements
E.S.B. acknowledges the NIH (1R01NS102727, 1R01EB024261, 1R41MH112318, 1R01MH110932, 1RM1HG008525, and Director's Pioneer Award 1DP1NS087724), the Open Philanthropy Project, DARPA, John Doerr, the NSF (grant 1734870), the MIT Aging Brain Initiative/Ludwig Foundation, the HHMI-Simons Faculty Scholars Program, IARPA D16PC00008, the US Army Research Laboratory and the US Army Research Office under contract/grant number W911NF1510548, the US–Israel Binational Science Foundation (grant 2014509), the MIT Media Lab, the MIT Brain and Cognitive Sciences Department, and the McGovern Institute. A.T.W. acknowledges the Hertz Foundation Fellowship.
Author information
Authors and Affiliations
Contributions
A.T.W., Y.Z., and E.S.B. all contributed to the writing of the manuscript and have read and agreed to its content.
Corresponding author
Ethics declarations
Competing interests
E.S.B. is a co-inventor on multiple patents related to ExM and is also a co-founder of a company (http://extbio.com/) commercializing ExM. Y.Z. and A.T.W. are inventors on several inventions related to ExM.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Information
Supplementary Tables 1 and 2, and Supplementary Notes 1 and 2
Rights and permissions
About this article
Cite this article
Wassie, A.T., Zhao, Y. & Boyden, E.S. Expansion microscopy: principles and uses in biological research. Nat Methods 16, 33–41 (2019). https://doi.org/10.1038/s41592-018-0219-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-018-0219-4
This article is cited by
-
Control of polymers’ amorphous-crystalline transition enables miniaturization and multifunctional integration for hydrogel bioelectronics
Nature Communications (2024)
-
Morphological analysis of descending tracts in mouse spinal cord using tissue clearing, tissue expansion and tiling light sheet microscopy techniques
Scientific Reports (2023)
-
iU-ExM: nanoscopy of organelles and tissues with iterative ultrastructure expansion microscopy
Nature Communications (2023)
-
Light-sheets and smart microscopy, an exciting future is dawning
Communications Biology (2023)
-
Multielement Z-tag imaging by X-ray fluorescence microscopy for next-generation multiplex imaging
Nature Methods (2023)