Abstract
The nucleosome acidic patch is a major interaction hub for chromatin, providing a platform for enzymes to dock and orient for nucleosome-targeted activities. To define the molecular basis of acidic patch recognition proteome wide, we performed an amino acid resolution acidic patch interactome screen. We discovered that the histone H3 lysine 36 (H3K36) demethylase KDM2A, but not its closely related paralog, KDM2B, requires the acidic patch for nucleosome binding. Despite fundamental roles in transcriptional repression in health and disease, the molecular mechanisms governing nucleosome substrate specificity of KDM2A/B, or any related JumonjiC (JmjC) domain lysine demethylase, remain unclear. We used a covalent conjugate between H3K36 and a demethylase inhibitor to solve cryogenic electron microscopy structures of KDM2A and KDM2B trapped in action on a nucleosome substrate. Our structures show that KDM2–nucleosome binding is paralog specific and facilitated by dynamic nucleosomal DNA unwrapping and histone charge shielding that mobilize the H3K36 sequence for demethylation.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD032791. Cryo-EM maps have been deposited in the Electron Microscopy Data Bank under accession codes EMD-26809 (KDM2A:H3K36C-UNC8015 nucleosome complex) and EMD-26810 (KDM2B:H3K36C-UNC8015 nucleosome complex). Atomic coordinates of the KDM2A:H3K36C-UNC8015 nucleosome and KMD2A:H3(29-41)K36C-UNC8015 complexes have been deposited in the Protein Data Bank with accession codes 7UV9 and 7UVA, respectively. Homology model of KDM2B:H3K36C-UNC8015 nucleosome is included as Supplementary File 3. Other structures used in this study are available through the Protein Data Bank with accession codes 4QX7, 6ESF, 6NZO, 7CRR and 7JO9. All materials in this study are available upon reasonable request. Source data are provided with this paper.
Code availability
In-house R script used for proteomics data analysis is available for download at GitHub (https://github.com/GoldfarbLab/NucleosomeProteomeAnalysis).
References
Hyun, K., Jeon, J., Park, K. & Kim, J. Writing, erasing and reading histone lysine methylations. Exp. Mol. Med. 49, e324 (2017).
Tsukada, Y.-I. et al. Histone demethylation by a family of JmjC domain-containing proteins. Nature 439, 811–816 (2006).
Shi, Y. et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119, 941–953 (2004).
Klose, R. J. & Zhang, Y. Regulation of histone methylation by demethylimination and demethylation. Nat. Rev. Mol. Cell Biol. 8, 307–318 (2007).
Shi, Y. & Whetstine, J. R. Dynamic regulation of histone lysine methylation by demethylases. Mol. Cell 25, 1–14 (2007).
Black, J. C., Van Rechem, C. & Whetstine, J. R. Histone lysine methylation dynamics: establishment, regulation, and biological impact. Mol. Cell 48, 491–507 (2012).
Punnia-Moorthy, G., Hersey, P., Emran, A. A. & Tiffen, J. Lysine demethylases: promising drug targets in melanoma and other cancers. Front. Genet. 12, 680633 (2021).
Borgel, J. et al. KDM2A integrates DNA and histone modification signals through a CXXC/PHD module and direct interaction with HP1. Nucleic Acids Res. 45, 1114–1129 (2017).
Frescas, D. et al. KDM2A represses transcription of centromeric satellite repeats and maintains the heterochromatic state. Cell Cycle 7, 3539–3547 (2008).
Bartke, T. et al. Nucleosome-interacting proteins regulated by DNA and histone methylation. Cell 143, 470–484 (2010).
Gao, Z. et al. PCGF homologs, CBX proteins, and RYBP define functionally distinct PRC1 family complexes. Mol. Cell 45, 344–356 (2012).
Gearhart, M. D., Corcoran, C. M., Wamstad, J. A. & Bardwell, V. J. Polycomb group and SCF ubiquitin ligases are found in a novel BCOR complex that is recruited to BCL6 targets. Mol. Cell. Biol. 26, 6880–6889 (2006).
Lagarou, A. et al. dKDM2 couples histone H2A ubiquitylation to histone H3 demethylation during Polycomb group silencing. Genes Dev. 22, 2799–2810 (2008).
Sánchez, C. et al. Proteomics analysis of Ring1B/Rnf2 interactors identifies a novel complex with the Fbxl10/Jhdm1B histone demethylase and the Bcl6 interacting corepressor. Mol. Cell. Proteom. 6, 820–834 (2007).
Wu, X., Johansen, J. V. & Helin, K. Fbxl10/Kdm2b recruits polycomb repressive complex 1 to CpG islands and regulates H2A ubiquitylation. Mol. Cell 49, 1134–1146 (2013).
Farcas, A. M. et al. KDM2B links the Polycomb Repressive Complex 1 (PRC1) to recognition of CpG islands. eLife 1, e00205–e00205 (2012).
Blackledge, N. P. et al. CpG islands recruit a histone H3 lysine 36 demethylase. Mol. Cell 38, 179–190 (2010).
Vacík, T., Lađinović, D. & Raška, I. KDM2A/B lysine demethylases and their alternative isoforms in development and disease. Nucleus 9, 431–441 (2018).
Yan, M., Yang, X., Wang, H. & Shao, Q. The critical role of histone lysine demethylase KDM2B in cancer. Am. J. Transl. Res 10, 2222–2233 (2018).
Cheng, Z. et al. A molecular threading mechanism underlies Jumonji lysine demethylase KDM2A regulation of methylated H3K36. Genes Dev. 28, 1758–1771 (2014).
Skrajna, A. et al. Comprehensive nucleosome interactome screen establishes fundamental principles of nucleosome binding. Nucleic Acids Res. 48, 9415–9432 (2020).
Nacev, B. A. et al. The expanding landscape of ‘oncohistone’ mutations in human cancers. Nature 567, 473–478 (2019).
McGinty, R. K. & Tan, S. Principles of nucleosome recognition by chromatin factors and enzymes. Curr. Opin. Struct. Biol. 71, 16–26 (2021).
Bennett, R. L. et al. A mutation in histone H2B represents a new class of oncogenic driver. Cancer Disco. 9, 1438–1451 (2019).
Wan, Y. C. E. et al. Cancer-associated histone mutation H2BG53D disrupts DNA–histone octamer interaction and promotes oncogenic phenotypes. Signal Transduct. Target Ther. 5, 27–4 (2020).
Arimura, Y. et al. Cancer-associated mutations of histones H2B, H3.1 and H2A.Z.1 affect the structure and stability of the nucleosome. Nucleic Acids Res. 46, 10007–10018 (2018).
Weinberg, D. N., Allis, C. D. & Lu, C. Oncogenic mechanisms of histone H3 mutations. Cold Spring Harb. Perspect. Med. 7, a026443 (2017).
Zhou, J. C., Blackledge, N. P., Farcas, A. M. & Klose, R. J. Recognition of CpG island chromatin by KDM2A requires direct and specific interaction with linker DNA. Mol. Cell. Biol. 32, 479–489 (2012).
Hamada, S. et al. Design, synthesis, enzyme-inhibitory activity, and effect on human cancer cells of a novel series of jumonji domain-containing protein 2 histone demethylase inhibitors. J. Med. Chem. 53, 5629–5638 (2010).
Suzuki, T. et al. Identification of the KDM2/7 histone lysine demethylase subfamily inhibitor and its antiproliferative activity. J. Med. Chem. 56, 7222–7231 (2013).
Luo, X. et al. A selective inhibitor and probe of the cellular functions of Jumonji C domain-containing histone demethylases. J. Am. Chem. Soc. 133, 9451–9456 (2011).
Wesley, N. A. et al. Time resolved-fluorescence resonance energy transfer platform for quantitative nucleosome binding and footprinting. Protein Sci. 31, e4339 (2022).
Li, W. et al. Molecular basis of nucleosomal H3K36 methylation by NSD methyltransferases. Nature 590, 498–503 (2021).
Bilokapic, S. & Halic, M. Nucleosome and ubiquitin position Set2 to methylate H3K36. Nat. Commun. 10, 3795 (2019).
Bagert, J. D. et al. Oncohistone mutations enhance chromatin remodeling and alter cell fates. Nat. Chem. Biol. 17, 403–411 (2021).
Spangler, C. J. et al. DOT1L activity in leukemia cells requires interaction with ubiquitylated H2B that promotes productive nucleosome binding. Cell Rep. 38, 110369 (2022).
Anderson, C. J. et al. Structural basis for recognition of ubiquitylated nucleosome by Dot1L methyltransferase. Cell Rep. 26, 1681–1690 (2019).
Tan, S., Kern, R. C. & Selleck, W. The pST44 polycistronic expression system for producing protein complexes in Escherichia coli. Protein Expr. Purif. 40, 385–395 (2005).
Weissmann, F. et al. biGBac enables rapid gene assembly for the expression of large multisubunit protein complexes. Proc. Natl Acad. Sci. USA 113, E2564–E2569 (2016).
Lowary, P. T. & Widom, J. New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning. J. Mol. Biol. 276, 19–42 (1998).
Kim, S.-A., Chatterjee, N., Jennings, M. J., Bartholomew, B. & Tan, S. Extranucleosomal DNA enhances the activity of the LSD1/CoREST histone demethylase complex. Nucleic Acids Res. 43, 4868–4880 (2015).
Makde, R. D., England, J. R., Yennawar, H. P. & Tan, S. Structure of RCC1 chromatin factor bound to the nucleosome core particle. Nature 467, 562–566 (2010).
Luger, K., Rechsteiner, T. J. & Richmond, T. J. Preparation of nucleosome core particle from recombinant histones. Methods Enzymol. 304, 3–19 (1999).
Luger, K., Rechsteiner, T. J., Flaus, A. J., Waye, M. M. & Richmond, T. J. Characterization of nucleosome core particles containing histone proteins made in bacteria. J. Mol. Biol. 272, 301–311 (1997).
Dignam, J. D., Lebovitz, R. M. & Roeder, R. G. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res. 11, 1475–1489 (1983).
Pieters, B. J. G. E. et al. Installation of trimethyllysine analogs on intact histones via cysteine alkylation. Bioconjug. Chem. 30, 952–958 (2019).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Evans, P. R. An introduction to data reduction: space-group determination, scaling and intensity statistics. Acta Crystallogr. D Biol. Crystallogr. 67, 282–292 (2011).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Zivanov, J., Nakane, T. & Scheres, S. H. W. Estimation of high-order aberrations and anisotropic magnification from cryo-EM data sets in RELION-3.1. IUCrJ 7, 253–267 (2020).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Zhong, E. D., Bepler, T., Berger, B. & Davis, J. H. CryoDRGN: reconstruction of heterogeneous cryo-EM structures using neural networks. Nat. Methods 18, 176–185 (2021).
Afonine, P. V. et al. New tools for the analysis and validation of cryo-EM maps and atomic models. Acta Crystallogr D Struct. Biol. 74, 814–840 (2018).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Acknowledgements
Single-particle cryo-EM data were collected with assistance from J. Peck and J. Strauss at the University of North Carolina at Chapel Hill School of Medicine Cryo-Electron Microscopy Facility, which is partially supported by National Institutes of Health (NIH) grant P30CA016086 and is part of the Molecular Microscopy Consortium of the University of North Carolina at Chapel Hill, Duke University and the National Institute of Environmental Health Sciences. Crystallography data were collected at the Southeast Regional Collaborative Access Team (SER-CAT) 22-ID beamline at the Advanced Photon Source, Argonne National Laboratory. SER-CAT is supported by its member institutions and NIH equipment grants S10_RR25528, S10_RR028976 and S10_OD027000. Plasmids containing 601 nucleosome positioning DNA fragments and NSD2 were gifts from S. Tan. We thank members of the McGinty, Goldfarb, Frye and James laboratories for critical comments on the manuscript. This work was supported by NIH grants R35GM133498 to R.K.M., F99CA253730 to C.J.S., R35GM139514 to S.V.F., R01CA010305 to L.I.J. and R01GM132299 to D.K.; American Cancer Society grant 132609-PF-18-153-01-DMC to A.S.; and National Science Foundation grant DGE-1650116 to N.A.W.
Author information
Authors and Affiliations
Contributions
R.K.M., D.G., S.V.F. and L.I.J. supervised the studies. C.J.S., A.S. and R.K.M. conceived the study and designed the experiments. C.J.S. prepared samples, grids, collected and processed the cryo-EM data and performed in vitro functional assays. A.S. prepared samples and performed proteomics experiments and in vitro functional assays. C.A.F. helped design and prepared cofactor analogs. A.N. and D.G. processed proteomics data. G.R.B. and D.N.A. prepared samples and solved the crystal structure. E.C.A., H.C.S., C.B.S. and J.-M.E.M. prepared samples. E.M.W. collected proteomics data. N.A.W. developed the nucleosome binding assay and performed nucleosome interaction studies. D.K. modeled substrate analogs. C.J.S., A.S. and R.K.M. wrote the manuscript, with contributions from all coauthors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Cheryl Arrowsmith and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
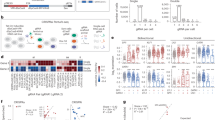
Extended Data Fig. 1 Amino acid resolution nucleosome affinity proteomics screen.
a, Schematic of nucleosome affinity proteomics screen. b, Coomassie-stained SDS-polyacrylamide denaturing gel (top) and ethidium bromide-stained native gel (bottom) of biotinylated nucleosome library. These experiments were repeated independently two times with similar results. c, Triplicate pulldowns run briefly on SDS polyacrylamide gels and stained with Coomassie blue prior to excision and in-gel trypsin digestion. Unused replicates of wild-type nucleosome pulldowns marked with an asterisk. d, Western blots following pulldowns from HEK293T nuclear lysates using indicated biotinylated nucleosomes. These experiments were performed once. WT, wild-type.
Extended Data Fig. 2 Oncohistone mutations have distinct effects on nucleosome binding.
a, Full heat map showing log2-fold change relative to wild-type nucleosome of quantified proteins listed at right across screen, clustered by protein and nucleosome mutant. b, Heat map of oncohistone mutant nucleosomes, clustered as in a. c, Heat map of each oncohistone mutation relative to the corresponding alanine mutation, clustered as in a.
Extended Data Fig. 3 KDM2A/B-nucleosome complex purification and 2D classification.
a, Chromatograms displaying normalized 280 nm absorbance from size exclusion purification of KDM2B/WT nucleosome (left), KDM2B/H3K36C-UNC8015 nucleosome (middle), and KDM2A/H3K36C-UNC8015 nucleosome (right) complex reconstitutions. These experiments were performed once. b, Highest populated 2D classes attained from cryo-EM analysis of indicated samples, with selected top views of KDM2A/B-unbound and bound nucleosomes.
Extended Data Fig. 4 Design and synthesis of H3K36-JmjC inhibitor covalent conjugate.
a, Chemical structures of JmjC inhibitors. b, Inhibition assays using H3K36me2 peptide substrates with KDM2B(2–734). c, Dose response inhibition of KDM2B(2–734) by UNC8016 on H3K36me2 peptide substrates. In b and c, mean ± SD are shown (n = 3 assay technical replicates). d, Synthetic scheme for alkylation reaction of H3K36C mutant with UNC8015. e, LCMS spectra and mass deconvolution (determined using ESIprot) of unreacted H3K36C (left) and reacted H3K36C-UNC8015 (right; 5-hour endpoint).
Extended Data Fig. 5 Crystal structure of KDM2A bound to H3K36-UNC8015 peptide covalent conjugate.
a, Structure of KDM2A catalytic domain in complex with H3K36C-UNC8015. b, Identical view of structure of KDM2A catalytic domain in complex with H3K36me2 peptide and α-ketoglutarate (α-KG). c-e, Zoomed views of c, H3K36C-UNC8015 peptide, d, H3K36me2 peptide and α-KG, and e, overlayed aligned structures. f, 2Fo-Fc electron density map of H3K36C-UNC8015 peptide at 1σ.
Extended Data Fig. 6 Cryo-EM data analysis for KDM2A-nucleosome complex.
a, Representative micrograph (left), CTF estimation (middle), and 2D classes (right) from KDM2A-nucleosome complex dataset. b, Cryo-EM data analysis workflow carried out in RELION-3.1. c, Multibody refinement analysis results from RELION. d, Final postprocessed map colored by local resolution (top), angular distribution of particles (middle), and FSC curve (bottom) for KDM2A-nucleosome complex.
Extended Data Fig. 7 Cryo-EM data analysis for KDM2B-nucleosome complex.
a, Representative micrograph (left), CTF estimation (middle), and 2D classes (right) from KDM2B-nucleosome complex dataset. b, Cryo-EM data analysis workflow carried out in RELION-3.1. c, Multibody refinement analysis results from RELION. d, Final postprocessed map colored by local resolution (top), angular distribution of particles (middle), and FSC curve (bottom) for KDM2B-nucleosome complex.
Extended Data Fig. 8 Unwrapped DNA-binding interfaces of KDM2A/B.
a, Sequence alignment of KDM2A and KDM2B DNA-binding interfaces with closely related KDM7A and PHF8 (KDM7B), with conserved basic amino acids and charge swapped amino acids colored blue and red, respectively. b, Zoomed view of unwrapped DNA-binding interface of KDM2A structure (left) and KDM2B structure (right). c, TR-FRET binding isotherms with KDM2A DNA interface mutants and unmodified wild-type 185 bp nucleosomes. d, TR-FRET nucleosome binding competition experiments using unmodified wildtype 185, 147, or 119 bp nucleosomes. In c and d, mean ± SD are shown (n = 3 assay technical replicates).
Extended Data Fig. 9 Acidic patch-localized density of KDM2A/B.
a, b, Acidic patch-localized density in nucleosome-focused cryo-EM maps from multibody analysis of a KDM2A- and b KDM2B-nucleosome complexes at indicated map thresholds. c, Overlay of acidic patch-localized density from nucleosome-focused map shown in a with equivalent region of modeled KDM2A. d, Sequence alignment of KDM2A and KDM2B N-termini, with basic amino acids colored blue.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2, Supplementary Figs. 1–7 and Supplementary Note
Supplementary Data 1
Nucleosome affinity proteomics data and statistics
Supplementary Data 2
Profile plots from nucleosome affinity proteomics
Supplementary Data 3
KDM2B–nucleosome model
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Spangler, C.J., Skrajna, A., Foley, C.A. et al. Structural basis of paralog-specific KDM2A/B nucleosome recognition. Nat Chem Biol 19, 624–632 (2023). https://doi.org/10.1038/s41589-023-01256-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-023-01256-y
This article is cited by
-
Interrogating epigenetic mechanisms with chemically customized chromatin
Nature Reviews Genetics (2024)
-
Paralog-specific recognition
Nature Chemical Biology (2023)