Abstract
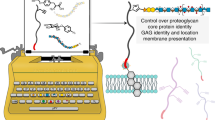
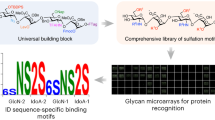
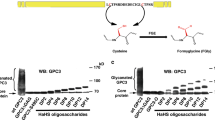
Glycosaminoglycans (GAGs) are essential polysaccharides in normal physiology and disease. However, understanding of the contribution of specific GAG structures to specific biological functions is limited, largely because of the great structural heterogeneity among GAGs themselves, as well as technical limitations in the structural characterization and chemical synthesis of GAGs. Here we describe a cell-based method to produce and display distinct GAGs with a broad repertoire of modifications, a library we refer to as the GAGOme. By using precise gene editing, we engineered a large panel of Chinese hamster ovary cells with knockout or knock-in of the genes encoding most of the enzymes involved in GAG biosynthesis, to generate a library of isogenic cell lines that differentially display distinct GAG features. We show that this library can be used for cell-based binding assays, recombinant expression of proteoglycans with distinct GAG structures, and production of distinct GAG chains on metabolic primers that may be used for the assembly of GAG glycan microarrays.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Esko, J. D. & Selleck, S. B. Order out of chaos: assembly of ligand binding sites in heparan sulfate. Annu. Rev. Biochem. 71, 435–471 (2002).
Mizumoto, S., Yamada, S. & Sugahara, K. Molecular interactions between chondroitin-dermatan sulfate and growth factors/receptors/matrix proteins. Curr. Opin. Struct. Biol. 34, 35–42 (2015).
Xu, D. & Esko, J. D. Demystifying heparan sulfate-protein interactions. Annu. Rev. Biochem. 83, 129–157 (2014).
Mizumoto, S., Yamada, S. & Sugahara, K. Human genetic disorders and knockout mice deficient in glycosaminoglycan. Biomed. Res. Int. 2014, 495764 (2014).
Gama, C. I. et al. Sulfation patterns of glycosaminoglycans encode molecular recognition and activity. Nat. Chem. Biol. 2, 467–473 (2006).
Fu, L., Suflita, M. & Linhardt, R. J. Bioengineered heparins and heparan sulfates. Adv. Drug Deliv. Rev. 97, 237–249 (2016).
Esko, J. D., Stewart, T. E. & Taylor, W. H. Animal cell mutants defective in glycosaminoglycan biosynthesis. Proc. Natl. Acad. Sci. USA 82, 3197–3201 (1985).
Zhang, L., Lawrence, R., Frazier, B. A. & Esko, J. D. CHO glycosylation mutants: proteoglycans. Methods Enzymol. 416, 205–221 (2006).
Okayama, M., Kimata, K. & Suzuki, S. The influence of p-nitrophenyl β-d-xyloside on the synthesis of proteochondroitin sulfate by slices of embryonic chick cartilage. J. Biochem. 74, 1069–1073 (1973).
Li, G. et al. Glycosaminoglycanomics of cultured cells using a rapid and sensitive LC-MS/MS approach. ACS Chem. Biol. 10, 1303–1310 (2015).
Yang, Z. et al. Engineered CHO cells for production of diverse, homogeneous glycoproteins. Nat. Biotechnol. 33, 842–844 (2015).
Koike, T., Izumikawa, T., Tamura, J. & Kitagawa, H. FAM20B is a kinase that phosphorylates xylose in the glycosaminoglycan-protein linkage region. Biochem. J. 421, 157–162 (2009).
Koike, T., Izumikawa, T., Sato, B. & Kitagawa, H. Identification of phosphatase that dephosphorylates xylose in the glycosaminoglycan-protein linkage region of proteoglycans. J. Biol. Chem. 289, 6695–6708 (2014).
Sugahara, K., Yamashina, I., De Waard, P., Van Halbeek, H. & Vliegenthart, J. F. Structural studies on sulfated glycopeptides from the carbohydrate-protein linkage region of chondroitin 4-sulfate proteoglycans of swarm rat chondrosarcoma. Demonstration of the structure Gal(4-O-sulfate)β-1-3Gal-β-1-4XYL-β-1-O-Ser. J. Biol. Chem. 263, 10168–10174 (1988).
Ueno, M., Yamada, S., Zako, M., Bernfield, M. & Sugahara, K. Structural characterization of heparan sulfate and chondroitin sulfate of syndecan-1 purified from normal murine mammary gland epithelial cells. Common phosphorylation of xylose and differential sulfation of galactose in the protein linkage region tetrasaccharide sequence. J. Biol. Chem. 276, 29134–29140 (2001).
de Waard, P., Vliegenthart, J. F., Harada, T. & de Sugahara, K. Structural studies on sulfated oligosaccharides derived from the carbohydrate-protein linkage region of chondroitin 6-sulfate proteoglycans of shark cartilage. II. Seven compounds containing 2 or 3 sulfate residues. J. Biol. Chem. 267, 6036–6043 (1992).
Kitagawa, H. et al. Sulfation of the galactose residues in the glycosaminoglycan-protein linkage region by recombinant human chondroitin 6-O-sulfotransferase 1. J. Biol. Chem. 283, 27438–27443 (2008).
Wen, J. et al. Xylose phosphorylation functions as a molecular switch to regulate proteoglycan biosynthesis. Proc. Natl. Acad. Sci. USA 111, 15723–15728 (2014).
Tone, Y. et al. 2-O-phosphorylation of xylose and 6-O-sulfation of galactose in the protein linkage region of glycosaminoglycans influence the glucuronyltransferase-I activity involved in the linkage region synthesis. J. Biol. Chem. 283, 16801–16807 (2008).
Uyama, T., Kitagawa, H., Tamura Ji, J. & Sugahara, K. Molecular cloning and expression of human chondroitin N-acetylgalactosaminyltransferase: the key enzyme for chain initiation and elongation of chondroitin/dermatan sulfate on the protein linkage region tetrasaccharide shared by heparin/heparan sulfate. J. Biol. Chem. 277, 8841–8846 (2002).
Uyama, T. et al. Molecular cloning and expression of a second chondroitin N-acetylgalactosaminyltransferase involved in the initiation and elongation of chondroitin/dermatan sulfate. J. Biol. Chem. 278, 3072–3078 (2003).
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P. M. & Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 42, D490–D495 (2014).
Izumikawa, T., Uyama, T., Okuura, Y., Sugahara, K. & Kitagawa, H. Involvement of chondroitin sulfate synthase-3 (chondroitin synthase-2) in chondroitin polymerization through its interaction with chondroitin synthase-1 or chondroitin-polymerizing factor. Biochem. J. 403, 545–552 (2007).
Izumikawa, T. et al. Identification of chondroitin sulfate glucuronyltransferase as chondroitin synthase-3 involved in chondroitin polymerization: chondroitin polymerization is achieved by multiple enzyme complexes consisting of chondroitin synthase family members. J. Biol. Chem. 283, 11396–11406 (2008).
Kitagawa, H., Shimakawa, H. & Sugahara, K. The tumor suppressor EXT-like gene EXTL2 encodes an α1,4-N-acetylhexosaminyltransferase that transfers N-acetylgalactosamine and N-acetylglucosamine to the common glycosaminoglycan-protein linkage region. The key enzyme for the chain initiation of heparan sulfate. J. Biol. Chem. 274, 13933–13937 (1999).
Kim, B. T. et al. Human tumor suppressor EXT gene family members EXTL1 and EXTL3 encode α1,4- N-acetylglucosaminyltransferases that likely are involved in heparan sulfate/ heparin biosynthesis. Proc. Natl. Acad. Sci. USA 98, 7176–7181 (2001).
Lind, T., Tufaro, F., McCormick, C., Lindahl, U. & Lidholt, K. The putative tumor suppressors EXT1 and EXT2 are glycosyltransferases required for the biosynthesis of heparan sulfate. J. Biol. Chem. 273, 26265–26268 (1998).
McCormick, C., Duncan, G., Goutsos, K. T. & Tufaro, F. The putative tumor suppressors EXT1 and EXT2 form a stable complex that accumulates in the Golgi apparatus and catalyzes the synthesis of heparan sulfate. Proc. Natl. Acad. Sci. USA 97, 668–673 (2000).
Pacheco, B., Malmström, A. & Maccarana, M. Two dermatan sulfate epimerases form iduronic acid domains in dermatan sulfate. J. Biol. Chem. 284, 9788–9795 (2009).
Maccarana, M. et al. Biosynthesis of dermatan sulfate: chondroitin-glucuronate C5-epimerase is identical to SART2. J. Biol. Chem. 281, 11560–11568 (2006).
Pinhal, M. A. et al. Enzyme interactions in heparan sulfate biosynthesis: uronosyl 5-epimerase and 2-O-sulfotransferase interact in vivo. Proc. Natl. Acad. Sci. USA 98, 12984–12989 (2001).
Salanti, A. et al. Targeting human cancer by a glycosaminoglycan binding malaria protein. Cancer Cell 28, 500–514 (2015).
Turnbull, J. E., Fernig, D. G., Ke, Y., Wilkinson, M. C. & Gallagher, J. T. Identification of the basic fibroblast growth factor binding sequence in fibroblast heparan sulfate. J. Biol. Chem. 267, 10337–10341 (1992).
Pye, D. A., Vives, R. R., Turnbull, J. E., Hyde, P. & Gallagher, J. T. Heparan sulfate oligosaccharides require 6-O-sulfation for promotion of basic fibroblast growth factor mitogenic activity. J. Biol. Chem. 273, 22936–22942 (1998).
Avnur, Z. & Geiger, B. Immunocytochemical localization of native chondroitin sulfate in tissues and cultured cells using specific monoclonal antibody. Cell 38, 811–822 (1984).
Ito, Y. et al. Structural characterization of the epitopes of the monoclonal antibodies 473HD, CS-56, and MO-225 specific for chondroitin sulfate D-type using the oligosaccharide library. Glycobiology 15, 593–603 (2005).
Persson, A., Ellervik, U. & Mani, K. Fine-tuning the structure of glycosaminoglycans in living cells using xylosides. Glycobiology 28, 499–511 (2018).
Esko, J. D. et al. Inhibition of chondroitin and heparan sulfate biosynthesis in Chinese hamster ovary cell mutants defective in galactosyltransferase I. J. Biol. Chem. 262, 12189–12195 (1987).
Kolset, S. O. & Tveit, H. Serglycin—structure and biology. Cell. Mol. Life Sci. 65, 1073–1085 (2008).
Datta, P. et al. Bioengineered Chinese hamster ovary cells with Golgi-targeted 3-O-sulfotransferase-1 biosynthesize heparan sulfate with an antithrombin-binding site. J. Biol. Chem. 288, 37308–37318 (2013).
Blixt, O. et al. Printed covalent glycan array for ligand profiling of diverse glycan binding proteins. Proc. Natl. Acad. Sci. USA 101, 17033–17038 (2004).
Stopschinski, B. E. et al. Specific glycosaminoglycan chain length and sulfation patterns are required for cell uptake of tau versus α-synuclein and β-amyloid aggregates. J. Biol. Chem. 293, 10826–10840 (2018).
Baik, J. Y. et al. Metabolic engineering of Chinese hamster ovary cells: towards a bioengineered heparin. Metab. Eng. 14, 81–90 (2012).
Paulson, J. C., Blixt, O. & Collins, B. E. Sweet spots in functional glycomics. Nat. Chem. Biol. 2, 238–248 (2006).
Song, X., Heimburg-Molinaro, J., Cummings, R. D. & Smith, D. F. Chemistry of natural glycan microarrays. Curr. Opin. Chem. Biol. 18, 70–77 (2014).
Rogers, C. J. et al. Elucidating glycosaminoglycan–protein–protein interactions using carbohydrate microarray and computational approaches. Proc. Natl. Acad. Sci. USA 108, 9747–9752 (2011).
Shipp, E. L. & Hsieh-Wilson, L. C. Profiling the sulfation specificities of glycosaminoglycan interactions with growth factors and chemotactic proteins using microarrays. Chem. Biol. 14, 195–208 (2007).
Geissner, A. & Seeberger, P. H. Glycan arrays: from basic biochemical research to bioanalytical and biomedical applications. Annu. Rev. Anal. Chem. 9, 223–247 (2016).
Kudelka, M. R. et al. Cellular O-glycome reporter/amplification to explore O-glycans of living cells. Nat. Methods 13, 81–86 (2016).
Varki, A. et al. Symbol nomenclature for graphical representations of glycans. Glycobiology 25, 1323–1324 (2015).
Narimatsu, Y. et al. A validated gRNA library for CRISPR–Cas9 targeting of the human glycosyltransferase genome. Glycobiology 28, 295–305 (2018).
Lonowski, L. A. et al. Genome editing using FACS enrichment of nuclease-expressing cells and indel detection by amplicon analysis. Nat. Protoc. 12, 581–603 (2017).
Yang, Z. et al. Fast and sensitive detection of indels induced by precise gene targeting. Nucleic Acids Res. 43, e59 (2015).
Bahr, S., Cortner, L., Ladley, S. & Borgschulte, T. Evaluating the effect of chromosomal context on zinc finger nuclease efficiency. BMC Proc. 7, 3 (2013).
Maresca, M., Lin, V. G., Guo, N. & Yang, Y. Obligate ligation-gated recombination (ObLiGaRe): custom-designed nuclease-mediated targeted integration through nonhomologous end joining. Genome Res. 23, 539–546 (2013).
Volpi, N., Galeotti, F., Yang, B. & Linhardt, R. J. Analysis of glycosaminoglycan-derived, precolumn, 2-aminoacridone-labeled disaccharides with LC-fluorescence and LC-MS detection. Nat. Protoc. 9, 541–558 (2014).
Lawrence, R., Lu, H., Rosenberg, R. D., Esko, J. D. & Zhang, L. Disaccharide structure code for the easy representation of constituent oligosaccharides from glycosaminoglycans. Nat. Methods 5, 291–292 (2008).
Czajkowsky, D. M., Hu, J., Shao, Z. & Pleass, R. J. Fc-fusion proteins: new developments and future perspectives. EMBO Mol. Med. 4, 1015–1028 (2012).
Acknowledgements
This work was supported by the Lundbeck Foundation, Læge Sofus Carl Emil Friis og hustru Olga Doris Friis’ Legat, the Novo Nordisk Foundation, Lund University, Kirsten og Freddy Johansen Fonden, the European Commission (GlycoImaging H2020-MSCA-ITN-721297; BioCapture H2020-MSCA-ITN-722171), the UCPH Excellence Programme for Interdisciplinary Research (CDO2016), and the Danish National Research Foundation (DNRF107) (all to H.C.). C.G was supported through FCT, POPH (Programa Operacional Potencial Humano) SFRH/BPD/96510/2013. A.S., T.M.C., and T.G. were supported through the ERC MalOnco Program, the Danish Cancer Society, and the Lundbeck Foundation.
Author information
Authors and Affiliations
Contributions
Y.-H.C., Y.M., Y.N., T.M.C., H.C., and Z.Y. conceived and designed the study; Y.-H.C., Y.M., Y.N., C.G., R.K., C.S., A.M., E.P.B., and Z.Y. contributed with experimental data and data interpretation; T.M.C., C.B.S., T.G., and A.S. contributed to the VAR2CSA studies; A.P., D.W., and U.E. contributed to the xyloside-priming studies; Y.-H.C., Y.M., H.C., and Z.Y. wrote the manuscript; and all authors edited and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The University of Copenhagen has filed patent application EP/2017/061385 on the basis of this work. Y.N., C.S., E.P.B., H.C., and Y.Z. are named inventors on the PCT application.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1–9 and Supplementary Note 1
Rights and permissions
About this article
Cite this article
Chen, YH., Narimatsu, Y., Clausen, T.M. et al. The GAGOme: a cell-based library of displayed glycosaminoglycans. Nat Methods 15, 881–888 (2018). https://doi.org/10.1038/s41592-018-0086-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-018-0086-z
This article is cited by
-
Structure and function of Semaphorin-5A glycosaminoglycan interactions
Nature Communications (2024)
-
A family of di-glutamate mucin-degrading enzymes that bridges glycan hydrolases and peptidases
Nature Catalysis (2024)
-
Molecular mechanism of decision-making in glycosaminoglycan biosynthesis
Nature Communications (2023)
-
Glycoengineered keratinocyte library reveals essential functions of specific glycans for all stages of HSV-1 infection
Nature Communications (2023)
-
Identification of global inhibitors of cellular glycosylation
Nature Communications (2023)