Abstract
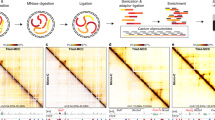
Long-range chromatin interactions play critical roles in genome organization and regulation of transcription. We now report transposase-mediated analysis of chromatin looping (Trac-looping) for simultaneous detection of multiscale genome-wide chromatin interactions among regulatory elements and chromatin accessibility. With this technique, a bivalent oligonucleotide linker is inserted between two interacting regions such that the chromatin interactions are captured without prior chromatin fragmentation and proximity-based ligation. Application of Trac-looping to human CD4+ T cells revealed substantial reorganization of enhancer–promoter interactions associated with changes in gene expression after T cell receptor stimulation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Vakoc, C. R. et al. Proximity among distant regulatory elements at the beta-globin locus requires GATA-1 and FOG-1. Mol. Cell 17, 453–462 (2005).
Zhang, Y. et al. Chromatin connectivity maps reveal dynamic promoter-enhancer long-range associations. Nature 504, 306–310 (2013).
Wei, Z. et al. Klf4 organizes long-range chromosomal interactions with the oct4 locus in reprogramming and pluripotency. Cell Stem Cell 13, 36–47 (2013).
Phillips-Cremins, J. E. et al. Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell 153, 1281–1295 (2013).
Tolhuis, B., Palstra, R. J., Splinter, E., Grosveld, F. & de Laat, W. Looping and interaction between hypersensitive sites in the active beta-globin locus. Mol. Cell 10, 1453–1465 (2002).
Palstra, R. J. et al. The beta-globin nuclear compartment in development and erythroid differentiation. Nat. Genet. 35, 190–194 (2003).
Drissen, R. et al. The active spatial organization of the beta-globin locus requires the transcription factor EKLF. Genes Dev. 18, 2485–2490 (2004).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Duan, Z. et al. A three-dimensional model of the yeast genome. Nature 465, 363–367 (2010).
Tanizawa, H. et al. Mapping of long-range associations throughout the fission yeast genome reveals global genome organization linked to transcriptional regulation. Nucleic Acids Res. 38, 8164–8177 (2010).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Mifsud, B. et al. Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C. Nat. Genet. 47, 598–606 (2015).
Davies, J. O. et al. Multiplexed analysis of chromosome conformation at vastly improved sensitivity. Nat. Methods 13, 74–80 (2016).
Chepelev, I., Wei, G., Wangsa, D., Tang, Q. & Zhao, K. Characterization of genome-wide enhancer-promoter interactions reveals co-expression of interacting genes and modes of higher order chromatin organization. Cell Res. 22, 490–503 (2012).
Tang, Z. et al. CTCF-mediated human 3D genome architecture reveals chromatin topology for transcription. Cell 163, 1611–1627 (2015).
Mumbach, M. R. et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat. Methods 13, 919–922 (2016).
Mumbach, M. R. et al. Enhancer connectome in primary human cells identifies target genes of disease-associated DNA elements. Nat. Genet. 49, 1602–1612 (2017).
Reznikoff, W. S. Tn5 as a model for understanding DNA transposition. Mol. Microbiol. 47, 1199–1206 (2003).
Reznikoff, W. S. The Tn5 transposon. Annu. Rev. Microbiol. 47, 945–963 (1993).
Adey, A. et al. Rapid, low-input, low-bias construction of shotgun fragment libraries by high-density in vitro transposition. Genome. Biol. 11, R119 (2010).
Gangadharan, S., Mularoni, L., Fain-Thornton, J., Wheelan, S. J. & Craig, N. L. DNA transposon Hermes inserts into DNA in nucleosome-free regions in vivo. Proc. Natl. Acad. Sci. USA 107, 21966–21972 (2010).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Cao, Q. et al. Reconstruction of enhancer-target networks in 935 samples of human primary cells, tissues and cell lines. Nat. Genet. 49, 1428–1436 (2017).
He, B., Chen, C., Teng, L. & Tan, K. Global view of enhancer-promoter interactome in human cells. Proc. Natl. Acad. Sci. USA 111, E2191–E2199 (2014).
Li, G. et al. Extensive promoter-centered chromatin interactions provide a topological basis for transcription regulation. Cell 148, 84–98 (2012).
Ma, W. et al. Fine-scale chromatin interaction maps reveal the cis-regulatory landscape of human lincRNA genes. Nat. Methods 12, 71–78 (2015).
Hughes, J. R. et al. Analysis of hundreds of cis-regulatory landscapes at high resolution in a single, high-throughput experiment. Nat. Genet. 46, 205–212 (2014).
Luger, K., Dechassa, M. L. & Tremethick, D. J. New insights into nucleosome and chromatin structure: an ordered state or a disordered affair? Nat. Rev. Mol. Cell Biol. 13, 436–447 (2012).
Hsieh, T. H. et al. Mapping nucleosome resolution chromosome folding in yeast by Micro-C. Cell 162, 108–119 (2015).
Risca, V. I., Denny, S. K., Straight, A. F. & Greenleaf, W. J. Variable chromatin structure revealed by in situ spatially correlated DNA cleavage mapping. Nature 541, 237–241 (2017).
Dorigo, B. et al. Nucleosome arrays reveal the two-start organization of the chromatin fiber. Science 306, 1571–1573 (2004).
Song, F. et al. Cryo-EM study of the chromatin fiber reveals a double helix twisted by tetranucleosomal units. Science 344, 376–380 (2014).
Hsieh, T. S., Fudenberg, G., Goloborodko, A. & Rando, O. J. Micro-C XL: assaying chromosome conformation from the nucleosome to the entire genome. Nat. Methods 13, 1009–1011 (2016).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Bonev, B. et al. Multiscale 3D genome rewiring during mouse neural development. Cell 171, 557–572 (2017).
Cremer, M. et al. Multicolor 3D fluorescence in situ hybridization for imaging interphase chromosomes. Methods Mol. Biol. 463, 205–239 (2008).
Javierre, B. M. et al. Lineage-specific genome architecture links enhancers and non-coding disease variants to target gene promoters. Cell 167, 1369–1384 (2016).
Kim, H. P., Imbert, J. & Leonard, W. J. Both integrated and differential regulation of components of the IL-2/IL-2 receptor system. Cytokine Growth Factor Rev. 17, 349–366 (2006).
Beagrie, R. A. et al. Complex multi-enhancer contacts captured by genome architecture mapping. Nature 543, 519–524 (2017).
Quinodoz, S. A. et al. Higher-order inter-chromosomal hubs shape 3D genome organization in the nucleus. Cell 174, 744–757 (2018).
Fang, R. et al. Mapping of long-range chromatin interactions by proximity ligation-assisted ChIP-seq. Cell Res. 26, 1345–1348 (2016).
Goryshin, I. Y. & Reznikoff, W. S. Tn5 in vitro transposition. J. Biol. Chem. 273, 7367–7374 (1998).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome. Biol. 9, R137 (2008).
Phanstiel, D. H., Boyle, A. P., Heidari, N. & Snyder, M. P. Mango: a bias-correcting ChIA-PET analysis pipeline. Bioinformatics 31, 3092–3098 (2015).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Wang, Z. et al. Combinatorial patterns of histone acetylations and methylations in the human genome. Nat. Genet. 40, 897–903 (2008).
Bernstein, B. E. et al. The NIH Roadmap Epigenomics Mapping Consortium. Nat. Biotechnol. 28, 1045–1048 (2010).
Li, W., Gong, K., Li, Q., Alber, F. & Zhou, X. J. Hi-Corrector: a fast, scalable and memory-efficient package for normalizing large-scale Hi-C data. Bioinformatics 31, 960–962 (2015).
Shin, H. et al. TopDom: an efficient and deterministic method for identifying topological domains in genomes. Nucleic Acids Res. 44, e70 (2016).
Sofueva, S. et al. Cohesin-mediated interactions organize chromosomal domain architecture. EMBO J. 32, 3119–3129 (2013).
Zheng, X. & Zheng, Y. CscoreTool: fast Hi-C compartment analysis at high resolution. Bioinformatics 34, 1568–1570 (2018).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Huang, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Zhao, K., Tang, Q. & Lai, B. Trac-looping protocol. Protoc. Exch. https://doi.org/10.1038/protex.2018.093 (2018).
Acknowledgements
We thank the NHLBI DNA Sequencing Core facility for sequencing the libraries; the NIH Biowulf High Performance Computing Systems for computing service; and the Department of Transfusion Medicine for preparing lymphocyte cells procured by apheresis from donor blood. We thank W.L. Ku for critical reading of the manuscript. The dCas9–KRAB plasmid was a gift from S. Qi (Department of Bioengineering, Department of Chemical and Systems Biology, Stanford University, Stanford, CA, USA). This work was supported by the Division of Intramural Research of NHLBI, NIH and CCR, NCI, NIH.
Author information
Authors and Affiliations
Contributions
K.Z conceived the project. J.S., T.R., and K.Z. directed the study. Q.T. performed the experiments. B.L., W.J., and G.H. did all data analysis. D.W. did the DNA-FISH experiments and analyzed the data. K.C., B.Z.S., G.R., Y.D., M.Z., and S.L. contributed to experimental design and data. B.L., W.J., G.H., and K.Z. wrote the paper with help from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Scheme of Trac-looping and chromatography analysis of the Tn5 complexes.
(a) DNA transposition mediated by Mosaic ends (ME) sequence and Tn5 transposase complex. (b) The monovalent ME linker and dimeric Tn5 complex used for DNA fragmentation. (c) The bivalent ME linker, monovalent ME adapter and Tnp form tetrameric Tn5 complex used for detection of chromatin looping. (d) Elution profiles of the Tn5 complexes from Superdex 200 column. Both the absorptions by protein (UV280, blue) and DNA (UV254, red) were monitored. The eluate fractions are indicated at the bottom of the plot. The size standards are indicated in the plots. (e) The A7 fraction contains both the bivalent and half linkers. The DNA was purified from the input (control), A7 and A11 fractions and was analyzed by 2% E-gel. Two repeated experiments were performed independently with similar results.
Supplementary Figure 2 Characterizing Trac-looping PETs.
(a) False-positive rate for H3K4me2 ChIA-PET, in situ Hi-C, and Trac-looping for naïve CD4+ T cells. Data are shown as mean ± s.d. of repeated experiments for each method (ChIA-PET, n=2; HiC, n=2; Trac-looping, n=4). The detail of the false-positive rate calculation was described in online Methods. (b) Fraction of intra-chromosomal PETs for H3K4me2 ChIA-PET, in situ Hi-C, and Trac-looping for naïve CD4+ T cells. Data are shown as mean ± s.d. of repeated experiments for each method (ChIA-PET, n=2; HiC, n=2; Trac-looping, n=4). (c) Fraction of intra-chromosomal PETs by Trac-looping in unfixed and fixed cells. Data are shown as mean ± s.d. of 4 repeated experiments respectively. (d) Fraction of intra-TAD PETs by Trac-looping in unfixed and fixed cells. Data are shown as mean ± s.d. of 4 repeated experiments respectively. (e) Fraction of super long PETs (>2M bp) detected by Trac-looping in unfixed and fixed cells. Data are shown as mean ± s.d. of 4 repeated experiments respectively.
Supplementary Figure 3 High consistency between Trac-looping and ATAC-seq in detection of chromatin accessibility.
(a) Distribution of PET-end density around transcription start sites (TSSs) for PETs sorted by distance. (b) Scatter plot showing tag density for ATAC-seq in fixed cells (x-axis) and unfixed cells (y-axis) at all accessible sites (n=64,316). (c) Scatter plot showing tag density for ATAC-seq and Trac-looping in fixed cells at all accessible sites (n=64,316). (d) Scatter plot showing tag density of ATAC-seq and Trac-looping in fixed cells at promoters (left; n=8,098), enhancers (middle; n=18,425) and CTCF sites (right; n=12,906). (b-d): The R value is the Pearson’ correlation coefficient.
Supplementary Figure 4 Trac-looping PETs reveal high order of chromatin organization.
(a) Heat map generated by Juicer-box (Durant et al. 2016) showing the contact matrix (resolution = 500K bp) on whole chromosome 4 for Trac-looping and in situ Hi-C in resting CD4+ T cells. (b) WashU genome browser showing the read density of PETs with distance less than 150 bp (corresponding to accessible region), interaction matrices from Trac-looping data and from in situ Hi-C data, and distributions of compartment scores and boundary scores overlaid between pooled Trac-looping data (red lines) and the in situ Hi-C data (blue lines) for a 20M bp genomic region in chromosome 19 of resting CD4+ T cells. The interaction matrices, compartment scores and boundary scores were visualized at a resolution of 20K bp. Only informative PETs longer than 200K bp were used to infer compartment scores: 10M in total from Trac-loop data sets and 76M in total from in situ Hi-C, explaining the more fluctuation on the curve estimated for Trac-looping than for in situ Hi-C. (c) Smoothed scatter plot of the compartment scores calculated from Trac-looping data and in situ Hi-C data for all 20K genomic bins (n=154,783) of resting CD4 T cells. The r value is the Pearson’s correlation coefficient. (d) Smoothed scatter plot of the boundary scores calculated from Trac-looping data and in situ Hi-C data for all 20K genomic bins (n=154,783) of resting CD4 T cells. The boundary score is calculated between two neighboring windows (m and n) of 200K bp, sliding along the genome at a step of 20K bps, as the ratio between interacting PETs within m or within n to the interacting PETs between m and n. The r value is the Pearson’s correlation coefficient. (e) Percentages of CTCF motif orientations at CTCF anchored chromatin loops predicted by Trac-looping as a function of distance.
Supplementary Figure 5 DNA-FISH validation of the Trac-looping results.
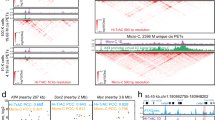
(a) The interaction landscapes and probe sets for 3D DNA-FISH experiments. Genome Browser display showing the chromatin interactions detected by Trac-looping in resting and activated T cells, respectively. The BAC clones used as probes in the DNA-FISH experiments are indicated below the panels: Clones RP11-133J18 and RP11-138B14 are used for positive region 2 on chr17 in both resting and stimulated CD4 T cell. The BAC clones RP11-138J18, RP11-12F14 and RP11-138J18, RP11-139I13 are used as probes for a negative control interaction. (b) The interaction landscapes and DNA-FISH probe set (RP11-101N10 and RP11-1130A7) for positive region 1 on chr5 in both resting and activated CD4+ T cell. (c) Summary of DNA-FISH validation results in resting (left panel) and activated (right panel) CD4 T cells. Y-axis shows the percentage of co-localization of the two pairs of interacting chromatin regions (positive) and one non-interacting pair of chromatin regions (negative control) measured by two sets of probes. The data shown represents 38, 39, and 59 measurements for two positive region and one negative region in resting T cells, and 38, 41, and 61 measurements for two positive region and one negative region in activated T cells. The center line indicates the mean value for the represented group; error bars indicate ±s.d. The P value was calculated by one-sided T test.
Supplementary Figure 6 Characterizing and validating interactions identified by Trac-looping.
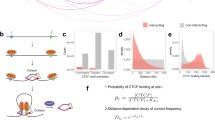
(a) Ratio of informative PETs associated with promoters of active genes to all informative PETs for data generated by the original Trac-looping (black bars), the PETs generated by the improved Trac-looping (red bars), and PETs from in-house Hi-C data and publicly available Hi-C data (blue bars). (b) Venn-diagram shows the comparison of called interactions from the number of PETs by Trac-looping and H3K4me2 ChIA-PET. (c) Cumulative distribution of number of predicted interaction by Trac-looping as a function of interaction distance. (d) Percentage of Trac-looping predicted interactions confirmed by alternative methods as a function of distance in resting CD4+ T cells. Alternative methods include PCHi-C (Javierre et al., 2016), H3K27ac Hi-ChIP (Mumbach et al., 2017), H3K4me2 CHIA-PET (Chepelev et al., 2012) and in situ Hi-C (Javierre et al., 2016). Predictions for Hi-ChIP and H3K4me2 ChIA-PET were made with Mango (bin size=2K, FDR<=0.05), the same method used for Trac-looping. Predictions for the Hi-C data were made with Homer (bin size=2K, FDR<=0.05). Predictions for PCHi-C are from published data. (e) Distribution of interactions predicted from 216M informative PETs, sorted by significance (FDR). (f) Saturation analysis shows interactions with high significance (FDR<1e-5) reached saturation by sequencing 60-80% of the total PETS, while interactions with low significance do not reach saturation at the current sequencing depth. (g) Percentage of functional interactions (enhancer-enhancer, enhancer-promoter and promoter-promoter) increases as the significance of the interactions increases.
Supplementary Figure 7 Associating Trac-looping interactions with chromatin states and gene expression in resting CD4+ T cells.
(a) Percentage of peaks involved in Trac-looping interactions for different histone modifications. (b) Percentage of in situ HiC-derived interactions anchored at different histone modification regions that were also detected by Trac-looping. (c) Heat map showing percentage of Trac-looping interactions anchored at different pairs of histone modification peaks. (d) Genomic distribution of Trac-looping interactions. (e) Distribution of interacting regions in active and poised promoters and enhancers associated with different histone modifications. (f) The expression level of a gene is positively correlated with the number of DHSs interacting with its promoter in resting CD4+ T cells. The genes are grouped according to the number of enhancer DHSs interacting with their promoters as detected by Trac-looping, with the group size n=19,416, 2,973, 1,731, 1,002, 623, 386, 222, 134, 424 for interaction number from 0 to ≥8. Center line, median; boxes, first and third quartiles; whiskers, 1.5× the interquartile range; notch, 95% confidence interval of the median. (g) Barplot showing the IL2RA expression measured by qRT-PCR and normalized to 18S RNA in transfected T cells. T cells were transfected with constructs expressing sgRNAs that target dCAS9-KRAB to the selected regions interacting with the IL2RA promoter. Three potential enhancers targeted by dCAS9-KRAB are indicated in Fig. 4a as sg1, 2 and 3. A sgRNA sequence from a different chromosome was used as a control. Data shown represents four independently repeated experiments (n=4). The P value was calculated by one-sided T test. (h) Interaction intensity of active and poised promoters and enhancers. Data shown represents 12,305 active promoters, 2,203 bivalent promoters, 23,524 active enhancers, and 49,760 primed enhancers. The center line, median; boxes, first and third quartiles; whiskers, 1.5× the interquartile range. (i) Identification of super enhancers in resting CD4+ T cells using H3K27ac ChIP-seq data. (j) Comparison of interaction intensity between regular and super enhancers. Data shown represents 24,040 HSs at regular enhancers and 8,344 HSs at super enhancers. The center line, median; boxes, first and third quartiles; whiskers, 1.5× the interquartile range. (k) Percentage of regular and super enhancers that interact with potential target promoters.
Supplementary Figure 8 Chromatin accessibility and proximity are not sufficient to allow interaction.
(a) The nearest DHS interacting with a promoter is identified and the number of DHSs that are looped out by the interaction are counted and shown in the table. (b) The nearest DHS that interacts with the reference DHS is identified and the number of DHSs that are looped out by the interaction are counted. (c) The fraction of enhancers that either interact with the nearest promoters or skip the nearest promoters. (d) Genome Browser image showing an enhancer that interacts with promoters other than the nearest promoter.
Supplementary Figure 9 Chromatin interaction changes are associated with gene expression changes and biological function of T cell activation.
(a) Scatter plot of the observed fold change of interacting PETs vs. fold change of accessibility at anchors for all the identified interactions. (b) Scatter plot of the normalized fold change of interacting PETs vs. fold change of accessibility at anchors for all the identified interactions. (c) Cumulative density of in situ Hi-C PETs density in resting (blue line) and stimulated T cell (red line) for loops (n=3,517) that showed an increase in interaction as predicted by Trac-looping. The P value was calculated by one-sided T test. (d) Cumulative density graph showing the expression levels of genes (n=2,122) with increased enhancer-promoter interactions in resting (black line) and activated (red line) T cells. A shift to right indicates increased expression in activated CD4+ T cells. The P value was calculated by one-sided T test. (e) GO analysis for genes (n=2,122) associated with an increase in interaction with HS(s). The FDR value calculation are performed by David Bioinformatics Resources (v6.8). (f) Cumulative density of in situ Hi-C PETs density in resting (blue line) and stimulated T cells (red line) for loops (n=748) that showed a decrease in interaction as predicted by Trac-looping. The P value was calculated by one-sided T test. (g) Cumulative density graph showing the expression levels of genes (n=725) with decreased enhancer-promoter interactions in resting (black line) and activated (red line) T cells. A shift to left indicates decreased expression in activated CD4+ T cells. The P value was calculated by one-sided T test. (h) GO analysis for genes (n=725) associated with a decrease in interaction with HS(s). The FDR value calculation are performed by David Bioinformatics Resources (v6.8).
Supplementary Figure 10 Detected enhancer switch after TCR stimulation of CD4+ T cells.
(a) Summary of changes of promoter-enhancer interactions for 7 different promoter groups during T cell activation. There are three kinds of interactions: no change, decreased, or increased after stimulation of CD4+ T cells. Each promoter may have multiple enhancer-promoter interactions, and thus may have more than one type of interactions. There are total 7 groups of promoters according to the types of interactions. For example, the 7th group of promoters has all three kinds of interactions. We annotated the 6th and the 7th groups as having the switched enhancer because some linked enhancer interaction were decreased while others were increased. (b) Genome Browser images showing examples of enhancer switch during T cell activation.
Supplementary Figure 11 Characterizing interaction networks during T cell stimulation.
(a) Most interaction networks are formed within TADs. Shown are two interaction networks localized to TAD113 and TAD114, respectively. (b) Examples of decreased and increased connectivity in interaction networks upon T cell activation. Red circle denotes promoters and green and blue circles denote potential enhancers. (c) The cumulative density graph of gene expression associated with 425 reduced interaction networks in resting T cells (red) and activated T cells (blue) is shown on the left and the box plot of gene expression is shown on the right. (d) The cumulative density graph of gene expression associated with 304 increased network in resting T cells (red) and activated T cells (blue) is shown on the left and the box plot of gene expression is shown on the right. (c-d) The P value was calculated by one-sided T test. For boxplot: center line, median; boxes, first and third quartiles; whiskers, 1.5× the interquartile range. (e) The number of promoters interacting with one promoter in resting and activated CD4+ T cells. (f) Genome Browser image showing the promoters that interact with the VTRNA1-2 in resting CD4 T cells. (g) Genome Browser image showing the promoters that interact with the C12orf57 promoter in activated CD4+ T cells.
Supplementary Figure 12 Trac-looping data predict transcription factors involved in the regulation of interaction networks.
(a) The top motif enriched in the increased interacting sites belongs to the FOS family of transcription factors. The numbers in brackets indicate the fold of either down-regulation or up-regulation by T cell activation as measured by RNA-seq. (b) The DNA surrounding the FOS motifs in the regions with increased interactions exhibits footprints in stimulated CD4+ T cell. All sequencing reads including both short PETs and long PETs from the Trac-looping libraries were pooled for this analysis. Blue and red lines represent reads from resting and stimulated T cell, respectively. (c) FOSL1 binding is enriched in the chromatin regions with increased interactions in stimulated CD4+ T cells, while no enrichment is detected in resting CD4+ T cells. ChIP-Seq assays were performed using chromatin from both resting and stimulated CD4+ T cells and the average profile of FOSL1 binding was plotted surrounding the center of chromatin regions with increased interactions. (d) Heatmaps of FOSL1 density at identified FOSL1 motif in interaction-increased loci in both resting and stimulated CD4+ T cells. These identified FOSL1 motifs were sorted based on their similarity to FOSL1 motif from JASPAR (MA0477.1). (e) Genome browser snapshot of an interaction-increased locus, which contains a potential enhancer of the BCAT1 gene. FOSL1 binding was detected after CD4+ T cell stimulation.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12, Supplementary Tables 1 and 2
Rights and permissions
About this article
Cite this article
Lai, B., Tang, Q., Jin, W. et al. Trac-looping measures genome structure and chromatin accessibility. Nat Methods 15, 741–747 (2018). https://doi.org/10.1038/s41592-018-0107-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-018-0107-y
This article is cited by
-
Comparative study on chromatin loop callers using Hi-C data reveals their effectiveness
BMC Bioinformatics (2024)
-
3C methods in cancer research: recent advances and future prospects
Experimental & Molecular Medicine (2024)
-
ChIATAC is an efficient strategy for multi-omics mapping of 3D epigenomes from low-cell inputs
Nature Communications (2023)
-
Three-dimensional genome organization in immune cell fate and function
Nature Reviews Immunology (2023)
-
Simultaneous profiling of chromatin architecture and transcription in single cells
Nature Structural & Molecular Biology (2023)